

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0299.14
(88 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
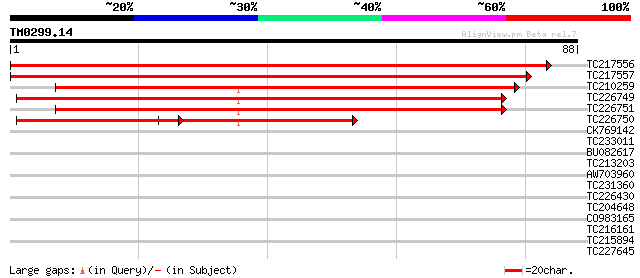
Score E
Sequences producing significant alignments: (bits) Value
TC217556 152 3e-38
TC217557 151 6e-38
TC210259 weakly similar to UP|Q6YU65 (Q6YU65) Hypoxia-responsive... 84 9e-18
TC226749 79 3e-16
TC226751 79 4e-16
TC226750 37 9e-04
CK769142 27 2.3
TC233011 similar to UP|Q9SDX4 (Q9SDX4) Dynamin homolog, partial ... 26 3.0
BU082617 26 3.9
TC213203 25 5.1
AW703960 similar to GP|7682677|gb| beta galactosidase {Vigna rad... 25 6.7
TC231360 similar to GB|CAE03203.1|32488346|OSJN00070 OSJNBb0060M... 25 6.7
TC226430 similar to GB|AAM70520.1|21700793|AY124811 At2g22250/T2... 25 6.7
TC204648 homologue to UP|Q9M5J4 (Q9M5J4) Beta galactosidase, com... 25 6.7
CO983165 25 6.7
TC216161 similar to UP|NO93_SOYBN (Q02921) Early nodulin 93 (N-9... 25 6.7
TC215894 similar to UP|Q96569 (Q96569) L-lactate dehydrogenase ... 25 8.7
TC227645 similar to UP|Q9FZK1 (Q9FZK1) F17L21.13, partial (61%) 25 8.7
>TC217556
Length = 654
Score = 152 bits (384), Expect = 3e-38
Identities = 73/84 (86%), Positives = 79/84 (93%)
Frame = +1
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
M+EAKTQ+ESIRKWVV+HKLRTVGCLWLSGI+GSIAYNWSRPNMKTSVKIIHARLHAQAL
Sbjct: 202 MSEAKTQIESIRKWVVEHKLRTVGCLWLSGISGSIAYNWSRPNMKTSVKIIHARLHAQAL 381
Query: 61 TLAALAGAAVVEYYDHRAEAKRAK 84
TL ALAGAA+VEYYD A AK +K
Sbjct: 382 TLGALAGAALVEYYDRNAGAKASK 453
>TC217557
Length = 748
Score = 151 bits (381), Expect = 6e-38
Identities = 72/81 (88%), Positives = 77/81 (94%)
Frame = +1
Query: 1 MAEAKTQVESIRKWVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 60
M+EAKTQ+ESIRKWVV+HKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL
Sbjct: 274 MSEAKTQIESIRKWVVEHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQAL 453
Query: 61 TLAALAGAAVVEYYDHRAEAK 81
TL ALAGAA+VEYYD + AK
Sbjct: 454 TLGALAGAALVEYYDRKTGAK 516
>TC210259 weakly similar to UP|Q6YU65 (Q6YU65) Hypoxia-responsive family
protein-like, partial (36%)
Length = 424
Score = 84.3 bits (207), Expect = 9e-18
Identities = 42/73 (57%), Positives = 56/73 (76%), Gaps = 1/73 (1%)
Frame = +2
Query: 8 VESIRKWVVDHKLRTVGCLWLSGITGS-IAYNWSRPNMKTSVKIIHARLHAQALTLAALA 66
+++I+ WV HKL TVG LW SGI S +AY+ +R MK S+++IHARLHAQALTLA L+
Sbjct: 38 MDAIQLWVSKHKLATVGGLWASGIGASLVAYSRTRSPMKPSLRLIHARLHAQALTLAVLS 217
Query: 67 GAAVVEYYDHRAE 79
GAA YY++RA+
Sbjct: 218 GAAAYRYYENRAD 256
>TC226749
Length = 631
Score = 79.3 bits (194), Expect = 3e-16
Identities = 36/77 (46%), Positives = 55/77 (70%), Gaps = 1/77 (1%)
Frame = +1
Query: 2 AEAKTQVESIRKWVVDHKLRTVGCLWLSGITGS-IAYNWSRPNMKTSVKIIHARLHAQAL 60
++ + +E+++ WV HKL ++G LW SGI + +AY+ + MK S+++IHAR+HAQAL
Sbjct: 52 SDQRYNMEALQSWVSKHKLASIGALWASGIGATLVAYSCKKSPMKPSLRLIHARMHAQAL 231
Query: 61 TLAALAGAAVVEYYDHR 77
TLA L+GAA YY+ R
Sbjct: 232 TLAVLSGAAAYHYYEKR 282
>TC226751
Length = 529
Score = 79.0 bits (193), Expect = 4e-16
Identities = 36/71 (50%), Positives = 53/71 (73%), Gaps = 1/71 (1%)
Frame = +1
Query: 8 VESIRKWVVDHKLRTVGCLWLSGITGS-IAYNWSRPNMKTSVKIIHARLHAQALTLAALA 66
+E+++ WV HKL ++G LW SGI + +AY+ + MK S+++IHAR+HAQALTLA L+
Sbjct: 55 METVQSWVSKHKLASIGALWASGIGATLVAYSCKKSPMKPSLRLIHARMHAQALTLAVLS 234
Query: 67 GAAVVEYYDHR 77
GAAV +Y+ R
Sbjct: 235 GAAVYHFYEKR 267
>TC226750
Length = 746
Score = 36.6 bits (83), Expect = 0.002
Identities = 16/24 (66%), Positives = 19/24 (78%)
Frame = +3
Query: 54 RLHAQALTLAALAGAAVVEYYDHR 77
R+HAQALTLA L+GAA YY+ R
Sbjct: 459 RMHAQALTLAVLSGAAAYHYYEKR 530
Score = 32.3 bits (72), Expect(2) = 9e-04
Identities = 15/32 (46%), Positives = 22/32 (67%), Gaps = 1/32 (3%)
Frame = +2
Query: 24 GCLWLSGITGS-IAYNWSRPNMKTSVKIIHAR 54
G LW SGI + +AY+ + MK S+++IHAR
Sbjct: 212 GALWASGIGATLVAYSCKKSPMKPSLRLIHAR 307
Score = 24.6 bits (52), Expect(2) = 9e-04
Identities = 8/26 (30%), Positives = 18/26 (68%)
Frame = +3
Query: 2 AEAKTQVESIRKWVVDHKLRTVGCLW 27
++ + +E+++ WV HKL ++G L+
Sbjct: 63 SDQRYNMEALQSWVSKHKLASIG*LY 140
>CK769142
Length = 753
Score = 26.6 bits (57), Expect = 2.3
Identities = 17/61 (27%), Positives = 27/61 (43%), Gaps = 1/61 (1%)
Frame = -1
Query: 3 EAKTQVESIRK-WVVDHKLRTVGCLWLSGITGSIAYNWSRPNMKTSVKIIHARLHAQALT 61
++KTQ ES V D + + W+S +T SIA+ + + I+ H T
Sbjct: 633 KSKTQTESFNSSCVTDPIMNGIRRAWISSLTNSIAFVRIATLPSSKLAILRCNFHDSPST 454
Query: 62 L 62
L
Sbjct: 453 L 451
>TC233011 similar to UP|Q9SDX4 (Q9SDX4) Dynamin homolog, partial (3%)
Length = 347
Score = 26.2 bits (56), Expect = 3.0
Identities = 14/49 (28%), Positives = 21/49 (42%), Gaps = 5/49 (10%)
Frame = -1
Query: 7 QVESIRKWVVDHKLRTVGCLWLSG-----ITGSIAYNWSRPNMKTSVKI 50
+VE R W + H+LR G WL +T + W +M +I
Sbjct: 281 KVEVFRNWRIRHRLRRGGVSWLGASNQRVLTSPQSLPWMNQSMDPKRRI 135
>BU082617
Length = 430
Score = 25.8 bits (55), Expect = 3.9
Identities = 17/49 (34%), Positives = 27/49 (54%), Gaps = 5/49 (10%)
Frame = +3
Query: 26 LWLSG--ITGSIAYNWSRPNMKTSVKIIHARLHA---QALTLAALAGAA 69
LW+ +T + +++ PNM TS+ +H RL A + LA L GA+
Sbjct: 12 LWVVSLHVTFVLCFHFVAPNMFTSLLNVHTRLTACFVKGKELAFLGGAS 158
>TC213203
Length = 459
Score = 25.4 bits (54), Expect = 5.1
Identities = 10/25 (40%), Positives = 13/25 (52%)
Frame = +3
Query: 50 IIHARLHAQALTLAALAGAAVVEYY 74
+IH+RLH T+ G A YY
Sbjct: 261 VIHSRLHRSFTTIRPRGGTATSNYY 335
>AW703960 similar to GP|7682677|gb| beta galactosidase {Vigna radiata},
partial (16%)
Length = 370
Score = 25.0 bits (53), Expect = 6.7
Identities = 7/14 (50%), Positives = 11/14 (78%)
Frame = +2
Query: 25 CLWLSGITGSIAYN 38
CLW+ G+T S+ Y+
Sbjct: 35 CLWVCGVTASVTYD 76
>TC231360 similar to GB|CAE03203.1|32488346|OSJN00070 OSJNBb0060M15.15 {Oryza
sativa (japonica cultivar-group);} , partial (17%)
Length = 707
Score = 25.0 bits (53), Expect = 6.7
Identities = 8/19 (42%), Positives = 16/19 (84%)
Frame = +1
Query: 8 VESIRKWVVDHKLRTVGCL 26
+ S+RKW++D K +++GC+
Sbjct: 262 IYSLRKWLLDCK*QSLGCV 318
>TC226430 similar to GB|AAM70520.1|21700793|AY124811 At2g22250/T26C19.9
{Arabidopsis thaliana;} , partial (33%)
Length = 686
Score = 25.0 bits (53), Expect = 6.7
Identities = 11/29 (37%), Positives = 17/29 (57%), Gaps = 1/29 (3%)
Frame = +1
Query: 9 ESIRKWVVDHKLRTVGCL-WLSGITGSIA 36
E+IR W+ HK++ CL W IT ++
Sbjct: 559 ENIRLWIFFHKMK*YNCL*WFKIITTKLS 645
>TC204648 homologue to UP|Q9M5J4 (Q9M5J4) Beta galactosidase, complete
Length = 2817
Score = 25.0 bits (53), Expect = 6.7
Identities = 7/14 (50%), Positives = 11/14 (78%)
Frame = +3
Query: 25 CLWLSGITGSIAYN 38
CLW+ G+T S+ Y+
Sbjct: 114 CLWVCGVTASVTYD 155
>CO983165
Length = 855
Score = 25.0 bits (53), Expect = 6.7
Identities = 8/19 (42%), Positives = 16/19 (84%)
Frame = -1
Query: 8 VESIRKWVVDHKLRTVGCL 26
+ S+RKW++D K +++GC+
Sbjct: 390 IYSLRKWLLDCK*QSLGCV 334
>TC216161 similar to UP|NO93_SOYBN (Q02921) Early nodulin 93 (N-93), partial
(93%)
Length = 534
Score = 25.0 bits (53), Expect = 6.7
Identities = 17/48 (35%), Positives = 22/48 (45%)
Frame = +2
Query: 39 WSRPNMKTSVKIIHARLHAQALTLAALAGAAVVEYYDHRAEAKRAKNS 86
W+R N+ S AQAL ++ +AGAA D A KNS
Sbjct: 251 WARANLNHS---------AQALIISTVAGAAYFIVADKTVLATARKNS 367
>TC215894 similar to UP|Q96569 (Q96569) L-lactate dehydrogenase , complete
Length = 1675
Score = 24.6 bits (52), Expect = 8.7
Identities = 10/25 (40%), Positives = 14/25 (56%)
Frame = -2
Query: 32 TGSIAYNWSRPNMKTSVKIIHARLH 56
T S+ WS PN+ SV +H+ H
Sbjct: 1197 TESVLREWSWPNL*ASVSPLHSSAH 1123
>TC227645 similar to UP|Q9FZK1 (Q9FZK1) F17L21.13, partial (61%)
Length = 1671
Score = 24.6 bits (52), Expect = 8.7
Identities = 14/49 (28%), Positives = 27/49 (54%), Gaps = 2/49 (4%)
Frame = +3
Query: 18 HKLRTVGCLWLSGITGSIAYNWSRP-NMKTSVKI-IHARLHAQALTLAA 64
+K+ VGC + S+ +WSRP NM +K+ + +QA+++ +
Sbjct: 795 YKILWVGCDGEYEVYDSVRNSWSRPGNMPAGMKLPLSINFRSQAVSIGS 941
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.127 0.376
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,827,863
Number of Sequences: 63676
Number of extensions: 43948
Number of successful extensions: 266
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 262
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 263
length of query: 88
length of database: 12,639,632
effective HSP length: 64
effective length of query: 24
effective length of database: 8,564,368
effective search space: 205544832
effective search space used: 205544832
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0299.14


