

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
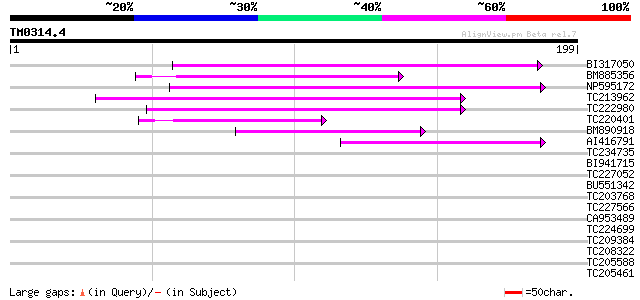
Query= TM0314.4
(199 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BI317050 85 2e-17
BM885356 81 4e-16
NP595172 polyprotein [Glycine max] 74 4e-14
TC213962 66 9e-12
TC222980 similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, partial (4%) 58 3e-09
TC220401 49 1e-06
BM890918 44 4e-05
AI416791 43 8e-05
TC234735 38 0.004
BI941715 31 0.33
TC227052 weakly similar to GB|AAM19768.1|20453045|AY094388 AT3g5... 30 0.56
BU551342 weakly similar to PIR|C86478|C8 protein F15O4.13 [impor... 30 0.96
TC203768 similar to UP|EF1D_ORYSA (P29545) Elongation factor 1-b... 28 2.1
TC227566 similar to UP|Q9MA20 (Q9MA20) T5E21.13, partial (7%) 28 2.1
CA953489 similar to GP|21553000|gb Drm4 {Pisum sativum}, partial... 28 3.6
TC224699 similar to UP|Q8LKG0 (Q8LKG0) Drm4, partial (95%) 28 3.6
TC209384 homologue to UP|Q71SQ1 (Q71SQ1) MYC1, partial (41%) 27 4.8
TC208322 27 4.8
TC205588 similar to UP|Q9FH57 (Q9FH57) GATA-binding transcriptio... 27 6.2
TC205461 similar to GB|AAN28790.1|23505861|AY143851 At5g64200/MS... 27 6.2
>BI317050
Length = 425
Score = 85.1 bits (209), Expect = 2e-17
Identities = 44/130 (33%), Positives = 69/130 (52%)
Frame = +3
Query: 58 VDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERAW 117
+D+P FD ++A I ++ +FF R E+ERL++ ++G+AL WFQW +W
Sbjct: 12 LDVPRFDDTNAPTLIFKISQFFDYHRTPEDERLQVTSFYLDGEALSWFQWLYRNDQLTSW 191
Query: 118 EPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADREILKGVFLN 177
F AL RF ++ +P G L + Q GSV ++ FE A + G L F++
Sbjct: 192 SSFLQALEMRFAQSMYDDPRGTLFKLTQTGSVNSFLAMFEALANWIVGLPTPFLLSCFIS 371
Query: 178 GLQEEIRAEM 187
GL +IR E+
Sbjct: 372 GLLPDIRREV 401
>BM885356
Length = 420
Score = 80.9 bits (198), Expect = 4e-16
Identities = 39/94 (41%), Positives = 51/94 (53%)
Frame = -3
Query: 45 PHRVHLRAVTGRRVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGW 104
PHR+ L D+P FDGSDA GWI ++ +FF+ + ERL + MEG+AL W
Sbjct: 259 PHRLKL--------DVPRFDGSDATGWIFKITQFFEYHTTPDHERLTIASFYMEGQALAW 104
Query: 105 FQWWEEQAPERAWEPFKHALIRRFQPTLVQNPFG 138
FQW +W F HAL RF T ++P G
Sbjct: 103 FQWMHRNGQLSSWPAFLHALHSRFASTSYEDPTG 2
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 73.9 bits (180), Expect = 4e-14
Identities = 37/132 (28%), Positives = 63/132 (47%)
Frame = +1
Query: 57 RVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERA 116
++D P FDG + WI + E+FF + +RL + + ++ + W+Q ++ P +
Sbjct: 295 KLDFPRFDGKNVMDWIFKAEQFFDYYATPDADRLIIASVHLDQDVVPWYQMLQKTEPFSS 474
Query: 117 WEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADREILKGVFL 176
W+ F AL F P+ P L + Q +V Y F + G E + F+
Sbjct: 475 WQAFTRALELDFGPSAYDCPRATLFKLNQSATVNEYYMQFTALVNRVDGLSAEAILDCFV 654
Query: 177 NGLQEEIRAEMK 188
+GLQEEI ++K
Sbjct: 655 SGLQEEISRDVK 690
>TC213962
Length = 799
Score = 66.2 bits (160), Expect = 9e-12
Identities = 36/130 (27%), Positives = 57/130 (43%)
Frame = -2
Query: 31 GSRFSDHSRERFDHPHRVHLRAVTGRRVDIPMFDGSDAYGWINRVERFFQLSRVGEEERL 90
G+ + S + F + T ++++P FDG D GWI ++ +F + E L
Sbjct: 456 GTICCNSSHQHFAQIPTIFPHPCTHMKLEVPWFDGQDPLGWIFKITQFSNYQGIPNVEHL 277
Query: 91 EMVMIAMEGKALGWFQWWEEQAPERAWEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVM 150
+ MEG AL W++W W HAL F P+ +P L ++Q+G V
Sbjct: 276 IVGSFNMEGPALCWYRWMSWNGFLTTWSAMLHALESCFTPSF*DDPHDALFKLQQRGFVN 97
Query: 151 AYRESFEEAA 160
Y F+ A
Sbjct: 96 DYLTEFKRFA 67
>TC222980 similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, partial (4%)
Length = 662
Score = 58.2 bits (139), Expect = 3e-09
Identities = 32/112 (28%), Positives = 52/112 (45%)
Frame = +3
Query: 49 HLRAVTGRRVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWFQWW 108
H V ++D P+ DGSD WI + E+FF + E+RL + I ++ + + +Q
Sbjct: 321 HPFQVRNVKIDFPILDGSDVLQWIFKAEQFFNYHKTPGEQRLIIASIHLDKEVVPCYQMM 500
Query: 109 EEQAPERAWEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAA 160
+ + W F AL F P+ + P L + + GSV +Y F A
Sbjct: 501 TRENSFKTWIAFTRALETEFGPSPYECPRSQLFKLAKIGSVQSYYVQFTALA 656
>TC220401
Length = 1286
Score = 49.3 bits (116), Expect = 1e-06
Identities = 23/66 (34%), Positives = 35/66 (52%)
Frame = +1
Query: 46 HRVHLRAVTGRRVDIPMFDGSDAYGWINRVERFFQLSRVGEEERLEMVMIAMEGKALGWF 105
HR H++ +D P F+G D GWI ++ +FF + E+E L + M+G AL +
Sbjct: 232 HRPHMK------LDAPRFNGHDPLGWIFKISQFFDYQTILEQEPLTVASFYMDGSALH*Y 393
Query: 106 QWWEEQ 111
QW Q
Sbjct: 394 QWMSRQ 411
>BM890918
Length = 421
Score = 44.3 bits (103), Expect = 4e-05
Identities = 21/67 (31%), Positives = 34/67 (50%)
Frame = +1
Query: 80 QLSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERAWEPFKHALIRRFQPTLVQNPFGP 139
++ + L+ ++MEG A WF W + P+ WE F ALI+RF+ + F
Sbjct: 76 EVQEIPSFHNLQYAFMSMEGAATHWFYIWSQNNPDSNWESFSTALIKRFRDHHGNHIFER 255
Query: 140 LLSIKQK 146
L +KQ+
Sbjct: 256 LSILKQE 276
>AI416791
Length = 420
Score = 43.1 bits (100), Expect = 8e-05
Identities = 25/72 (34%), Positives = 37/72 (50%)
Frame = -3
Query: 117 WEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGADREILKGVFL 176
WE K AL+ R+ + + L +KQ+GSV Y FE A + + +G FL
Sbjct: 373 WESLKIALLERYGGHGDGDVYEQLTELKQEGSVEDYITEFEYLIAQIPRLPEKQFQGYFL 194
Query: 177 NGLQEEIRAEMK 188
+GLQ EIR ++
Sbjct: 193 HGLQSEIRGRVR 158
>TC234735
Length = 553
Score = 37.7 bits (86), Expect = 0.004
Identities = 14/26 (53%), Positives = 20/26 (76%)
Frame = +1
Query: 88 ERLEMVMIAMEGKALGWFQWWEEQAP 113
E++ VM+ MEG+ALGWF+ WE +P
Sbjct: 181 EKVPAVMVVMEGRALGWFRRWESCSP 258
>BI941715
Length = 282
Score = 31.2 bits (69), Expect = 0.33
Identities = 12/49 (24%), Positives = 24/49 (48%)
Frame = +1
Query: 81 LSRVGEEERLEMVMIAMEGKALGWFQWWEEQAPERAWEPFKHALIRRFQ 129
+ R +E ++++ + MEG + WF E W + A+I R++
Sbjct: 31 VQRTSKEVKVKLTKLNMEGSTIHWFNSLYETEDHLTWLELRQAMIERYR 177
>TC227052 weakly similar to GB|AAM19768.1|20453045|AY094388
AT3g56200/F18O21_160 {Arabidopsis thaliana;} , partial
(68%)
Length = 1627
Score = 30.4 bits (67), Expect = 0.56
Identities = 17/38 (44%), Positives = 20/38 (51%), Gaps = 1/38 (2%)
Frame = +2
Query: 3 RGRSNTPRFRQASRERDR-EASSHSGSRTGSRFSDHSR 39
R R R Q SR R+ ++ H GSR SRF D SR
Sbjct: 146 RPRRGVQRGHQHSRRRNHVDSGDHEGSRRSSRFRDDSR 259
>BU551342 weakly similar to PIR|C86478|C8 protein F15O4.13 [imported] -
Arabidopsis thaliana, partial (5%)
Length = 467
Score = 29.6 bits (65), Expect = 0.96
Identities = 25/88 (28%), Positives = 39/88 (43%), Gaps = 4/88 (4%)
Frame = -3
Query: 116 AWEPFKHALIRRFQPTLVQNPFGPLLSIKQKGSVMAYRESFEEAAAPMRGA----DREIL 171
+WE K + RRF P+L Q L +GS + E ++E M A D E
Sbjct: 447 SWEXMKXLMRRRFVPSLYQRELHNKLQRLTQGS-RSVDEYYKEMEVSMIRANVMED*EAT 271
Query: 172 KGVFLNGLQEEIRAEMKLYPAADLAGLM 199
FL+GL ++R ++ +L L+
Sbjct: 270 MARFLHGLNSDVRDVVEWQNHVELENLV 187
>TC203768 similar to UP|EF1D_ORYSA (P29545) Elongation factor 1-beta'
(EF-1-beta'), partial (88%)
Length = 1419
Score = 28.5 bits (62), Expect = 2.1
Identities = 18/65 (27%), Positives = 31/65 (47%)
Frame = +2
Query: 10 RFRQASRERDREASSHSGSRTGSRFSDHSRERFDHPHRVHLRAVTGRRVDIPMFDGSDAY 69
R R+ RER+RE S + R PHR+ R+ RR +P+++G +
Sbjct: 89 RERERERERERERERERESSNSCHSLQNGRYLLKSPHRI--RSQVPRR--LPLWEGLRFW 256
Query: 70 GWINR 74
G +++
Sbjct: 257 GSVHK 271
>TC227566 similar to UP|Q9MA20 (Q9MA20) T5E21.13, partial (7%)
Length = 819
Score = 28.5 bits (62), Expect = 2.1
Identities = 9/24 (37%), Positives = 14/24 (57%)
Frame = -2
Query: 99 GKALGWFQWWEEQAPERAWEPFKH 122
G++L W+QWW +R W +H
Sbjct: 308 GRSLLWWQWWWHSLRQRWWMHHRH 237
>CA953489 similar to GP|21553000|gb Drm4 {Pisum sativum}, partial (94%)
Length = 426
Score = 27.7 bits (60), Expect = 3.6
Identities = 19/56 (33%), Positives = 26/56 (45%), Gaps = 7/56 (12%)
Frame = -1
Query: 8 TPRFRQASRERDREASSHSGSRTGSR-------FSDHSRERFDHPHRVHLRAVTGR 56
T F + + R R + + GSR+GS F DH R H HR+ R +T R
Sbjct: 321 TEGFSSSRKRRHRRSRTGGGSRSGSTGTLVTRWFHDHYTSR--HSHRILGRILTVR 160
>TC224699 similar to UP|Q8LKG0 (Q8LKG0) Drm4, partial (95%)
Length = 750
Score = 27.7 bits (60), Expect = 3.6
Identities = 19/56 (33%), Positives = 26/56 (45%), Gaps = 7/56 (12%)
Frame = -2
Query: 8 TPRFRQASRERDREASSHSGSRTGSR-------FSDHSRERFDHPHRVHLRAVTGR 56
T F + + R R + + GSR+GS F DH R H HR+ R +T R
Sbjct: 305 TEGFSSSRKRRHRRSRTGGGSRSGSTGTLVTRWFHDHYTSR--HSHRILGRILTVR 144
>TC209384 homologue to UP|Q71SQ1 (Q71SQ1) MYC1, partial (41%)
Length = 1054
Score = 27.3 bits (59), Expect = 4.8
Identities = 8/19 (42%), Positives = 9/19 (47%)
Frame = -1
Query: 102 LGWFQWWEEQAPERAWEPF 120
+GW WW R W PF
Sbjct: 145 IGWQWWWRRGPRRRRWSPF 89
>TC208322
Length = 1090
Score = 27.3 bits (59), Expect = 4.8
Identities = 14/38 (36%), Positives = 19/38 (49%)
Frame = +3
Query: 102 LGWFQWWEEQAPERAWEPFKHALIRRFQPTLVQNPFGP 139
+G ++ E PE PF LIRRF + + NP P
Sbjct: 711 MGGIMYYLEHEPE-TMAPFLRGLIRRFLASRISNPSPP 821
>TC205588 similar to UP|Q9FH57 (Q9FH57) GATA-binding transcription
factor-like protein (At5g66320), partial (25%)
Length = 1084
Score = 26.9 bits (58), Expect = 6.2
Identities = 16/38 (42%), Positives = 20/38 (52%)
Frame = -1
Query: 2 TRGRSNTPRFRQASRERDREASSHSGSRTGSRFSDHSR 39
T G +T RF AS ER S SGS + ++ HSR
Sbjct: 367 TEGSGSTNRF--ASSERTGYKSDSSGSESSTKCETHSR 260
>TC205461 similar to GB|AAN28790.1|23505861|AY143851 At5g64200/MSJ1_4
{Arabidopsis thaliana;} , partial (69%)
Length = 1148
Score = 26.9 bits (58), Expect = 6.2
Identities = 16/50 (32%), Positives = 22/50 (44%)
Frame = +3
Query: 3 RGRSNTPRFRQASRERDREASSHSGSRTGSRFSDHSRERFDHPHRVHLRA 52
R RS +PR R RDR+ S SR+ R+ D +R R+
Sbjct: 435 RSRSRSPRKRHRDDYRDRDYRRRSRSRSYDRYERDRHRGRDKDYRRRSRS 584
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.136 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,851,218
Number of Sequences: 63676
Number of extensions: 104208
Number of successful extensions: 752
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 738
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 746
length of query: 199
length of database: 12,639,632
effective HSP length: 92
effective length of query: 107
effective length of database: 6,781,440
effective search space: 725614080
effective search space used: 725614080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Lotus: description of TM0314.4


