

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0301a.2
(80 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
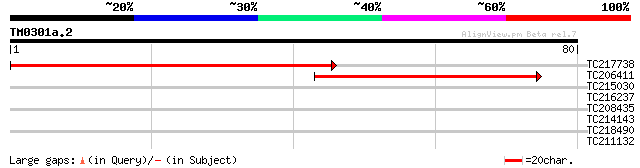
TC217738 homologue to UP|YCF2_LOTJA (Q9B1K6) Protein ycf2, parti... 79 3e-16
TC206411 homologue to UP|Q6L372 (Q6L372) Cytochrome b6 , partial... 38 8e-04
TC215030 31 0.13
TC216237 similar to UP|Q70WD8 (Q70WD8) RAC-ROP-like G-protein, p... 28 0.84
TC208435 28 1.1
TC214143 UP|Q39832 (Q39832) Ribulose-1,5-bisphosphate carboxylas... 27 2.4
TC218490 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (7%) 27 2.4
TC211132 homologue to UP|Q9XGX6 (Q9XGX6) Cellulose synthase cata... 26 3.2
>TC217738 homologue to UP|YCF2_LOTJA (Q9B1K6) Protein ycf2, partial (13%)
Length = 941
Score = 79.3 bits (194), Expect = 3e-16
Identities = 39/46 (84%), Positives = 41/46 (88%)
Frame = +1
Query: 1 FLLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLLD 46
F LLIH QRWLR N+SLSNGFFRSNTLSESYQYLSN+FLSN LLD
Sbjct: 700 FELLIHCQRWLRINSSLSNGFFRSNTLSESYQYLSNLFLSNEALLD 837
>TC206411 homologue to UP|Q6L372 (Q6L372) Cytochrome b6 , partial (95%)
Length = 2549
Score = 38.1 bits (87), Expect = 8e-04
Identities = 21/32 (65%), Positives = 25/32 (77%)
Frame = +3
Query: 44 LLDNTKMK*IIENSRGR*GSR*RQMSRLDILA 75
++ N KMK*I+E SRG SR*R+ SRLDILA
Sbjct: 3 IIHNKKMK*IVEYSRGLKRSR*RRRSRLDILA 98
>TC215030
Length = 493
Score = 30.8 bits (68), Expect = 0.13
Identities = 14/30 (46%), Positives = 20/30 (66%)
Frame = +2
Query: 25 NTLSESYQYLSNMFLSNGTLLDNTKMK*II 54
+TLS +Y Y+S +F SNG L T+M +I
Sbjct: 185 DTLSSTYMYISCIFFSNGAKLIQTRMPCVI 274
>TC216237 similar to UP|Q70WD8 (Q70WD8) RAC-ROP-like G-protein, partial (90%)
Length = 1651
Score = 28.1 bits (61), Expect = 0.84
Identities = 11/20 (55%), Positives = 16/20 (80%)
Frame = +1
Query: 6 HRQRWLRPNNSLSNGFFRSN 25
H QR+ RP++SL+NGF + N
Sbjct: 1138 HYQRFSRPSSSLANGFIQIN 1197
>TC208435
Length = 740
Score = 27.7 bits (60), Expect = 1.1
Identities = 15/35 (42%), Positives = 19/35 (53%)
Frame = -3
Query: 11 LRPNNSLSNGFFRSNTLSESYQYLSNMFLSNGTLL 45
+R LSN F+SNT Y+ + FLSN LL
Sbjct: 561 IRAGQILSNSLFKSNT------YVKHFFLSNAKLL 475
>TC214143 UP|Q39832 (Q39832) Ribulose-1,5-bisphosphate carboxylase small
subunit precursor (Ribulose-1,5-bisphosphate carboxylase
small subunit rbcS1), complete
Length = 1249
Score = 26.6 bits (57), Expect = 2.4
Identities = 11/14 (78%), Positives = 13/14 (92%)
Frame = +1
Query: 27 LSESYQYLSNMFLS 40
L ESYQY+SN+FLS
Sbjct: 715 LFESYQYVSNLFLS 756
>TC218490 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (7%)
Length = 689
Score = 26.6 bits (57), Expect = 2.4
Identities = 13/28 (46%), Positives = 18/28 (63%)
Frame = -2
Query: 21 FFRSNTLSESYQYLSNMFLSNGTLLDNT 48
FF + TL E +YLS +FLS +L+ T
Sbjct: 373 FFLNLTLQECTKYLSYVFLSFNLILEGT 290
>TC211132 homologue to UP|Q9XGX6 (Q9XGX6) Cellulose synthase catalytic
subunit, partial (19%)
Length = 615
Score = 26.2 bits (56), Expect = 3.2
Identities = 12/36 (33%), Positives = 20/36 (55%)
Frame = +3
Query: 2 LLLIHRQRWLRPNNSLSNGFFRSNTLSESYQYLSNM 37
LL H + WLR + L NGF + ++ + +LS +
Sbjct: 405 LLC*HLKLWLRHQSLLGNGFLSARNIALNLGHLSGI 512
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.350 0.154 0.482
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,912,927
Number of Sequences: 63676
Number of extensions: 26606
Number of successful extensions: 341
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 340
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 341
length of query: 80
length of database: 12,639,632
effective HSP length: 56
effective length of query: 24
effective length of database: 9,073,776
effective search space: 217770624
effective search space used: 217770624
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.9 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0301a.2


