

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
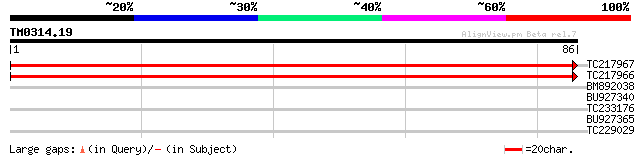
Query= TM0314.19
(86 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC217967 homologue to UP|Q9LX13 (Q9LX13) (3R)-hydroxymyristoyl-[... 152 4e-38
TC217966 homologue to UP|Q9LX13 (Q9LX13) (3R)-hydroxymyristoyl-[... 149 3e-37
BM892038 25 5.2
BU927340 similar to GP|18087570|gb AT3g54650/T5N23_10 {Arabidops... 25 6.8
TC233176 weakly similar to GB|AAL07033.1|15810291|AY056184 auxin... 25 6.8
BU927365 25 8.9
TC229029 similar to UP|Q84XI2 (Q84XI2) Glycosyl hydrolase, parti... 25 8.9
>TC217967 homologue to UP|Q9LX13 (Q9LX13) (3R)-hydroxymyristoyl-[acyl carrier
protein] dehydratase-like protein (At5g10160), partial
(49%)
Length = 590
Score = 152 bits (383), Expect = 4e-38
Identities = 78/86 (90%), Positives = 80/86 (92%)
Frame = +3
Query: 1 LLLQAMAQVGVLVMLQPEGGGSRENFFFVGIDKVRFCKPVIAGDTLVMRMTLTKLQKSFE 60
LLLQAMAQVG LVMLQPE GGSRENFFF GIDKVRF KPVIAGDTLVMRMTLTKLQK F
Sbjct: 180 LLLQAMAQVGGLVMLQPEVGGSRENFFFAGIDKVRFRKPVIAGDTLVMRMTLTKLQKRFG 359
Query: 61 LAKMEGKAYVGGEVVCEGEFLMAIGS 86
+AKMEGKAYVGGEVVCEGEFLMA+GS
Sbjct: 360 IAKMEGKAYVGGEVVCEGEFLMAMGS 437
>TC217966 homologue to UP|Q9LX13 (Q9LX13) (3R)-hydroxymyristoyl-[acyl carrier
protein] dehydratase-like protein (At5g10160), partial
(71%)
Length = 910
Score = 149 bits (375), Expect = 3e-37
Identities = 75/86 (87%), Positives = 80/86 (92%)
Frame = +2
Query: 1 LLLQAMAQVGVLVMLQPEGGGSRENFFFVGIDKVRFCKPVIAGDTLVMRMTLTKLQKSFE 60
L+++AMAQVG LVMLQPE GGSRENFFF GIDKVRF KPVIAGDTLVMRMTLTKLQK F
Sbjct: 503 LMVEAMAQVGGLVMLQPEVGGSRENFFFAGIDKVRFRKPVIAGDTLVMRMTLTKLQKRFG 682
Query: 61 LAKMEGKAYVGGEVVCEGEFLMAIGS 86
+AKMEGKAYVGGEVVCEGEFLMA+GS
Sbjct: 683 IAKMEGKAYVGGEVVCEGEFLMAMGS 760
>BM892038
Length = 422
Score = 25.4 bits (54), Expect = 5.2
Identities = 18/63 (28%), Positives = 30/63 (47%), Gaps = 5/63 (7%)
Frame = -3
Query: 17 PEGGG----SRENFFFVGIDKVRFCKPVIAGDTLVMRMTLTKLQK-SFELAKMEGKAYVG 71
P GGG + F V KV F ++ G + R ++++ SF +EG+ +G
Sbjct: 309 PGGGGRFSGEKREFMVVL*KKVVFLLVMVLGRVQIGRRRRRRIRRRSFGNGDLEGRRCIG 130
Query: 72 GEV 74
GE+
Sbjct: 129 GEM 121
>BU927340 similar to GP|18087570|gb AT3g54650/T5N23_10 {Arabidopsis
thaliana}, partial (23%)
Length = 446
Score = 25.0 bits (53), Expect = 6.8
Identities = 15/43 (34%), Positives = 24/43 (54%), Gaps = 6/43 (13%)
Frame = +2
Query: 40 VIAGDTLVMRMTLTKL------QKSFELAKMEGKAYVGGEVVC 76
+ DT V R++ +L +KS + KMEG + +GG V+C
Sbjct: 68 ISVSDTAVNRISGVELARFVADKKSLKSLKMEGCSNLGGFVLC 196
>TC233176 weakly similar to GB|AAL07033.1|15810291|AY056184 auxin response
factor ARF17 {Arabidopsis thaliana;} , partial (17%)
Length = 958
Score = 25.0 bits (53), Expect = 6.8
Identities = 11/28 (39%), Positives = 17/28 (60%)
Frame = +2
Query: 20 GGSRENFFFVGIDKVRFCKPVIAGDTLV 47
G R +FF +G K K ++AGDT++
Sbjct: 392 GTPRHHFFTIGWSKFVNHKKLVAGDTVI 475
>BU927365
Length = 445
Score = 24.6 bits (52), Expect = 8.9
Identities = 12/28 (42%), Positives = 17/28 (59%)
Frame = -1
Query: 52 LTKLQKSFELAKMEGKAYVGGEVVCEGE 79
+ KL+ +F LAK G + + E VCE E
Sbjct: 274 VAKLRSNFVLAKPGGTSGLAAEGVCERE 191
>TC229029 similar to UP|Q84XI2 (Q84XI2) Glycosyl hydrolase, partial (12%)
Length = 808
Score = 24.6 bits (52), Expect = 8.9
Identities = 11/38 (28%), Positives = 20/38 (51%)
Frame = +3
Query: 40 VIAGDTLVMRMTLTKLQKSFELAKMEGKAYVGGEVVCE 77
++ G+TL + ++ Q + G Y GGE++CE
Sbjct: 570 LVPGETLPINISFDVPQGVTPRVILHGWNYDGGEIICE 683
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.142 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,864,156
Number of Sequences: 63676
Number of extensions: 25283
Number of successful extensions: 179
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 179
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 179
length of query: 86
length of database: 12,639,632
effective HSP length: 62
effective length of query: 24
effective length of database: 8,691,720
effective search space: 208601280
effective search space used: 208601280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0314.19


