

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
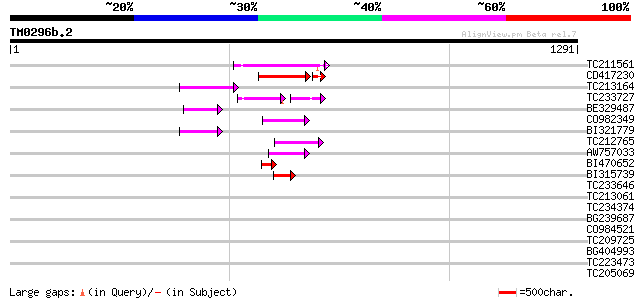
Query= TM0296b.2
(1291 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC211561 weakly similar to UP|O22675 (O22675) Reverse transcript... 106 6e-23
CD417230 weakly similar to GP|2244915|emb| reverse transcriptase... 91 3e-19
TC213164 87 4e-17
TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR ret... 60 9e-13
BE329487 weakly similar to PIR|T14619|T146 reverse transcriptase... 64 3e-10
CO982349 55 2e-07
BI321779 54 6e-07
TC212765 50 6e-06
AW757033 47 4e-05
BI470652 similar to PIR|T17102|T171 RNA-directed DNA polymerase ... 45 2e-04
BI315739 45 3e-04
TC233646 35 0.28
TC213061 34 0.48
TC234374 31 0.82
BG239687 32 1.8
CO984521 32 1.8
TC209725 similar to UP|Q6N4V2 (Q6N4V2) 50S ribosomal protein L15... 31 3.1
BG404993 30 5.3
TC223473 30 5.3
TC205069 similar to UP|Q9SJ31 (Q9SJ31) Expressed protein, partia... 30 6.9
>TC211561 weakly similar to UP|O22675 (O22675) Reverse transcriptase
(Fragment) , partial (59%)
Length = 1077
Score = 106 bits (265), Expect = 6e-23
Identities = 81/223 (36%), Positives = 111/223 (49%), Gaps = 5/223 (2%)
Frame = +1
Query: 511 MDNAFLA*EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVIIDLIM 570
+ +A +A E+I + RS K L FK+D EKAYDS+SW+FL + +F I I
Sbjct: 1 LHSALIANEVIDEAKRS--NKSCLVFKVDYEKAYDSVSWNFLMYMMWRMDFSPKWIKWIE 174
Query: 571 CYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGRWKPL 630
V S+ IS+L NG+P F P RGLRQGDP +P+ F + E L+ ++ VE +KP
Sbjct: 175 ECVKSASISVLVNGSPTAEFLPQRGLRQGDPLTPFLFNIVAEGLNGLMRRAVEENLYKPY 354
Query: 631 HRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAITSIGV 690
A G IS L +A D + F +A+ V + L+ S LRIN KS GV
Sbjct: 355 LVGANGVPISILQYADDTIFFGEAAKENVEAIKVILRSFELVSNLRINFAKS-CFGVFGV 531
Query: 691 APEVRTEIG-----LRALIPFVNDLGKYLGFPLQGRRLNSERC 728
+ + E PF+ YLG P+ NS RC
Sbjct: 532 TDQWKQEAANYLHCSLLAFPFI-----YLGIPIGA---NSRRC 636
>CD417230 weakly similar to GP|2244915|emb| reverse transcriptase like
protein {Arabidopsis thaliana}, partial (7%)
Length = 688
Score = 90.5 bits (223), Expect(2) = 3e-19
Identities = 50/120 (41%), Positives = 75/120 (61%), Gaps = 1/120 (0%)
Frame = -3
Query: 566 IDLIMCYVSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPG 625
+ L+ +S++ +SLLWNG L F P G+RQ DP +PY FVLC+ERLS H+ NLV
Sbjct: 686 VQLVWYCMSTTTMSLLWNGEVLDHFSPNIGVRQRDPIAPYLFVLCLERLS-HLINLVMNK 510
Query: 626 RW-KPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKA 684
+ + + GG ISHL F D++LF +A++ QV V+ L+ + S ++NV K+K+
Sbjct: 509 KL*RSIQVNRGGLLISHLAFVDDLILFVEANMDQVDVINLTLELFSDSSGAKVNVDKTKS 330
Score = 24.3 bits (51), Expect(2) = 3e-19
Identities = 11/29 (37%), Positives = 19/29 (64%)
Frame = -2
Query: 690 VAPEVRTEIGLRALIPFVNDLGKYLGFPL 718
V ++ + +GL++ +DLGKYLG P+
Sbjct: 300 VEEDISSGLGLQS----TDDLGKYLGVPI 226
>TC213164
Length = 446
Score = 87.4 bits (215), Expect = 4e-17
Identities = 47/135 (34%), Positives = 74/135 (54%)
Frame = +3
Query: 387 LLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
LLV K+EVR + + +PGPDG F K++W+ + + + + E + +G +
Sbjct: 15 LLVARFEKKEVRATMWDRGNAKSPGPDGLNFKFIKEFWDILKTDLLRFLDEFYANGVFLK 194
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLLVNRIRPFLNDIVGPMQNSFLP 506
R + LIPKV+ P S+ E++PISL YK++ KLL R + L I+ Q F+
Sbjct: 195 RSNAFFLALIPKVHDP*SLNEYKPISLIGCIYKIVAKLLSGRHKKVLPTIIDERQTVFME 374
Query: 507 GRGTMDNAFLA*EII 521
GR T+ N +A EI+
Sbjct: 375 GRHTLHNVVIANEIM 419
>TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR retroelement
reverse transcriptase, partial (7%)
Length = 956
Score = 60.5 bits (145), Expect(2) = 9e-13
Identities = 36/115 (31%), Positives = 63/115 (54%), Gaps = 6/115 (5%)
Frame = -3
Query: 519 EIIHQMHRSMARKGNLAFKIDLEKAYDSISWSFLRETLELYEFPLVIIDLIMCYVSSSQI 578
E+I+ + + + GNLA KID+ KA+D++ FL L + + + I + S+++
Sbjct: 807 EVINMLDKKVFG-GNLALKIDIVKAFDTLDRDFLISVLNKFGYSHTFCNWIRVILLSAKL 631
Query: 579 SLLWNGAPLPSFKPGRGLRQGDPKSPYFF-VLCMERLSIHIQ-----NLVEPGRW 627
S+ NG P+ F RG+RQGDP SP + C+E+ +I++ +P +W
Sbjct: 630 SISVNGEPVGFFSCQRGVRQGDPLSPSLYRQRCLEQGTIYVDF*RKVATYDPSQW 466
Score = 32.3 bits (72), Expect(2) = 9e-13
Identities = 24/80 (30%), Positives = 37/80 (46%), Gaps = 1/80 (1%)
Frame = -2
Query: 640 SHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSKAITSIGVAPEVRTEIG 699
SH+ A D+++FCK + V L++ Q + I+ KSK IG P R +
Sbjct: 451 SHVMHADDIMIFCKGTSRNVHNLINFFQLYKDSFGQVISKHKSKIF--IGSIPGQRLRVI 278
Query: 700 LRALIPFVNDLG-KYLGFPL 718
L F +D+ Y G P+
Sbjct: 277 SDLLQYFHSDIPFNYFGVPI 218
>BE329487 weakly similar to PIR|T14619|T146 reverse transcriptase - beet
retrotransposon (fragment), partial (15%)
Length = 376
Score = 64.3 bits (155), Expect = 3e-10
Identities = 29/88 (32%), Positives = 49/88 (54%)
Frame = -2
Query: 397 VRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDERLLEILVVLI 456
++ A+ S +PGPDG F K +WE++ + + ++E +G + + LI
Sbjct: 375 IKSAVWQCGSVKSPGPDGLNFKFIKHFWERLKPDIIRFLNEFHANGIFPKGGNASFIALI 196
Query: 457 PKVNHPTSIKEFRPISLCNVTYKLITKL 484
PKV HP ++ +FRPISL YK++ K+
Sbjct: 195 PKVKHPQALNDFRPISLIGCVYKIVAKI 112
>CO982349
Length = 795
Score = 55.5 bits (132), Expect = 2e-07
Identities = 34/109 (31%), Positives = 58/109 (53%)
Frame = +3
Query: 575 SSQISLLWNGAPLPSFKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGRWKPLHRTA 634
S+ +S+L N +P+ F P RGLRQGDP +P F + E L+ ++ V+ R+
Sbjct: 90 SASVSILVNESPINEFSPQRGLRQGDPLAPLLFNIVAEGLTGLMREAVDRKRFNSFLVGK 269
Query: 635 GGTGISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSK 683
+S L +A D + F +A++ + V+ L+ S L+IN +S+
Sbjct: 270 NKEPVSILQYADDTIFFEEATMENMKVIKTILRCFELASGLKINFAESR 416
>BI321779
Length = 421
Score = 53.5 bits (127), Expect = 6e-07
Identities = 27/97 (27%), Positives = 48/97 (48%)
Frame = +2
Query: 387 LLVQPVSKEEVRQALMSMKSFTAPGPDGFQPFFFKQYWEQVGDTVWKVVSEAFESGKVDE 446
+L+Q EE+++ + +S +PGPDG+ K +WE + + V E + +
Sbjct: 80 MLIQDFEVEEIKKVI*DCESSKSPGPDGYSLLLIKIFWEVIKEEVINFFKEFHKFDFLPR 259
Query: 447 RLLEILVVLIPKVNHPTSIKEFRPISLCNVTYKLITK 483
+ LI K ++P I ++ PISL YK++ K
Sbjct: 260 GTNRSFIALIAKCDNPLDIGQYCPISLVGCLYKIV*K 370
>TC212765
Length = 637
Score = 50.1 bits (118), Expect = 6e-06
Identities = 35/111 (31%), Positives = 51/111 (45%)
Frame = +3
Query: 604 PYFFVLCMERLSIHIQNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLFCKASVAQVTVLV 663
PY FVLCMERL++ I +L W+P+ + + +L D+ LF A +
Sbjct: 15 PYLFVLCMERLALCIHDLTSQ*IWRPILVSDDNRLVPYLMLVDDIFLFINAIGD*AYLAS 194
Query: 664 DALQFLCEGSVLRINVQKSKAITSIGVAPEVRTEIGLRALIPFVNDLGKYL 714
L + L++N+ KS S +AP +R I I LGKYL
Sbjct: 195 LTL*NFFQTFGLKLNLDKSSMYCS*RMAPVLRDLISNTLNIVHAPSLGKYL 347
>AW757033
Length = 441
Score = 47.4 bits (111), Expect = 4e-05
Identities = 31/94 (32%), Positives = 47/94 (49%)
Frame = -2
Query: 590 FKPGRGLRQGDPKSPYFFVLCMERLSIHIQNLVEPGRWKPLHRTAGGTGISHLFFAGDVL 649
F P RGLRQGDP +P F + E L+ ++ V +K +S L +A D +
Sbjct: 431 FSPHRGLRQGDPLAPLLFNIAAEGLTGLMREAVARNHYKSFAVGKYKKPVSILQYADDTI 252
Query: 650 LFCKASVAQVTVLVDALQFLCEGSVLRINVQKSK 683
F +A++ V V+ L+ S L+IN KS+
Sbjct: 251 FFGEATMENVRVIKTILRGFELASGLKINFAKSR 150
>BI470652 similar to PIR|T17102|T171 RNA-directed DNA polymerase (EC
2.7.7.49) (clone AmLi2) - garden snapdragon (fragment),
partial (22%)
Length = 111
Score = 45.4 bits (106), Expect = 2e-04
Identities = 20/34 (58%), Positives = 25/34 (72%)
Frame = +1
Query: 573 VSSSQISLLWNGAPLPSFKPGRGLRQGDPKSPYF 606
+SS+ IS+L NG+P FKP RGLRQG P P+F
Sbjct: 10 LSSASISILVNGSPTEEFKPXRGLRQGGPFGPFF 111
>BI315739
Length = 442
Score = 44.7 bits (104), Expect = 3e-04
Identities = 20/51 (39%), Positives = 32/51 (62%)
Frame = +3
Query: 601 PKSPYFFVLCMERLSIHIQNLVEPGRWKPLHRTAGGTGISHLFFAGDVLLF 651
P SPY F +CME+L+I IQ+ V W+P+ + G GIS ++ + ++ F
Sbjct: 48 PLSPYLF*ICMEKLAILIQSKVNEETWEPIKISRNGPGISPIYSSQIIVYF 200
Score = 39.3 bits (90), Expect = 0.011
Identities = 21/45 (46%), Positives = 29/45 (63%)
Frame = +2
Query: 639 ISHLFFAGDVLLFCKASVAQVTVLVDALQFLCEGSVLRINVQKSK 683
ISHLFF + LLF KA+ +QV ++ D LQ C L+ +QKS+
Sbjct: 164 ISHLFFPDNCLLFVKATSSQVKLVRDVLQQFCLT*GLKAGLQKSR 298
>TC233646
Length = 978
Score = 34.7 bits (78), Expect = 0.28
Identities = 21/62 (33%), Positives = 37/62 (58%), Gaps = 1/62 (1%)
Frame = -3
Query: 451 ILVVLIPKVNHPTSIKEFRPISLCNVTYKLITKLL-VNRIRPFLNDIVGPMQNSFLPGRG 509
I +VLIPK+++ +K + ISLCNVT +L+++ + F + + Q SF+P +G
Sbjct: 832 ISLVLIPKIDNLEILK*Y*SISLCNVTTRLLSRSS*PTSSKSFPS*PIARNQCSFIPSKG 653
Query: 510 TM 511
+
Sbjct: 652 AI 647
>TC213061
Length = 823
Score = 33.9 bits (76), Expect = 0.48
Identities = 14/41 (34%), Positives = 21/41 (51%)
Frame = +2
Query: 845 EDQWIWGPGSRGTYLVKEAYSWLHERNRVSAGAGNWRWLWR 885
EDQW+W S G+YL + AY + E ++ LW+
Sbjct: 125 EDQWVWAAESSGSYLARSAYRVIREGIPEEEQDREFKELWK 247
>TC234374
Length = 787
Score = 30.8 bits (68), Expect(2) = 0.82
Identities = 16/42 (38%), Positives = 24/42 (57%)
Frame = +1
Query: 416 QPFFFKQYWEQVGDTVWKVVSEAFESGKVDERLLEILVVLIP 457
Q FF+ Y + VGDTV+++V + F+S + L VL P
Sbjct: 637 QRIFFQHY*DAVGDTVYQLVLQLFKSRWILPNLNSNTAVLFP 762
Score = 20.8 bits (42), Expect(2) = 0.82
Identities = 9/24 (37%), Positives = 11/24 (45%)
Frame = +2
Query: 395 EEVRQALMSMKSFTAPGPDGFQPF 418
EEV+ + M PG DG F
Sbjct: 575 EEVKSLVFDMNGEGVPGTDGCNGF 646
>BG239687
Length = 443
Score = 32.0 bits (71), Expect = 1.8
Identities = 13/44 (29%), Positives = 21/44 (47%)
Frame = -3
Query: 844 QEDQWIWGPGSRGTYLVKEAYSWLHERNRVSAGAGNWRWLWRGS 887
+ED W+W G Y V++AY + V A + +W+ S
Sbjct: 234 EEDSWVWKSNESGGYNVRDAYMLILHDGLVDEHANILKQVWKAS 103
>CO984521
Length = 716
Score = 32.0 bits (71), Expect = 1.8
Identities = 15/47 (31%), Positives = 19/47 (39%)
Frame = -2
Query: 839 TLSTAQEDQWIWGPGSRGTYLVKEAYSWLHERNRVSAGAGNWRWLWR 885
T+ DQW W G Y K AY LH G ++ LW+
Sbjct: 667 TIRPELSDQWKWAAEPSGCYSTKSAYKALHHVTVGEEQDGKFKELWK 527
>TC209725 similar to UP|Q6N4V2 (Q6N4V2) 50S ribosomal protein L15, partial
(35%)
Length = 612
Score = 31.2 bits (69), Expect = 3.1
Identities = 9/14 (64%), Positives = 11/14 (78%)
Frame = +1
Query: 878 GNWRWLWRGSDCRE 891
G W W+W+G DCRE
Sbjct: 415 GAWDWVWKGQDCRE 456
>BG404993
Length = 412
Score = 30.4 bits (67), Expect = 5.3
Identities = 18/49 (36%), Positives = 21/49 (42%), Gaps = 3/49 (6%)
Frame = -3
Query: 840 LSTAQE---DQWIWGPGSRGTYLVKEAYSWLHERNRVSAGAGNWRWLWR 885
LST Q+ D W W G Y K AY L E R G + LW+
Sbjct: 359 LSTIQQHEADSWTWKADPSGNYSTKTAYELLQEAIRGDNEDGIFVQLWK 213
>TC223473
Length = 654
Score = 30.4 bits (67), Expect = 5.3
Identities = 11/31 (35%), Positives = 20/31 (64%)
Frame = +1
Query: 327 RKRNSVHRLKLSDGSWCSDTSKLQEEALGYF 357
R+ N +H+++ S G W +D S + +E + YF
Sbjct: 400 RQLNFIHKIQNSSGEWITDIS*IHQEMIVYF 492
>TC205069 similar to UP|Q9SJ31 (Q9SJ31) Expressed protein, partial (58%)
Length = 787
Score = 30.0 bits (66), Expect = 6.9
Identities = 24/80 (30%), Positives = 36/80 (45%), Gaps = 22/80 (27%)
Frame = +1
Query: 5 MEPKCQSLRVRRFWTS-------------------LGFSFAFVMEAVGFSGGIWV---LT 42
M PK L VRR+W + G S FV+ A+ F+ G W+ +
Sbjct: 385 MFPKAFLLNVRRYWQTAGEREGENPMKTPLPYIIIFGMSTPFVILAIAFANG-WIKVPVR 561
Query: 43 NNSSNLSFRLVDLHAQVVTF 62
+S++LS L+DLH + F
Sbjct: 562 *DSTSLSTLLLDLHMLIHEF 621
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.340 0.149 0.508
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 60,028,532
Number of Sequences: 63676
Number of extensions: 895645
Number of successful extensions: 6347
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 6139
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 6332
length of query: 1291
length of database: 12,639,632
effective HSP length: 108
effective length of query: 1183
effective length of database: 5,762,624
effective search space: 6817184192
effective search space used: 6817184192
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0296b.2


