

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0302a.8
(578 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
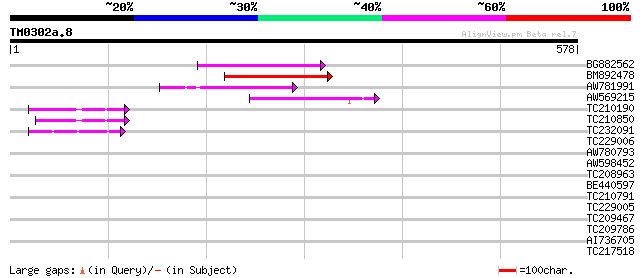
Score E
Sequences producing significant alignments: (bits) Value
BG882562 110 2e-24
BM892478 108 9e-24
AW781991 96 3e-20
AW569215 weakly similar to GP|4914326|gb| F14N23.12 {Arabidopsis... 80 2e-15
TC210190 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, part... 51 1e-06
TC210850 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, part... 49 5e-06
TC232091 45 9e-05
TC229006 42 0.001
AW780793 39 0.005
AW598452 weakly similar to GP|15982769|gb| At2g27110/T20P8.16 {A... 36 0.041
TC208963 35 0.070
BE440597 31 1.3
TC210791 31 1.3
TC229005 29 6.6
TC209467 similar to UP|XT32_ARATH (Q9SJL9) Probable xyloglucan e... 29 6.6
TC209786 similar to GB|AAS49116.1|44917583|BT011753 At2g28480 {A... 29 6.6
AI736705 similar to GP|11994736|db far-red impaired response pro... 28 8.6
TC217518 28 8.6
>BG882562
Length = 416
Score = 110 bits (274), Expect = 2e-24
Identities = 55/131 (41%), Positives = 75/131 (56%)
Frame = +3
Query: 192 KTDAEFAISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDATY 251
K D E +Y + +P + NDDG L +FW D S+ Y FGDVVAFD+T
Sbjct: 24 KGDPELISNYFCRIQLMNPNFFYVMDLNDDGQLRNVFWIDSRSRAAYSYFGDVVAFDSTC 203
Query: 252 GKNRYKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKAPLFVITDG 311
N Y+ P V F GVNHH KS + L+++E ETY+WL +L M G+ P +IT+
Sbjct: 204 LSNNYEIPLVAFVGVNHHGKSVLLGCGLLADETFETYIWLFRAWLTCMTGRPPQTIITNX 383
Query: 312 DKAMKAAIKQV 322
KAM++AI +V
Sbjct: 384 CKAMQSAIAEV 416
>BM892478
Length = 429
Score = 108 bits (269), Expect = 9e-24
Identities = 49/110 (44%), Positives = 69/110 (62%)
Frame = +3
Query: 220 DDGNLDRLFWSDGISQLNYQVFGDVVAFDATYGKNRYKCPFVVFYGVNHHNKSTIFSTAL 279
D+GN +FW+DG S+ + FGDV+ D +Y K Y PF F GVNHH + + AL
Sbjct: 93 DNGNCMNIFWADGRSRYSCSHFGDVLVLDTSYRKTVYLVPFATFVGVNHHKQPVLLGCAL 272
Query: 280 VSNEKIETYVWLLERFLEAMKGKAPLFVITDGDKAMKAAIKQVFPNAHHR 329
+++E E++ WL + +L AM G+ PL VI D D A++ AI +VFP HHR
Sbjct: 273 IADESEESFTWLFQTWLRAMSGRLPLTVIADQDIAIQRAIAKVFPVTHHR 422
>AW781991
Length = 419
Score = 96.3 bits (238), Expect = 3e-20
Identities = 46/141 (32%), Positives = 78/141 (54%)
Frame = +3
Query: 153 QIHNTFASQTGGYQKVRFSKGNMYNVQGRQRRMKKKEPEKTDAEFAISYIRGLGSKDPLL 212
+I + + GG KV F++ + N R R++ E D + + Y+R + +++P
Sbjct: 3 EIMSALIKEYGGISKVGFTEVDCRNYM-RNNRLRSLEG---DIQLVLDYLRQMHAENPNF 170
Query: 213 YCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDATYGKNRYKCPFVVFYGVNHHNKS 272
+ ++D ++ +FW+D +++NY FGD V FD TY NRY+ PF F GVNHH +
Sbjct: 171 FYAVQGDEDQSITNVFWADPKARMNYTFFGDTVTFDTTYRSNRYRLPFAFFTGVNHHGQP 350
Query: 273 TIFSTALVSNEKIETYVWLLE 293
+F A + NE ++VWL +
Sbjct: 351 VLFGCAFLINESEASFVWLFK 413
>AW569215 weakly similar to GP|4914326|gb| F14N23.12 {Arabidopsis thaliana},
partial (19%)
Length = 425
Score = 80.1 bits (196), Expect = 2e-15
Identities = 45/139 (32%), Positives = 68/139 (48%), Gaps = 6/139 (4%)
Frame = +1
Query: 245 VAFDATYGKNRYKCPFVVFYGVNHHNKSTIFSTALVSNEKIETYVWLLERFLEAMKGKAP 304
V FD TY Y ++ GV+++ + FS AL+ +E I+++ W L+ FL MKGKAP
Sbjct: 10 VVFDTTYRVEAYDMLLGIWLGVDNNGMTCFFSCALLRDENIQSFSWALKAFLGFMKGKAP 189
Query: 305 LFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNARKHVHI------RAFNTELRKCMFM 358
++TD + +K AI P H C WHIL + + E + +
Sbjct: 190 QTILTDHNMWLKEAIAVELPQTKHAFCIWHILSKFSDWFSLLLGSQYDEWKAEFHRLYNL 369
Query: 359 ELDVSEFEEHWAATVAKFG 377
E V +FEE W V ++G
Sbjct: 370 E-QVEDFEEGWRQMVDQYG 423
>TC210190 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, partial (65%)
Length = 866
Score = 51.2 bits (121), Expect = 1e-06
Identities = 35/106 (33%), Positives = 48/106 (45%), Gaps = 3/106 (2%)
Frame = +1
Query: 20 DFLDSEVAYLFYHWYG*VNGFAVR-KYIKIHNKEGLVIQRNFVCHKEGLREDRGLTMDKR 78
+F A+ FY+ Y GF VR + ++G I R VC+KEG R DKR
Sbjct: 4 EFGSEAAAHAFYNAYATEVGFIVRVSKLSRSRRDGTAIGRTLVCNKEGFR-----MADKR 168
Query: 79 QR--ESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHELEDDK 122
++ R + R GC A V ++G+W F EH H L K
Sbjct: 169 EKIVRQRAETRVGCRAMIMVR-KLSSGKWVVAKFVKEHTHPLTPGK 303
>TC210850 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, partial (63%)
Length = 1326
Score = 49.3 bits (116), Expect = 5e-06
Identities = 34/99 (34%), Positives = 46/99 (46%), Gaps = 3/99 (3%)
Frame = +1
Query: 27 AYLFYHWYG*VNGFAVR-KYIKIHNKEGLVIQRNFVCHKEGLREDRGLTMDKRQR--ESR 83
A+ FY+ Y GF VR + ++G I R VC+KEG R DKR++ R
Sbjct: 7 AHAFYNAYATDVGFIVRVSKLSRSRRDGTAIGRTLVCNKEGFR-----MADKREKIVRQR 171
Query: 84 TDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHELEDDK 122
+ R GC A V ++G+W F EH H L K
Sbjct: 172 AETRVGCRAMIMVR-KLSSGKWVIAKFVKEHTHPLTPGK 285
>TC232091
Length = 530
Score = 45.1 bits (105), Expect = 9e-05
Identities = 34/101 (33%), Positives = 44/101 (42%), Gaps = 2/101 (1%)
Frame = +1
Query: 20 DFLDSEVAYLFYHWYG*VNGFAVRKYIKIHNKE--GLVIQRNFVCHKEGLREDRGLTMDK 77
+F E A FY Y GF VR + H E +I R F C+K+G R K
Sbjct: 151 EFQSEEAAKNFYEEYARREGFVVR-LDRCHRSEVDKQIISRRFSCNKQGFHV-RVRNKTK 324
Query: 78 RQRESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHEL 118
+ R R GC A V ++ T G+W F EH+H L
Sbjct: 325 PVHKPRASIREGCEAMMYVKVN-TCGKWVVTKFVKEHSHLL 444
>TC229006
Length = 991
Score = 41.6 bits (96), Expect = 0.001
Identities = 30/101 (29%), Positives = 47/101 (45%), Gaps = 2/101 (1%)
Frame = +2
Query: 20 DFLDSEVAYLFYHWYG*VNGFAVRKY-IKIHNKEGLVIQRNFVCHKEGLRED-RGLTMDK 77
+F + A +FY Y GF +R + ++G ++ R C+KEG RG
Sbjct: 116 EFESEDAAKIFYDEYARRLGFVMRVMSCRRPERDGRILARRLGCNKEGYCVSIRGKFASV 295
Query: 78 RQRESRTDFRCGCTAKFRVHIDKTNGRWYAKLFSDEHNHEL 118
R+ + T R GC A + +K+ G+W F +HNH L
Sbjct: 296 RKPRAST--REGCKAMIHIKYNKS-GKWVITKFVKDHNHPL 409
>AW780793
Length = 419
Score = 39.3 bits (90), Expect = 0.005
Identities = 32/95 (33%), Positives = 42/95 (43%), Gaps = 4/95 (4%)
Frame = +3
Query: 20 DFLDSEVAYLFYHWYG*VNGFAVRKYIKIHNKEGLV----IQRNFVCHKEGLREDRGLTM 75
+F E A FY YG GF VR + HN+ V I ++FVC KEG R + L
Sbjct: 147 EFNSREEAREFYIAYGRRIGFTVRIH---HNRRSRVNNQVIGQDFVCSKEGFRAKKYLHR 317
Query: 76 DKRQRESRTDFRCGCTAKFRVHIDKTNGRWYAKLF 110
R R GC A R+ + + G+W F
Sbjct: 318 KDRVLPPPPATREGCQAMIRLAL-RDRGKWVVTKF 419
>AW598452 weakly similar to GP|15982769|gb| At2g27110/T20P8.16 {Arabidopsis
thaliana}, partial (7%)
Length = 256
Score = 36.2 bits (82), Expect = 0.041
Identities = 18/78 (23%), Positives = 38/78 (48%)
Frame = +2
Query: 194 DAEFAISYIRGLGSKDPLLYCRHVANDDGNLDRLFWSDGISQLNYQVFGDVVAFDATYGK 253
DA + Y + + +++ + ++D + + + S+ +Y+ GD + DA YGK
Sbjct: 17 DAHNLLEYCKRMQAENHGCFYAIQLDEDDRVSNVL*AGTRSRTSYKHMGDAESLDAAYGK 196
Query: 254 NRYKCPFVVFYGVNHHNK 271
+RY + + G N H +
Sbjct: 197 HRYAVSYAPYTGRNCHGR 250
>TC208963
Length = 1163
Score = 35.4 bits (80), Expect = 0.070
Identities = 14/37 (37%), Positives = 25/37 (66%)
Frame = +1
Query: 304 PLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNAR 340
P ++T GD A+ +A++ VFP++ + LC +HI N +
Sbjct: 22 PQVIVTVGDIALMSAVQVVFPSSSNLLCRFHINQNVK 132
>BE440597
Length = 443
Score = 31.2 bits (69), Expect = 1.3
Identities = 21/74 (28%), Positives = 35/74 (46%), Gaps = 9/74 (12%)
Frame = +2
Query: 109 LFSD-EHNHELEDDKMCGMIAAHRKMNESDIMHMNNMSRSGIST--------PQIHNTFA 159
LFS + NH +C +A++ +M++ D ++S S T P+ HN F
Sbjct: 74 LFSQIKKNHFQFLPLLCASMASNHEMDQDDDGGATDLSTSDAETTTTNLNGVPKQHNAFE 253
Query: 160 SQTGGYQKVRFSKG 173
+T G+Q + KG
Sbjct: 254 EETTGFQVLPLKKG 295
>TC210791
Length = 456
Score = 31.2 bits (69), Expect = 1.3
Identities = 14/37 (37%), Positives = 23/37 (61%)
Frame = +1
Query: 304 PLFVITDGDKAMKAAIKQVFPNAHHRLCAWHILDNAR 340
P +IT D A+ A++ VFP++ + LC +HI N +
Sbjct: 31 PPVIITVRDIALMDAVQVVFPSSSNLLCRFHISKNVK 141
>TC229005
Length = 679
Score = 28.9 bits (63), Expect = 6.6
Identities = 13/32 (40%), Positives = 17/32 (52%)
Frame = +1
Query: 87 RCGCTAKFRVHIDKTNGRWYAKLFSDEHNHEL 118
R GC A + DK+ G+W F +HNH L
Sbjct: 10 REGCKAMIHIKYDKS-GKWVIAKFVKDHNHPL 102
>TC209467 similar to UP|XT32_ARATH (Q9SJL9) Probable xyloglucan
endotransglucosylase/hydrolase protein 32 precursor
(At-XTH32) (XTH-32) , partial (93%)
Length = 1223
Score = 28.9 bits (63), Expect = 6.6
Identities = 10/29 (34%), Positives = 20/29 (68%)
Frame = +2
Query: 257 KCPFVVFYGVNHHNKSTIFSTALVSNEKI 285
+C ++V YG++HH + + ST L++N +
Sbjct: 671 QCGYMVQYGMHHHGQLRMESTKLITNTNL 757
>TC209786 similar to GB|AAS49116.1|44917583|BT011753 At2g28480 {Arabidopsis
thaliana;} , partial (64%)
Length = 1160
Score = 28.9 bits (63), Expect = 6.6
Identities = 24/68 (35%), Positives = 34/68 (49%), Gaps = 10/68 (14%)
Frame = +2
Query: 186 KKKEPEKTDAEFAISYIR---GLGSKDPLLYCRHVA--NDDGNLDRLFWSDGIS-----Q 235
KKK EK+ E ++ +R + K+ LYCRHVA D N + L D S +
Sbjct: 785 KKKALEKSKFEQSLESVRRFVAIAEKELELYCRHVALYGDPNNRNLLSMLDVPSGNSKEK 964
Query: 236 LNYQVFGD 243
N++V GD
Sbjct: 965 GNHEVSGD 988
>AI736705 similar to GP|11994736|db far-red impaired response protein;
Mutator-like transposase-like protein; phytochrome A
signaling, partial (14%)
Length = 383
Score = 28.5 bits (62), Expect = 8.6
Identities = 17/46 (36%), Positives = 26/46 (55%), Gaps = 3/46 (6%)
Frame = +3
Query: 76 DKRQRESRTDFR-CGCT-AKFRVHIDK-TNGRWYAKLFSDEHNHEL 118
+K+ E+ T R C T K +H+ + ++G+W F EHNHEL
Sbjct: 33 NKQDSENSTGRRSCSKTDCKASMHVKRRSDGKWVIHSFVKEHNHEL 170
>TC217518
Length = 1050
Score = 28.5 bits (62), Expect = 8.6
Identities = 11/23 (47%), Positives = 14/23 (60%)
Frame = +2
Query: 84 TDFRCGCTAKFRVHIDKTNGRWY 106
T C C K +V+ID T+G WY
Sbjct: 551 TFISCKCRRKMKVNIDFTSGLWY 619
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.340 0.149 0.499
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,332,342
Number of Sequences: 63676
Number of extensions: 366004
Number of successful extensions: 2396
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 2377
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2392
length of query: 578
length of database: 12,639,632
effective HSP length: 102
effective length of query: 476
effective length of database: 6,144,680
effective search space: 2924867680
effective search space used: 2924867680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0302a.8


