

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0316.5
(318 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
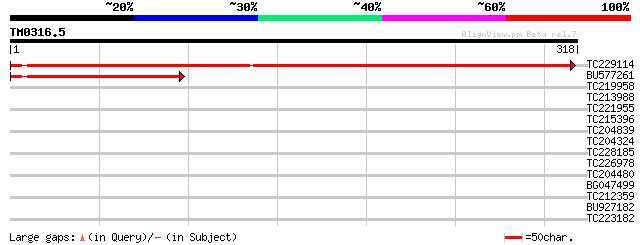
Score E
Sequences producing significant alignments: (bits) Value
TC229114 similar to UP|Q9LQQ6 (Q9LQQ6) F24B9.6, partial (42%) 485 e-137
BU577261 157 6e-39
TC219958 32 0.29
TC213988 32 0.50
TC221955 weakly similar to UP|Q6UAL1 (Q6UAL1) Myosin heavy chain... 30 1.4
TC215396 weakly similar to UP|Q8H9F4 (Q8H9F4) 41 kD chloroplast ... 29 2.5
TC204839 similar to UP|Q9LQL8 (Q9LQL8) F5D14.17 protein, partial... 29 2.5
TC204324 similar to UP|NTF2_ARATH (Q9C7F5) Nuclear transport fac... 29 3.2
TC228185 similar to UP|Q9LUA5 (Q9LUA5) Arabidopsis thaliana geno... 29 3.2
TC226978 28 4.2
TC204480 similar to UP|Q947C3 (Q947C3) 1-deoxy-D-xylulose-5-phos... 28 4.2
BG047499 28 5.5
TC212359 homologue to GB|AAP42741.1|30984556|BT008728 At3g58120 ... 28 5.5
BU927182 27 9.4
TC223182 similar to UP|Q84UX6 (Q84UX6) Nucleosome/chromatin asse... 27 9.4
>TC229114 similar to UP|Q9LQQ6 (Q9LQQ6) F24B9.6, partial (42%)
Length = 1476
Score = 485 bits (1249), Expect = e-137
Identities = 249/317 (78%), Positives = 277/317 (86%)
Frame = +2
Query: 1 MAETVNHLNNHDSKEKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYL 60
M T H N DSK+KEA LAALLKEMKEGLD VR KIQSLTA VKEGQYPTADGFSYL
Sbjct: 170 MENTAGH--NDDSKQKEAPKLAALLKEMKEGLDTVRRKIQSLTATVKEGQYPTADGFSYL 343
Query: 61 EAKNLLLLNYCQSLVYYLLRKAKGFSIEEHPVVRSIVEIRLYLEKIRPIDKKQQYQIQKL 120
EAKNLLLLNYCQSLVYYLLRKAKG SIE+HPVVRS+VEIRL+LEKIRPIDKKQQYQI KL
Sbjct: 344 EAKNLLLLNYCQSLVYYLLRKAKGLSIEDHPVVRSVVEIRLFLEKIRPIDKKQQYQIXKL 523
Query: 121 MKVGENAATSDLPSKKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPT 180
M+ ENA SD+ +K EPVAS+KSED SKYRPNPDMLVS+ DL Q ++ Y+PVKFAPT
Sbjct: 524 MQASENAIRSDILNK-EPVASNKSEDASKYRPNPDMLVSKVDLTSQADNEYYQPVKFAPT 700
Query: 181 SMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIA 240
SMD +K+SK ERNALRREKEILK++K S +IRTLMND+EE+PEEIRD+EG+S+EV+R+IA
Sbjct: 701 SMDLEKSSKHERNALRREKEILKQAKQSDYIRTLMNDMEERPEEIRDFEGASREVDRYIA 880
Query: 241 KMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLPFEDETGEQV 300
KM+ERA QEEELF RVPLTK ERKREKYLKKSRNGL GLTESF+DEIKTLPFED+TGEQV
Sbjct: 881 KMDERARQEEELFTRVPLTKQERKREKYLKKSRNGLQGLTESFYDEIKTLPFEDKTGEQV 1060
Query: 301 MGPSKGSNRNGRLKKRK 317
+G S GS N RLKKRK
Sbjct: 1061VGSSNGSRTNNRLKKRK 1111
>BU577261
Length = 428
Score = 157 bits (397), Expect = 6e-39
Identities = 81/98 (82%), Positives = 84/98 (85%)
Frame = +1
Query: 1 MAETVNHLNNHDSKEKEASPLAALLKEMKEGLDVVRHKIQSLTAKVKEGQYPTADGFSYL 60
M NH N DSK+KEA LAALLKEMKEGLD VRHKIQSLTA VKEG YPTADGFSYL
Sbjct: 139 MENIANH--NDDSKQKEAPQLAALLKEMKEGLDTVRHKIQSLTAMVKEGLYPTADGFSYL 312
Query: 61 EAKNLLLLNYCQSLVYYLLRKAKGFSIEEHPVVRSIVE 98
E KNLLLLNYCQSLVYYLLRKAKG SIE+HPVVRS+VE
Sbjct: 313 EVKNLLLLNYCQSLVYYLLRKAKGLSIEDHPVVRSVVE 426
>TC219958
Length = 989
Score = 32.3 bits (72), Expect = 0.29
Identities = 35/146 (23%), Positives = 65/146 (43%), Gaps = 7/146 (4%)
Frame = +1
Query: 135 KKEPVASDKSEDVSKYRPNPDMLVSREDLAPQEGHDAYRPVKFAPTSMDQDKTSKQERNA 194
K PVA ++E+ S++ + DL P R P +K KQ R
Sbjct: 139 KSPPVALKRAEERSEWWAVDGEVHEIGDLVP------LRERFVIPRENIPNKRRKQLREQ 300
Query: 195 -LRREKEILKESKHSSFIRTLM---NDIEEKPEEIRDYEGSSKEVERHIAKMEERAWQEE 250
+RR + +LKES+H + + M N++ E E + EG +K++ + A E +
Sbjct: 301 FMRRTRLVLKESEHDPWCKKYMELYNELRENWERLYWDEGYTKKLAQDHANYESAEDDDG 480
Query: 251 ELF---NRVPLTKHERKREKYLKKSR 273
+ +R P ++ E +++ ++R
Sbjct: 481 DFSPYRSRRPQSQMEHSKDQNFGRNR 558
>TC213988
Length = 776
Score = 31.6 bits (70), Expect = 0.50
Identities = 24/124 (19%), Positives = 52/124 (41%), Gaps = 15/124 (12%)
Frame = -2
Query: 186 KTSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKE----------- 234
K + E LR +++ +E HSSF + ND++ K ++ +++E
Sbjct: 526 KKADCEIRELRDLEDLCREVLHSSF--KVANDVKSKEAVFNSFDSATQERHPLKEEIDRF 353
Query: 235 ----VERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTL 290
+E+ ++ E + + V + KH Y +K ++ +DGL ++L
Sbjct: 352 SAPFIEKSVSSCSEGQTHDSKF---VDMEKHSAFENGYARKDQDTVDGLKHKIGQSHRSL 182
Query: 291 PFED 294
++
Sbjct: 181 ELDE 170
>TC221955 weakly similar to UP|Q6UAL1 (Q6UAL1) Myosin heavy chain class XI E3
protein, partial (6%)
Length = 728
Score = 30.0 bits (66), Expect = 1.4
Identities = 22/84 (26%), Positives = 43/84 (51%), Gaps = 7/84 (8%)
Frame = +1
Query: 214 LMNDIEEKPEEIRDYEGSSKEVE-RHIAKMEE------RAWQEEELFNRVPLTKHERKRE 266
L ++EE ++I+++E S E+E + A+++E +A Q +E R+ L+ + E
Sbjct: 190 LETEVEELKKKIKEFEESYSEIENENQARLKEAEEAQLKATQLQETIERLELSLSNLESE 369
Query: 267 KYLKKSRNGLDGLTESFFDEIKTL 290
+ + + E F+EIK L
Sbjct: 370 NQVLCQKALEEPKNEELFEEIKIL 441
>TC215396 weakly similar to UP|Q8H9F4 (Q8H9F4) 41 kD chloroplast nucleoid DNA
binding protein (CND41), partial (69%)
Length = 1808
Score = 29.3 bits (64), Expect = 2.5
Identities = 12/34 (35%), Positives = 20/34 (58%)
Frame = +3
Query: 6 NHLNNHDSKEKEASPLAALLKEMKEGLDVVRHKI 39
+ LNNHD K K +P + +L + KE + + +I
Sbjct: 252 SQLNNHDGKAKSTTPHSDILNQDKERVKYINSRI 353
>TC204839 similar to UP|Q9LQL8 (Q9LQL8) F5D14.17 protein, partial (94%)
Length = 1262
Score = 29.3 bits (64), Expect = 2.5
Identities = 23/80 (28%), Positives = 37/80 (45%), Gaps = 5/80 (6%)
Frame = -3
Query: 88 EEHPVVRSIVEIRLYLEKIRPI-----DKKQQYQIQKLMKVGENAATSDLPSKKEPVASD 142
+ ++RS + ++ ++I PI K QY++QKL K + TS P +A
Sbjct: 312 DSESIIRSFI---VFNKQINPIADHGQTSKGQYEVQKLQKAFPASPTSHPP--PPIIALQ 148
Query: 143 KSEDVSKYRPNPDMLVSRED 162
SK+RP P+ R D
Sbjct: 147 SLNQNSKWRPVPNTF*DRID 88
>TC204324 similar to UP|NTF2_ARATH (Q9C7F5) Nuclear transport factor 2
(NTF-2), complete
Length = 772
Score = 28.9 bits (63), Expect = 3.2
Identities = 12/43 (27%), Positives = 26/43 (59%)
Frame = -1
Query: 176 KFAPTSMDQDKTSKQERNALRREKEILKESKHSSFIRTLMNDI 218
KF+P+++ Q T + + + ++EILK+ +H+ T N++
Sbjct: 676 KFSPSNLHQSSTIYR*KRFDQSQQEILKQKRHNYAFNTNQNEL 548
>TC228185 similar to UP|Q9LUA5 (Q9LUA5) Arabidopsis thaliana genomic DNA,
chromosome 3, P1 clone: MPE11, partial (14%)
Length = 1102
Score = 28.9 bits (63), Expect = 3.2
Identities = 27/117 (23%), Positives = 53/117 (45%), Gaps = 8/117 (6%)
Frame = +3
Query: 198 EKEILKESKHSSFIRTLMNDIEEK-------PE-EIRDYEGSSKEVERHIAKMEERAWQE 249
+K + ES +S +T + D E PE +IR + S +++ + ++ +
Sbjct: 102 KKRVRDESSYSPESQTQLVDSPESESQRVDSPESKIRRVDSSESQLD---SVLDSEVRLQ 272
Query: 250 EELFNRVPLTKHERKREKYLKKSRNGLDGLTESFFDEIKTLPFEDETGEQVMGPSKG 306
+++FN + + +R+ S GLD + +SF DEI + +G+ P G
Sbjct: 273 DDIFNILDDADNVPERD-----SVQGLDSVIKSFEDEINAPGSDSGSGDPTQEPDSG 428
>TC226978
Length = 1994
Score = 28.5 bits (62), Expect = 4.2
Identities = 32/146 (21%), Positives = 56/146 (37%), Gaps = 4/146 (2%)
Frame = +3
Query: 131 DLPSKKEPVASDKSEDVSKYRPN--PDMLVSREDLAPQE--GHDAYRPVKFAPTSMDQDK 186
++P K E + + D K N D+ ++ED A G P +++DK
Sbjct: 240 NVPQKPESITGTRRTDSPKPEENFFSDLSKAKEDNASLNLNGRGFLVANSNKPVEIEEDK 419
Query: 187 TSKQERNALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERA 246
S+ EI + KH S + ++D EGS+ A+ E+ A
Sbjct: 420 RSEAANKRKMPFDEIRDQKKHDSDVHADLHDRARASHISITEEGST-------AENEDVA 578
Query: 247 WQEEELFNRVPLTKHERKREKYLKKS 272
E E P++ H + +++ S
Sbjct: 579 DSETENSTSRPISHHSDGSKGFIRVS 656
>TC204480 similar to UP|Q947C3 (Q947C3) 1-deoxy-D-xylulose-5-phosphate
reductoisomerase, partial (92%)
Length = 2110
Score = 28.5 bits (62), Expect = 4.2
Identities = 29/79 (36%), Positives = 36/79 (44%), Gaps = 7/79 (8%)
Frame = +2
Query: 167 EGHDAYRPVKFAPTS-----MDQDKTSKQERNALRREKEI--LKESKHSSFIRTLMNDIE 219
E D ++ V A S DQ K K + A+R E I LKE+ H D+E
Sbjct: 575 ENPDKFKVVALAAGSNVTLLADQVKRFKPQLVAVRNESLIAELKEALH---------DVE 727
Query: 220 EKPEEIRDYEGSSKEVERH 238
EKPE I +G EV RH
Sbjct: 728 EKPEIIPGEQGII-EVARH 781
>BG047499
Length = 422
Score = 28.1 bits (61), Expect = 5.5
Identities = 17/49 (34%), Positives = 27/49 (54%), Gaps = 2/49 (4%)
Frame = -1
Query: 3 ETVNHLNNHDSKEKEASP--LAALLKEMKEGLDVVRHKIQSLTAKVKEG 49
+++N N SK + P L+ +L +K L RHK+ SLT +V +G
Sbjct: 284 KSLNL*KNSSSKSEIDLPSLLSIILAALKRNLRRQRHKMHSLTVQVSDG 138
>TC212359 homologue to GB|AAP42741.1|30984556|BT008728 At3g58120 {Arabidopsis
thaliana;} , partial (34%)
Length = 664
Score = 28.1 bits (61), Expect = 5.5
Identities = 16/39 (41%), Positives = 24/39 (61%)
Frame = +3
Query: 107 RPIDKKQQYQIQKLMKVGENAATSDLPSKKEPVASDKSE 145
R I++ +Q Q+ +K ENAA S LPS K P+ ++E
Sbjct: 345 REIERLRQVYYQQSLKKMENAAGSPLPSPK-PICDAQTE 458
>BU927182
Length = 447
Score = 27.3 bits (59), Expect = 9.4
Identities = 15/55 (27%), Positives = 26/55 (47%)
Frame = +3
Query: 219 EEKPEEIRDYEGSSKEVERHIAKMEERAWQEEELFNRVPLTKHERKREKYLKKSR 273
E K E +++EG V + E+ ++E + K R+REK ++K R
Sbjct: 60 ERKKEFEKEHEGERVRVREKDRREREKEVEKERREREKEVEKDRREREKEVEKDR 224
>TC223182 similar to UP|Q84UX6 (Q84UX6) Nucleosome/chromatin assembly factor
group A, partial (29%)
Length = 442
Score = 27.3 bits (59), Expect = 9.4
Identities = 22/89 (24%), Positives = 41/89 (45%)
Frame = +3
Query: 193 NALRREKEILKESKHSSFIRTLMNDIEEKPEEIRDYEGSSKEVERHIAKMEERAWQEEEL 252
NAL+ + ++L +H + TL + ++ E +RD + EVE +EER+ E
Sbjct: 135 NALKDKLQLLA-GQHVDILETLPPKVRQRVEILRDLQSQHDEVEAKF--LEERSQLE--- 296
Query: 253 FNRVPLTKHERKREKYLKKSRNGLDGLTE 281
K+++ E K ++G+ E
Sbjct: 297 ------AKYQKLYEPLYSKRYEIVNGVVE 365
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.311 0.130 0.352
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,512,609
Number of Sequences: 63676
Number of extensions: 117089
Number of successful extensions: 564
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 561
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 563
length of query: 318
length of database: 12,639,632
effective HSP length: 97
effective length of query: 221
effective length of database: 6,463,060
effective search space: 1428336260
effective search space used: 1428336260
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0316.5


