

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
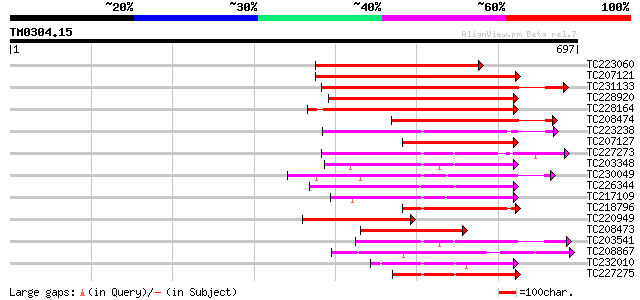
Query= TM0304.15
(697 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC223060 similar to PIR|T05137|T05137 protein kinase homolog F7H... 389 e-108
TC207121 similar to UP|O65833 (O65833) TCTR2 protein, partial (30%) 325 3e-89
TC231133 similar to UP|Q7X8K7 (Q7X8K7) CTR1-like kinase kinase k... 308 5e-84
TC228920 similar to UP|Q93XL9 (Q93XL9) CTR1-like protein kinase,... 295 6e-80
TC228164 similar to UP|CTR1_ARATH (Q05609) Serine/threonine-prot... 275 5e-74
TC208474 weakly similar to UP|O43283 (O43283) Leucine zipper bea... 256 2e-68
TC223238 similar to UP|Q94AB2 (Q94AB2) AT3g58640/F14P22_230, par... 228 7e-60
TC207127 weakly similar to UP|FUS3_YEAST (P16892) Mitogen-activa... 196 4e-50
TC227273 weakly similar to UP|KGS9_YEAST (P43637) Probable serin... 192 3e-49
TC203348 homologue to UP|Q9SAJ2 (Q9SAJ2) T8K14.1 protein, partia... 167 1e-41
TC230049 similar to UP|Q9MAV2 (Q9MAV2) F24O1.13, partial (47%) 159 3e-39
TC226344 homologue to UP|Q9AWA6 (Q9AWA6) Serine/threonine/tyrosi... 158 9e-39
TC217109 UP|Q39886 (Q39886) Protein kinase , complete 154 2e-37
TC218796 similar to UP|Q94AB2 (Q94AB2) AT3g58640/F14P22_230, par... 145 8e-35
TC220949 weakly similar to UP|Q13476 (Q13476) Kinase suppressor ... 144 2e-34
TC208473 similar to UP|Q9FGZ6 (Q9FGZ6) Protein kinase, partial (... 136 4e-32
TC203541 homologue to UP|Q9SAJ2 (Q9SAJ2) T8K14.1 protein, partia... 135 6e-32
TC208867 similar to UP|Q9LXT2 (Q9LXT2) Serine/threonine-specific... 129 4e-30
TC232010 homologue to UP|Q8L6Y9 (Q8L6Y9) Protein kinase ATN1-lik... 128 7e-30
TC227275 weakly similar to UP|RON_MOUSE (Q62190) Macrophage-stim... 128 7e-30
>TC223060 similar to PIR|T05137|T05137 protein kinase homolog F7H19.240 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(26%)
Length = 627
Score = 389 bits (1000), Expect = e-108
Identities = 186/207 (89%), Positives = 198/207 (94%)
Frame = +1
Query: 376 DNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKE 435
+++SV CEIHWE L LR+EIGQGS AVVYHGIWNGSDVA+KVYFGN YTEETLQDY+KE
Sbjct: 7 ESNSVSKCEIHWEHLQLREEIGQGSCAVVYHGIWNGSDVAVKVYFGNEYTEETLQDYRKE 186
Query: 436 IDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDI 495
IDIMKRLRHPNVLLFMGAVYS ERLAIVTELLPRGSLFK LH++NQTLDIRRRLRMALD+
Sbjct: 187 IDIMKRLRHPNVLLFMGAVYSQERLAIVTELLPRGSLFKNLHRNNQTLDIRRRLRMALDV 366
Query: 496 ARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWM 555
ARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKD TLLTTKSGRGTPQWM
Sbjct: 367 ARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDATLLTTKSGRGTPQWM 546
Query: 556 APEILRNEPSNEKSDVYSYGVVLWELM 582
APE+LRNEPSNEKSDVYS+GV+LWELM
Sbjct: 547 APEVLRNEPSNEKSDVYSFGVILWELM 627
>TC207121 similar to UP|O65833 (O65833) TCTR2 protein, partial (30%)
Length = 1452
Score = 325 bits (834), Expect = 3e-89
Identities = 151/252 (59%), Positives = 195/252 (76%)
Frame = +1
Query: 376 DNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKE 435
D+ V +CEI WEDL L + IG GSY VYH WNG++VA+K + ++ L ++K+E
Sbjct: 328 DDVDVGECEIPWEDLVLGERIGIGSYGEVYHADWNGTEVAVKKFLDQDFSGAALSEFKRE 507
Query: 436 IDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDI 495
+ IM+RLRHPN++LFMGAV L+I++E LPRGSL++ LH+SN +D +RR++MALD+
Sbjct: 508 VRIMRRLRHPNIVLFMGAVTRPPNLSIISEYLPRGSLYRILHRSNYQIDEKRRIKMALDV 687
Query: 496 ARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWM 555
ARGMN LH P IVHRDLKS NLLVDKNW VKV DFGLSRLK T L++KS GTP+WM
Sbjct: 688 ARGMNCLHTSTPTIVHRDLKSPNLLVDKNWNVKVCDFGLSRLKHNTFLSSKSTAGTPEWM 867
Query: 556 APEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDP 615
APE+LRNEPSNEK DVYS+GV+LWEL T +PW +N++QVVG VGF +RRLD+P+ +DP
Sbjct: 868 APEVLRNEPSNEKCDVYSFGVILWELATLRLPWSEMNTMQVVGAVGFQNRRLDIPKEVDP 1047
Query: 616 HVASIINDCWRR 627
VA II +CW++
Sbjct: 1048IVARIIWECWQQ 1083
>TC231133 similar to UP|Q7X8K7 (Q7X8K7) CTR1-like kinase kinase kinase,
partial (29%)
Length = 1061
Score = 308 bits (789), Expect = 5e-84
Identities = 153/304 (50%), Positives = 201/304 (65%)
Frame = +1
Query: 384 EIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLR 443
+I WE++ + + IG GSY VY G W+G++VA+K + + E L+++K E+ IMKRLR
Sbjct: 7 DIPWEEIAVGERIGLGSYGEVYRGEWHGTEVAVKKFLYQDISGELLEEFKSEVQIMKRLR 186
Query: 444 HPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLH 503
HPNV+LFMGAV L+IV+E LPRGSL++ +H+ N LD RRRLRMALD ARGMNYLH
Sbjct: 187 HPNVVLFMGAVTRPPNLSIVSEFLPRGSLYRLIHRPNNQLDERRRLRMALDAARGMNYLH 366
Query: 504 HRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNE 563
+ P IVHRDLKS NLLVDKNW VKV DFGLSR+K +T L+++S GT +WMAPE+LRNE
Sbjct: 367 NCTPVIVHRDLKSPNLLVDKNWVVKVCDFGLSRMKHSTFLSSRSTAGTAEWMAPEVLRNE 546
Query: 564 PSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIIND 623
S+EK DV+SYGV+LWEL T PW +N +QVVG VGF RRLD+P+ +DP +A II
Sbjct: 547 LSDEKCDVFSYGVILWELSTLQQPWGGMNPMQVVGAVGFQHRRLDIPDNVDPAIADIIRQ 726
Query: 624 CWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSFEELIQRMLFVLNRVTAE 683
CW+ +DP+ RP+F E++ + + +T
Sbjct: 727 CWQ-------------------------------TDPKLRPTFAEIMAALKPLQKPITVS 813
Query: 684 SVKR 687
V R
Sbjct: 814 QVHR 825
>TC228920 similar to UP|Q93XL9 (Q93XL9) CTR1-like protein kinase, partial
(31%)
Length = 1059
Score = 295 bits (754), Expect = 6e-80
Identities = 142/236 (60%), Positives = 178/236 (75%), Gaps = 2/236 (0%)
Frame = +1
Query: 392 LRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFM 451
LR++IG GS+ V+ WNGSDVA+K+ + E +++ +E+ IMKRLRHPN++LFM
Sbjct: 1 LREKIGSGSFGTVHRAEWNGSDVAVKILMEQDFHAERFKEFLREVAIMKRLRHPNIVLFM 180
Query: 452 GAVYSLERLAIVTELLPRGSLFKTLHKSN--QTLDIRRRLRMALDIARGMNYLHHRNPPI 509
GAV L+IVTE L RGSL++ LH+S + LD RRRL MA D+A+GMNYLH RNPPI
Sbjct: 181 GAVTQPPNLSIVTEYLSRGSLYRLLHRSGAKEVLDERRRLGMAYDVAKGMNYLHKRNPPI 360
Query: 510 VHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEPSNEKS 569
VHRDLKS NLLVDK +TVKV DFGLSRLK T L++KS GTP+WMAPE+L +EPSNEKS
Sbjct: 361 VHRDLKSPNLLVDKKYTVKVCDFGLSRLKANTFLSSKSAAGTPEWMAPEVLCDEPSNEKS 540
Query: 570 DVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCW 625
DVYS+GV+LWEL T PW NLN QVV VGF +RL++P ++P VA++I CW
Sbjct: 541 DVYSFGVILWELATLQQPWVNLNPAQVVAAVGFKRKRLEIPHDVNPQVAALIEACW 708
>TC228164 similar to UP|CTR1_ARATH (Q05609) Serine/threonine-protein kinase
CTR1 , partial (42%)
Length = 1571
Score = 275 bits (703), Expect = 5e-74
Identities = 137/261 (52%), Positives = 178/261 (67%), Gaps = 2/261 (0%)
Frame = +1
Query: 367 SHESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTE 426
SHE K D D I W +L L++ IG GS+ V W GSDVA+K+ G+
Sbjct: 448 SHEVDLDKEDLD------IPWSELILKENIGTGSFGTVLRADWRGSDVAVKILKVQGFDP 609
Query: 427 ETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSN--QTLD 484
+++ KE+ +MKRLRHPN++L MGAV +L+IVTE L RGSL++ LH N +L
Sbjct: 610 GRFEEFLKEVSLMKRLRHPNIVLLMGAVIQPPKLSIVTEYLSRGSLYELLHMPNVGSSLS 789
Query: 485 IRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLT 544
+RRL MA D+A GMNYLH PPIVHRDLKS NLLVD ++TVKV DFGLSR K T L+
Sbjct: 790 EKRRLSMAYDVASGMNYLHQMRPPIVHRDLKSPNLLVDDSYTVKVCDFGLSRTKANTFLS 969
Query: 545 TKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMD 604
+K+ GTP+WMAPE++R E S+EK DV+S+GV+LWEL+T PW LN QVV VGFM
Sbjct: 970 SKTAAGTPEWMAPEVIRGELSSEKCDVFSFGVILWELVTLQQPWRQLNPSQVVAAVGFMG 1149
Query: 605 RRLDLPEGLDPHVASIINDCW 625
+RL++P ++P VA++I CW
Sbjct: 1150KRLEIPGHVNPQVAALIELCW 1212
>TC208474 weakly similar to UP|O43283 (O43283) Leucine zipper bearing kinase,
partial (7%)
Length = 1206
Score = 256 bits (655), Expect = 2e-68
Identities = 123/204 (60%), Positives = 149/204 (72%)
Frame = +3
Query: 470 GSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKV 529
GSL + LH++ LD RRR+ MALDIARG+NYLHH NPPI+HRDLKSSNLLVDKNWTVKV
Sbjct: 201 GSLCRLLHRNTSKLDWRRRVHMALDIARGVNYLHHCNPPIIHRDLKSSNLLVDKNWTVKV 380
Query: 530 GDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWE 589
GDFGLSRLK T LTTK+GRGTPQWMAPE+LRNEPS+EKSDVY +GV+LWE++T+ IPW+
Sbjct: 381 GDFGLSRLKHETFLTTKTGRGTPQWMAPEVLRNEPSDEKSDVYGFGVILWEIVTEKIPWD 560
Query: 590 NLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTE 649
NLNS+QV+G VGFM++RL++P+ +DP ASII CW
Sbjct: 561 NLNSMQVIGAVGFMNQRLEIPKNVDPRWASIIESCWH----------------------- 671
Query: 650 SLLVLNMSSDPEQRPSFEELIQRM 673
SDP RP+F EL+ R+
Sbjct: 672 --------SDPACRPTFPELLDRL 719
>TC223238 similar to UP|Q94AB2 (Q94AB2) AT3g58640/F14P22_230, partial (33%)
Length = 876
Score = 228 bits (581), Expect = 7e-60
Identities = 121/292 (41%), Positives = 174/292 (59%), Gaps = 2/292 (0%)
Frame = +2
Query: 385 IHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRH 444
I + +L++ +G G + V GIWNG+DVAIKV+ T E ++D+ EI I+ RLRH
Sbjct: 20 IDFTELNVXTRVGIGFFGEVXRGIWNGTDVAIKVFLEQDLTAENMEDFCNEISILSRLRH 199
Query: 445 PNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQT--LDIRRRLRMALDIARGMNYL 502
PNV+LF+GA RL++VTE + GSLF +H S Q L RRRL+M DI RG+ ++
Sbjct: 200 PNVILFLGACTKPPRLSMVTEYMEMGSLFYLIHVSGQKKKLSWRRRLKMLRDICRGLMHI 379
Query: 503 HHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRN 562
H I+HRD+KS+N LVDK+W VK+ DFGLSR+ + + S GTP+WMAPE++RN
Sbjct: 380 HRMK--IIHRDVKSANCLVDKHWIVKICDFGLSRIITESPMRDSSSAGTPEWMAPELIRN 553
Query: 563 EPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIIN 622
EP +EK D++S GV++WEL T + PWE + +VV V RLD+PEG + +I+
Sbjct: 554 EPFSEKCDIFSLGVIMWELCTLNRPWEGVPPERVVYTVANEGARLDIPEG---PLGRLIS 724
Query: 623 DCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSFEELIQRML 674
+CW ++P +RPS EE++ R++
Sbjct: 725 ECW--------------------------------AEPHERPSCEEILSRLV 784
>TC207127 weakly similar to UP|FUS3_YEAST (P16892) Mitogen-activated protein
kinase FUS3 (MAP kinase FUS3) , partial (7%)
Length = 972
Score = 196 bits (497), Expect = 4e-50
Identities = 92/143 (64%), Positives = 110/143 (76%)
Frame = +3
Query: 483 LDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTL 542
LD RRRLRMA D+A+G+NYLH PPIVH DLK+ NLLVD+NWTVKV DFGLSR K T
Sbjct: 3 LDPRRRLRMA*DVAKGINYLHCLKPPIVHWDLKTPNLLVDRNWTVKVCDFGLSRFKANTF 182
Query: 543 LTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGF 602
L++KS GTP+WMAPE LR EPSNEKSDVYS+GV+LWEL+T PW L+ QVVG V F
Sbjct: 183 LSSKSVAGTPEWMAPEFLRGEPSNEKSDVYSFGVILWELVTLQQPWNGLSHAQVVGAVAF 362
Query: 603 MDRRLDLPEGLDPHVASIINDCW 625
+RRL +P + P +AS++ CW
Sbjct: 363 QNRRLAIPPNISPALASLMESCW 431
>TC227273 weakly similar to UP|KGS9_YEAST (P43637) Probable
serine/threonine-protein kinase YGL179C , partial (8%)
Length = 2373
Score = 192 bits (489), Expect = 3e-49
Identities = 115/314 (36%), Positives = 180/314 (56%), Gaps = 9/314 (2%)
Frame = +3
Query: 384 EIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLR 443
EI L +++G GS+ +Y G + DVAIKV + + L+++ +E+ IM+++R
Sbjct: 993 EIDTNQLKYENKVGSGSFGDLYRGTYCSQDVAIKVLKPERISTDMLREFAQEVYIMRKIR 1172
Query: 444 HPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGMNYLH 503
H NV+ F+GA L IVTE + RGSL+ LHK + L++A+D+++GMNYLH
Sbjct: 1173 HKNVVQFIGACTRPPNLCIVTEFMSRGSLYDFLHKQRGVFKLPSLLKVAIDVSKGMNYLH 1352
Query: 504 HRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNE 563
N I+HRDLK++NLL+D+N VKV DFG++R++ + + T + GT +WMAPE++ ++
Sbjct: 1353 QNN--IIHRDLKTANLLMDENEVVKVADFGVARVQTQSGVMT-AETGTYRWMAPEVIEHK 1523
Query: 564 PSNEKSDVYSYGVVLWELMTQSIPWENLNSLQ-VVGVVGFMDRRLDLPEGLDPHVASIIN 622
P ++K+DV+S+G+ LWEL+T +P+ L LQ VGVV +GL P +I
Sbjct: 1524 PYDQKADVFSFGIALWELLTGELPYSCLTPLQAAVGVV---------QKGLRP---TIPK 1667
Query: 623 DCWRRCFYMYSSLQLYYGRNHLL--------PQTESLLVLNMSSDPEQRPSFEELIQRML 674
+ R ++ QL + R L P+ LL DP QRP+F E+I+ +
Sbjct: 1668 NTHPRLS*LHYKQQLAWCRRDQLSTIPKNTHPRLSELLQRCWQQDPTQRPNFSEVIEILQ 1847
Query: 675 FVLNRVTAESVKRS 688
+ V K S
Sbjct: 1848 QIAKEVNDHKDKAS 1889
>TC203348 homologue to UP|Q9SAJ2 (Q9SAJ2) T8K14.1 protein, partial (23%)
Length = 951
Score = 167 bits (423), Expect = 1e-41
Identities = 103/252 (40%), Positives = 146/252 (57%), Gaps = 14/252 (5%)
Frame = +3
Query: 388 EDLHLRDEIGQGSYAVVYHGIWNGSDVAIK-----VYFGNGYTEETLQ-DYKKEIDIMKR 441
EDL E+G G++ VYHG W GSDVAIK + G +E L ++ +E DI+ +
Sbjct: 117 EDLEELRELGSGTFGTVYHGKWRGSDVAIKRIKKSCFAGRSSEQERLTIEFWREADILSK 296
Query: 442 LRHPNVLLFMGAVYSLE--RLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRMALDIARGM 499
L HPNV+ F G V LA V E + GSL L + ++ LD R+RL +A+D A GM
Sbjct: 297 LHHPNVVAFYGVVQDGPGATLATVAEYMVDGSLRNVLLRKDRYLDRRKRLIIAMDAAFGM 476
Query: 500 NYLHHRNPPIVHRDLKSSNLLVDKNWTV----KVGDFGLSRLKDTTLLTTKSGRGTPQWM 555
YLH +N IVH DLK NLLV+ + KVGDFGLS++K TL++ RGT WM
Sbjct: 477 EYLHSKN--IVHFDLKCDNLLVNLKDPIRPICKVGDFGLSKIKRNTLVSG-GVRGTLPWM 647
Query: 556 APEILRNEPS--NEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGL 613
APE+L + +EK DV+S+G+VLWE++T P+ N++ ++G + R +P
Sbjct: 648 APELLNGSSNKVSEKVDVFSFGIVLWEILTGDEPYANMHYGAIIGGIVNNTLRPTIPSYC 827
Query: 614 DPHVASIINDCW 625
D +++ CW
Sbjct: 828 DLEWKTLMEQCW 863
>TC230049 similar to UP|Q9MAV2 (Q9MAV2) F24O1.13, partial (47%)
Length = 1216
Score = 159 bits (403), Expect = 3e-39
Identities = 106/337 (31%), Positives = 163/337 (47%), Gaps = 8/337 (2%)
Frame = +2
Query: 342 HMAIEDEVQKQQGRLQLPSSRESVGSHESSSSKG---DNDSVVDCEIHWEDLHLRDEIGQ 398
H I + K RL L +V + S G + + L + +
Sbjct: 11 HKQISNSGNKLGRRLSLGEYNRAVSWSKYLVSPGAEIKGEGEEEWSADMSQLLIGSKFAS 190
Query: 399 GSYAVVYHGIWNGSDVAIKVYFGNGYTEETL----QDYKKEIDIMKRLRHPNVLLFMGAV 454
G ++ +Y G++ DVAIK+ E+ + + E+ ++ RL HPN++ F+ A
Sbjct: 191 GRHSRIYRGVYKQKDVAIKLISQPEEDEDLAAFLEKQFTSEVSLLLRLGHPNIITFIAAC 370
Query: 455 YSLERLAIVTELLPRGSLFKTLHKSNQT-LDIRRRLRMALDIARGMNYLHHRNPPIVHRD 513
I+TE L GSL K LH L ++ L++ALDIARGM YLH + I+HRD
Sbjct: 371 KKPPVFCIITEYLAGGSLGKFLHHQQPNILPLKLVLKLALDIARGMKYLHSQG--ILHRD 544
Query: 514 LKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYS 573
LKS NLL+ ++ VKV DFG+S L ++ + K GT +WMAPE+++ + +K DVYS
Sbjct: 545 LKSENLLLGEDMCVKVADFGISCL-ESQCGSAKGFTGTYRWMAPEMIKEKHHTKKVDVYS 721
Query: 574 YGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYS 633
+G+VLWEL+T P++N+ Q V + R LP + +IN CW
Sbjct: 722 FGIVLWELLTGKTPFDNMTPEQAAYAVSHKNARPPLPSKCPWAFSDLINRCW-------- 877
Query: 634 SLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSFEELI 670
SS+P++RP F+E++
Sbjct: 878 -----------------------SSNPDKRPHFDEIV 919
>TC226344 homologue to UP|Q9AWA6 (Q9AWA6) Serine/threonine/tyrosine kinase,
partial (98%)
Length = 1818
Score = 158 bits (399), Expect = 9e-39
Identities = 91/262 (34%), Positives = 155/262 (58%), Gaps = 5/262 (1%)
Frame = +2
Query: 369 ESSSSKGDNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKVYFG--NGYTE 426
++SS D+ + I L++ + QG++ +Y G +NG DVAIK+ N +
Sbjct: 596 DNSSPTEGLDNFDEWTIDLRKLNMGEPFAQGAFGKLYRGTYNGEDVAIKILERPENDPAK 775
Query: 427 ETL--QDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHK-SNQTL 483
L Q +++E+ ++ L+HPN++ F+GA IVTE GS+ + L K N+++
Sbjct: 776 AQLMEQQFQQEVTMLATLKHPNIVRFIGACRKPMVWCIVTEYAKGGSVRQFLMKRQNRSV 955
Query: 484 DIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLL 543
++ ++ ALD+ARGM Y+H ++HRDLKS NLL+ + ++K+ DFG++R++ T
Sbjct: 956 PLKLAVKQALDVARGMAYVH--GLLLIHRDLKSDNLLIFGDKSIKIADFGVARIEVQTEG 1129
Query: 544 TTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFM 603
T GT +WMAPE++++ P +K DVYS+G+VLWEL+T +P++N+ ++Q V
Sbjct: 1130MTPE-TGTYRWMAPEMIQHRPYTQKVDVYSFGIVLWELITGMLPFQNMTAVQAAFAVVNK 1306
Query: 604 DRRLDLPEGLDPHVASIINDCW 625
+ R +P P + I+ CW
Sbjct: 1307NVRPIIPNDCLPVLRDIMTRCW 1372
>TC217109 UP|Q39886 (Q39886) Protein kinase , complete
Length = 1940
Score = 154 bits (388), Expect = 2e-37
Identities = 80/238 (33%), Positives = 140/238 (58%), Gaps = 7/238 (2%)
Frame = +1
Query: 395 EIGQGSYAVVYHGIWNGSDVAIKVYF-----GNGYTEETLQ-DYKKEIDIMKRLRHPNVL 448
+ G+++ +YHG++ VA+K+ NG L+ + +E+ ++ RL H NV+
Sbjct: 814 KFAHGAHSRLYHGVYKEEAVAVKIIMVPEDDENGALASRLEKQFIREVTLLSRLHHQNVI 993
Query: 449 LFMGAVYSLERLAIVTELLPRGSLFKTLHK-SNQTLDIRRRLRMALDIARGMNYLHHRNP 507
F A I+TE L GSL LHK +QT+ +++ + ALDIARGM Y+H +
Sbjct: 994 KFSAACRKPPVYCIITEYLAEGSLRAYLHKLEHQTVSLQKLIAFALDIARGMEYIHSQG- 1170
Query: 508 PIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEPSNE 567
++HRDLK N+L++++ +K+ DFG++ ++ + GT +WMAPE+++ + +
Sbjct: 1171 -VIHRDLKPENVLINEDNHLKIADFGIA-CEEASCDLLADDPGTYRWMAPEMIKRKSYGK 1344
Query: 568 KSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCW 625
K DVYS+G++LWE++T +IP+E++N +Q V + R +P P + ++I CW
Sbjct: 1345 KVDVYSFGLILWEMLTGTIPYEDMNPIQAAFAVVNKNSRPIIPSNCPPAMRALIEQCW 1518
>TC218796 similar to UP|Q94AB2 (Q94AB2) AT3g58640/F14P22_230, partial (20%)
Length = 701
Score = 145 bits (365), Expect = 8e-35
Identities = 70/146 (47%), Positives = 98/146 (66%)
Frame = +2
Query: 483 LDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTL 542
L+ RRRLRM DI +G+ +H +VHRDLKS+N LV+K+WTVK+ DFGLSR+ +
Sbjct: 8 LNWRRRLRMLRDICKGLMCIHRMK--VVHRDLKSANCLVNKHWTVKICDFGLSRIMTESP 181
Query: 543 LTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGF 602
+ S GTP+WMAPE++RNEP EK D++S GV++WEL T + PWE + +VV V
Sbjct: 182 MRDSSSAGTPEWMAPELIRNEPFTEKCDIFSLGVIMWELCTLNRPWEGVPPERVVYSVAH 361
Query: 603 MDRRLDLPEGLDPHVASIINDCWRRC 628
RL++PEG + +I++CW C
Sbjct: 362 EGSRLEIPEG---PLGRLISECWAEC 430
>TC220949 weakly similar to UP|Q13476 (Q13476) Kinase suppressor of ras-1
(Fragment), partial (6%)
Length = 637
Score = 144 bits (362), Expect = 2e-34
Identities = 72/140 (51%), Positives = 99/140 (70%), Gaps = 2/140 (1%)
Frame = +2
Query: 361 SRESVGSHESSSSKG--DNDSVVDCEIHWEDLHLRDEIGQGSYAVVYHGIWNGSDVAIKV 418
S SV S++S+ S D+ V + +I WE++ L + IG GSY VYHG W+G+++A+K
Sbjct: 218 SDRSVVSNDSTKSDSALDDHEVAEVDIPWEEITLGERIGLGSYGEVYHGEWHGTEIAVKR 397
Query: 419 YFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHK 478
+ + E+L+++K E+ IMKRLRHPNV+LFMGAV L+IVTE LPRGSL++ LH+
Sbjct: 398 FLDQDISGESLEEFKTEVRIMKRLRHPNVVLFMGAVTRPPNLSIVTEFLPRGSLYRLLHR 577
Query: 479 SNQTLDIRRRLRMALDIARG 498
N LD RRRL+MALD ARG
Sbjct: 578 PNSQLDERRRLKMALDTARG 637
>TC208473 similar to UP|Q9FGZ6 (Q9FGZ6) Protein kinase, partial (11%)
Length = 397
Score = 136 bits (342), Expect = 4e-32
Identities = 65/132 (49%), Positives = 95/132 (71%)
Frame = +2
Query: 432 YKKEIDIMKRLRHPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSNQTLDIRRRLRM 491
+K+E+ +MKRLRHPN++LFMGAV S + L IVT+ LP LF L ++ ++ R++L +
Sbjct: 2 FKQEVSVMKRLRHPNIILFMGAVTSPQHLCIVTKFLPHKILFHLLQRNTSKIN*RQQLHI 181
Query: 492 ALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGRGT 551
L + + +NYLHH +PPI+H+DL SSN+LV+KN T++ +F LS LK T LTT++ + T
Sbjct: 182 TLYMTQSINYLHHYDPPIIHQDLNSSNILVNKN*TIRSHNFNLSHLKHKTYLTTETEKET 361
Query: 552 PQWMAPEILRNE 563
PQ M P+ L NE
Sbjct: 362 PQ*MRPKFLPNE 397
>TC203541 homologue to UP|Q9SAJ2 (Q9SAJ2) T8K14.1 protein, partial (19%)
Length = 850
Score = 135 bits (340), Expect = 6e-32
Identities = 89/273 (32%), Positives = 140/273 (50%), Gaps = 8/273 (2%)
Frame = +3
Query: 426 EETLQDYKKEIDIMKRLRHPNVLLFMGAVYSLE--RLAIVTELLPRGSLFKTLHKSNQTL 483
E ++ +E +I+ +L HPNV+ F G V +A V E + GSL L + ++ L
Sbjct: 24 ERLTVEFWREAEILSKLHHPNVVAFYGVVQDGPGGTMATVAEYMVDGSLRHVLLRKDRYL 203
Query: 484 DIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTV----KVGDFGLSRLKD 539
D R+RL +A+D A GM YLH +N IVH DLK NLLV+ + KVGDFGLS++K
Sbjct: 204 DRRKRLIIAMDAAFGMEYLHSKN--IVHFDLKCDNLLVNLKDPMRPICKVGDFGLSKIKR 377
Query: 540 TTLLTTKSGRGTPQWMAPEILRNEPS--NEKSDVYSYGVVLWELMTQSIPWENLNSLQVV 597
TL++ RGT WMAPE+L + +EK DV+S+G+VLWE++T P+ N++ ++
Sbjct: 378 NTLVSG-GVRGTLPWMAPELLNGSSNKVSEKVDVFSFGIVLWEILTGEEPYANMHYGAII 554
Query: 598 GVVGFMDRRLDLPEGLDPHVASIINDCWRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMS 657
G + R +P+ D +++ CW +
Sbjct: 555 GGIVNNTLRPTIPDHCDSEWRTLMEQCW-------------------------------A 641
Query: 658 SDPEQRPSFEELIQRMLFVLNRVTAESVKRSSE 690
+P RPSF E+ R+ + + +++S+
Sbjct: 642 PNPAARPSFTEIASRLRIMTAAASQTKTQKASK 740
>TC208867 similar to UP|Q9LXT2 (Q9LXT2) Serine/threonine-specific protein
kinase-like protein, partial (72%)
Length = 1118
Score = 129 bits (324), Expect = 4e-30
Identities = 97/310 (31%), Positives = 155/310 (49%), Gaps = 11/310 (3%)
Frame = +1
Query: 396 IGQGSYAVVYHGIWN-GSDVAIKVYFGNGYTEETLQDYKKEIDIMKRLRHPNVLLFMGAV 454
IG G + +VY G+ N G VAIK G E +++K E++++ RL P +L +G
Sbjct: 16 IGHGGFGLVYRGVLNDGRKVAIKFMDQAGKQGE--EEFKVEVELLSRLHSPYLLALLGYC 189
Query: 455 YSLERLAIVTELLPRGSLFKTLHKSNQT------LDIRRRLRMALDIARGMNYLH-HRNP 507
+V E + G L + L+ + + LD RLR+AL+ A+G+ YLH H +P
Sbjct: 190 SDSNHKLLVYEFMANGGLQEHLYPVSNSIITPVKLDWETRLRIALEAAKGLEYLHEHVSP 369
Query: 508 PIVHRDLKSSNLLVDKNWTVKVGDFGLSRLKDTTLLTTKSGR--GTPQWMAPEILRNEPS 565
P++HRD KSSN+L+DK + KV DFGL++L S R GT ++APE
Sbjct: 370 PVIHRDFKSSNILLDKKFHAKVSDFGLAKLGPDRAGGHVSTRVLGTQGYVAPEYALTGHL 549
Query: 566 NEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEG-LDPHVASIINDC 624
KSDVYSYGVVL EL+T +P +D + EG L ++ D
Sbjct: 550 TTKSDVYSYGVVLLELLTGRVP---------------VDMKRPPGEGVLVSWALPLLTDR 684
Query: 625 WRRCFYMYSSLQLYYGRNHLLPQTESLLVLNMSSDPEQRPSFEELIQRMLFVLNRVTAES 684
+ M SL+ Y ++ Q ++ + + + + RP +++Q ++ ++ + S
Sbjct: 685 EKVVKIMDPSLEGQYSMKEVV-QVAAIAAMCVQPEADYRPLMADVVQSLVPLVKTQRSPS 861
Query: 685 VKRSSETCNA 694
SS + N+
Sbjct: 862 KVGSSSSFNS 891
>TC232010 homologue to UP|Q8L6Y9 (Q8L6Y9) Protein kinase ATN1-like protein,
partial (62%)
Length = 674
Score = 128 bits (322), Expect = 7e-30
Identities = 79/192 (41%), Positives = 112/192 (58%), Gaps = 10/192 (5%)
Frame = +2
Query: 444 HPNVLLFMGAVYSLERLAIVTELLPRGSLFKTLHKSN-QTLDIRRRLRMALDIARGMNYL 502
H N++ F+GA + IVTE+LP SL K L + LD ++ ALDIAR M++L
Sbjct: 71 HENLVKFIGACKD-PLMVIVTEMLPGLSLRKYLTTIRPKQLDPYVAIKFALDIARAMDWL 247
Query: 503 HHRNPPIVHRDLKSSNLLVDKNW-TVKVGDFGLSRLKDTTLLTTKSGRGTPQWMAPEIL- 560
H I+HRDLK NLL+ +N +VK+ DFGL+R + T + T GT +WMAPE+
Sbjct: 248 HANG--IIHRDLKPDNLLLTENQKSVKLADFGLAREESVTEMMTAE-TGTYRWMAPELYS 418
Query: 561 -------RNEPSNEKSDVYSYGVVLWELMTQSIPWENLNSLQVVGVVGFMDRRLDLPEGL 613
+ N K DVYS+G+VLWEL+T +P+E +++LQ F R +LP+ +
Sbjct: 419 TVTLRQGEKKHYNNKVDVYSFGIVLWELLTNRMPFEGMSNLQAAYAAAFKQERPNLPDDI 598
Query: 614 DPHVASIINDCW 625
P +A II CW
Sbjct: 599 SPDLAFIIQSCW 634
>TC227275 weakly similar to UP|RON_MOUSE (Q62190) Macrophage-stimulating
protein receptor precursor (MSP receptor) (p185-Ron)
(Stem cell-derived tyrosine kinase) , partial (4%)
Length = 737
Score = 128 bits (322), Expect = 7e-30
Identities = 63/157 (40%), Positives = 100/157 (63%)
Frame = +2
Query: 471 SLFKTLHKSNQTLDIRRRLRMALDIARGMNYLHHRNPPIVHRDLKSSNLLVDKNWTVKVG 530
S++ LHK ++A+D+++GMNYLH N I+HRDLK++NLL+D+N TVKV
Sbjct: 5 SVYDYLHKQKGFXXFXTXXKVAIDVSKGMNYLHQHN--IIHRDLKAANLLMDENCTVKVA 178
Query: 531 DFGLSRLKDTTLLTTKSGRGTPQWMAPEILRNEPSNEKSDVYSYGVVLWELMTQSIPWEN 590
DFG++R+K + + T GT +WMAPE++ ++P + K+DV+S+G+VLWEL+T +P+E
Sbjct: 179 DFGVARVKAQSGVMTAE-TGTYRWMAPEVIEHKPYDHKADVFSFGIVLWELLTGKLPYEY 355
Query: 591 LNSLQVVGVVGFMDRRLDLPEGLDPHVASIINDCWRR 627
L LQ V R +P+ P ++ W++
Sbjct: 356 LTPLQAAIGVVQKGLRPTIPKNTHPKFVELLERSWQQ 466
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.132 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,961,260
Number of Sequences: 63676
Number of extensions: 431358
Number of successful extensions: 3937
Number of sequences better than 10.0: 819
Number of HSP's better than 10.0 without gapping: 3240
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3307
length of query: 697
length of database: 12,639,632
effective HSP length: 104
effective length of query: 593
effective length of database: 6,017,328
effective search space: 3568275504
effective search space used: 3568275504
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0304.15


