

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0296a.5
(754 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
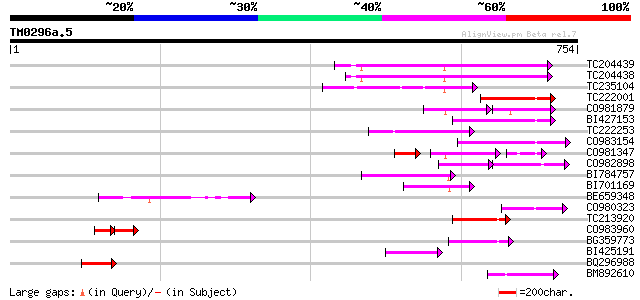
Score E
Sequences producing significant alignments: (bits) Value
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 149 4e-36
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 145 6e-35
TC235104 weakly similar to UP|Q850H8 (Q850H8) Gag-pol polyprotei... 113 4e-25
TC222001 similar to UP|C716_NEPRA (O04164) Cytochrome P450 71A6 ... 102 8e-22
CO981879 72 3e-19
BI427153 91 2e-18
TC222253 similar to UP|Q9ZQE4 (Q9ZQE4) Copia-like retroelement p... 90 3e-18
CO983154 84 3e-16
CO981347 58 1e-14
CO982898 51 3e-12
BI784757 69 1e-11
BI701169 67 3e-11
BE659348 weakly similar to PIR|JC7809|JC78 sulfakinin receptor p... 64 2e-10
CO980323 53 4e-07
TC213920 53 4e-07
CO983960 38 5e-07
BG359773 47 2e-05
BI425191 weakly similar to GP|14586969|gb| pol polyprotein {Citr... 47 4e-05
BQ296988 similar to GP|21740616|em OSJNBb0089K24.12 {Oryza sativ... 46 5e-05
BM892610 45 9e-05
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 149 bits (376), Expect = 4e-36
Identities = 94/306 (30%), Positives = 151/306 (48%), Gaps = 15/306 (4%)
Frame = +1
Query: 432 SVNMSTVSSSNKLPNIWHHRLGHPSQPCYISLQQM-----------YPPITTSPCKNCDT 480
S + + +SS IWH R GH + L+ M P + + C
Sbjct: 2026 SYSSTCLSSKEDEVRIWHQRFGH------LHLRGMKKIIDKGAVRGIPNLKIEEGRICGE 2187
Query: 481 CHFAKQKRLSFS-LRNTIYGTVLELLHVDIWGPYSVASITGAKYFHIIVYDKSRFTWIKL 539
C KQ ++S L++ VLELLH+D+ GP V S+ G +Y +++V D SRFTW+
Sbjct: 2188 CQIGKQVKMSHQKLQHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTWVNF 2367
Query: 540 MKAKSEARAALQTFILYSQRQYGSLVKIVRLDNGVEF---AMTDFYSQHGVQHQLSCVKT 596
++ KSE + L QR+ ++K +R D+G EF T+F + G+ H+ S T
Sbjct: 2368 IREKSETFEVFKELSLRLQREKDCVIKRIRSDHGREFENSRFTEFCTSEGITHEFSAAIT 2547
Query: 597 PQQNGVVERKHQHILNIARALMLQSNLTLEFWSFAVIHAVCLMNLLPTKVLSYSSPFIIL 656
PQQNG+VERK++ + AR ++ L W+ A+ A + N + + + ++ + I
Sbjct: 2548 PQQNGIVERKNRTLQEAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLYEIW 2727
Query: 657 HNLKPDIEHLKVFGSLCYATSLTAHRTKFPLRARKCIVLGQKEGGVKGYLMFDLKTREVF 716
KP ++H +FGS CY + R K ++ I LG + Y +F+ +TR V
Sbjct: 2728 KGRKPSVKHFHIFGSPCYILADREQRRKMDPKSDAGIFLGYSTNS-RAYRVFNSRTRTVM 2904
Query: 717 LSRNVI 722
S NV+
Sbjct: 2905 ESINVV 2922
Score = 30.8 bits (68), Expect = 2.3
Identities = 14/51 (27%), Positives = 21/51 (40%)
Frame = +1
Query: 171 TGNTQGTHSNSNRGRSNSQYSSSKSNV*RLCTHCGRQNHTVETCFLKHGYP 221
TG T H + + G + K C +CG+ H C+ HG+P
Sbjct: 1432 TGATMSQHRSRHHGMQQKKSKRKKWR----CHYCGKYGHIKPFCYHLHGHP 1572
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 145 bits (366), Expect = 6e-35
Identities = 90/291 (30%), Positives = 145/291 (48%), Gaps = 15/291 (5%)
Frame = +1
Query: 447 IWHHRLGHPSQPCYISLQQM-----------YPPITTSPCKNCDTCHFAKQKRLSFS-LR 494
IWH R GH + L+ M P + + C C KQ ++S L+
Sbjct: 2074 IWHQRFGH------LHLRGMKKIIDKGAVRGIPNLKIEEGRICGECQIGKQVKMSHQKLQ 2235
Query: 495 NTIYGTVLELLHVDIWGPYSVASITGAKYFHIIVYDKSRFTWIKLMKAKSEARAALQTFI 554
+ VLELLH+D+ GP V S+ G +Y +++V D SRFTW+ ++ KS+ +
Sbjct: 2236 HQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTWVNFIREKSDTFEVFKELS 2415
Query: 555 LYSQRQYGSLVKIVRLDNGVEF---AMTDFYSQHGVQHQLSCVKTPQQNGVVERKHQHIL 611
L QR+ ++K +R D+G EF T+F + G+ H+ S TPQQNG+VERK++ +
Sbjct: 2416 LRLQREKDCVIKRIRSDHGREFENSKFTEFCTSEGITHEFSAAITPQQNGIVERKNRTLQ 2595
Query: 612 NIARALMLQSNLTLEFWSFAVIHAVCLMNLLPTKVLSYSSPFIILHNLKPDIEHLKVFGS 671
AR ++ L W+ A+ A + N + + + ++ + I KP ++H +FGS
Sbjct: 2596 EAARVMLHAKELPYNLWAEAMNTACYIHNRVTLRRGTPTTLYEIWKGRKPTVKHFHIFGS 2775
Query: 672 LCYATSLTAHRTKFPLRARKCIVLGQKEGGVKGYLMFDLKTREVFLSRNVI 722
CY + R K ++ I LG + Y +F+ +TR V S NV+
Sbjct: 2776 PCYILADREQRRKMDPKSDAGIFLGYSTNS-RAYRVFNSRTRTVMESINVV 2925
Score = 31.6 bits (70), Expect = 1.4
Identities = 17/51 (33%), Positives = 24/51 (46%)
Frame = +1
Query: 171 TGNTQGTHSNSNRGRSNSQYSSSKSNV*RLCTHCGRQNHTVETCFLKHGYP 221
TG T H + + G +Q SK R C +CG+ H C+ HG+P
Sbjct: 1435 TGATMSQHRSRHHG---TQQKKSKRKKWR-CHYCGKYGHIKPFCYHLHGHP 1575
>TC235104 weakly similar to UP|Q850H8 (Q850H8) Gag-pol polyprotein
(Fragment), partial (28%)
Length = 865
Score = 113 bits (282), Expect = 4e-25
Identities = 70/211 (33%), Positives = 113/211 (53%), Gaps = 5/211 (2%)
Frame = +2
Query: 417 TSWSFHLSLVSN--HFHSVNMSTVSSSNKLPNIWHHRLGHPSQPCYISLQQMYPPITTSP 474
T W+ + + S+ ++ N+S V S+ P + H RLGHP L+ M P +
Sbjct: 215 TGWTIGVGIESHGLYYLKPNLSWVCSAVTSPKLLHERLGHPHLS---KLKIMVPSLEKIK 385
Query: 475 CKNCDTCHFAKQKRLSFSLRNTIYGTVLELLHVDIWGPYSVASITGAKYFHIIVYDKSRF 534
C++C K R S + + ++H DIWGP V+S++ +YF + + S+
Sbjct: 386 DLFCESCQLGKHVRSSXRHVESRVDSPFLVIHXDIWGPNRVSSMS-YRYFVTFIDEFSQC 562
Query: 535 TWIKLMKAKSEARAALQTFILYSQRQYGSLVKIVRLDNGVEF---AMTDFYSQHGVQHQL 591
T + LMK +SE + L T + + Q+G +KI+R DN E+ ++ F S G+ HQ
Sbjct: 563 TRVFLMKERSEILSFL-TSVNKIKTQFGKTIKILRSDNAKEYFSSVISPFXSAQGILHQF 739
Query: 592 SCVKTPQQNGVVERKHQHILNIARALMLQSN 622
SC TPQQN + ERK++H++ AR L+L +N
Sbjct: 740 SCPHTPQQNDIAERKNRHLVETARTLLLHAN 832
>TC222001 similar to UP|C716_NEPRA (O04164) Cytochrome P450 71A6 (Fragment)
, partial (21%)
Length = 912
Score = 102 bits (253), Expect = 8e-22
Identities = 46/99 (46%), Positives = 67/99 (67%)
Frame = -2
Query: 627 FWSFAVIHAVCLMNLLPTKVLSYSSPFIILHNLKPDIEHLKVFGSLCYATSLTAHRTKFP 686
FW++A++HA L+N +PT L +SP+ LH PDI HL++FG LCYA+++ A+R K
Sbjct: 899 FWNYALLHAAYLINCIPTPFLQNTSPYERLHGHIPDISHLRIFGCLCYASTIKANRKKLE 720
Query: 687 LRARKCIVLGQKEGGVKGYLMFDLKTREVFLSRNVIFYE 725
RA CI +G K KGY+++DL + + SRNV+FYE
Sbjct: 719 PRAHPCIFIGFKP-NTKGYMLYDLHSHNIITSRNVVFYE 606
>CO981879
Length = 576
Score = 71.6 bits (174), Expect(2) = 3e-19
Identities = 36/93 (38%), Positives = 54/93 (57%), Gaps = 3/93 (3%)
Frame = -1
Query: 551 QTFILYSQRQYGSLVKIVRLDNGVEFA---MTDFYSQHGVQHQLSCVKTPQQNGVVERKH 607
+TF Q Q+ +K+ R DNG E+ ++ ++G+ HQ SCV TPQQNGV ERK+
Sbjct: 564 KTFFQMIQTQFQVKIKVFRSDNGREYFNKHLSKXXLENGIIHQSSCVDTPQQNGVAERKN 385
Query: 608 QHILNIARALMLQSNLTLEFWSFAVIHAVCLMN 640
+H+ +ARAL+ Q+ W A++ L N
Sbjct: 384 RHLXEVARALLFQNKAPKYXWGEAILTGTYLKN 286
Score = 42.4 bits (98), Expect(2) = 3e-19
Identities = 24/89 (26%), Positives = 47/89 (51%), Gaps = 5/89 (5%)
Frame = -2
Query: 642 LPTKVLSYSSPFIILHNLKPDIE-----HLKVFGSLCYATSLTAHRTKFPLRARKCIVLG 696
+P+K+L++ +P + + P+ LK+FG + ++ K RA+KC+ +G
Sbjct: 281 MPSKILNFRTPLDVFTSAFPNNRLSCTLPLKIFGCTVFVHIHEPNQGKLEPRAKKCVFVG 102
Query: 697 QKEGGVKGYLMFDLKTREVFLSRNVIFYE 725
KGY FD +++ F++ +V F+E
Sbjct: 101 YAPNQ-KGYKCFDPTSKKTFVTIDVTFFE 18
>BI427153
Length = 422
Score = 90.5 bits (223), Expect = 2e-18
Identities = 53/138 (38%), Positives = 72/138 (51%), Gaps = 1/138 (0%)
Frame = +1
Query: 589 HQLSCVKTPQQNGVVERKHQHILNIARALMLQSNLTLEFWSFAVIHAVCLMNLLPTKVLS 648
HQ +C TPQQNG+ ERK+ H+L AR+LML SN+ W AV+ A L+N +P+ L
Sbjct: 1 HQSTCPHTPQQNGIAERKNHHLLETARSLMLNSNVPTHHWGDAVLTACFLINRMPSSSLE 180
Query: 649 YSSPF-IILHNLKPDIEHLKVFGSLCYATSLTAHRTKFPLRARKCIVLGQKEGGVKGYLM 707
P I+ N KVFG C+ L+ K R+ KC+ LG KGY
Sbjct: 181 NQIPHSIVFPNDLLFYVSPKVFGCTCFVHDLSPGLDKLSARSVKCVFLGYSR-LQKGYTC 357
Query: 708 FDLKTREVFLSRNVIFYE 725
+ R ++S NV F+E
Sbjct: 358 YFPNMRRYYMSANVTFFE 411
>TC222253 similar to UP|Q9ZQE4 (Q9ZQE4) Copia-like retroelement pol
polyprotein, partial (4%)
Length = 919
Score = 90.1 bits (222), Expect = 3e-18
Identities = 49/143 (34%), Positives = 84/143 (58%), Gaps = 2/143 (1%)
Frame = +1
Query: 478 CDTCHFAKQKRLSFSL-RNTIYGTVLELLHVDIWGPYSVASITGAKYFHIIVYDKSRFTW 536
CDTC K+ R SF ++ +L+++H+D+ + + YF + D S+ W
Sbjct: 331 CDTCEIGKKHRESFPTGKSWRMKKLLKIVHLDLC-TVEIPTHGDNNYFITFIDDFSKKMW 507
Query: 537 IKLMKAKSEARAALQTFILYSQRQYGSLVKIVRLDNGVEF-AMTDFYSQHGVQHQLSCVK 595
+ +K KSEA A + F ++++Q G VK + +D G E+ + T F+ +HG+QHQL+
Sbjct: 508 VYFLKQKSEACNAFKMFKAFAEKQNGCKVKALIIDKGQEYLSYTIFFEKHGIQHQLTTKY 687
Query: 596 TPQQNGVVERKHQHILNIARALM 618
TPQ NGV ERK++ I+++ R ++
Sbjct: 688 TPQHNGVTERKNKTIMDMVRCML 756
>CO983154
Length = 568
Score = 83.6 bits (205), Expect = 3e-16
Identities = 53/155 (34%), Positives = 79/155 (50%), Gaps = 4/155 (2%)
Frame = +3
Query: 596 TPQQNGVVERKHQHILNIARALMLQSNLTLEFWSFAVIHAVCLMNLLPTKVLSYSSPFII 655
TPQQNG+ ERK++H+L AR+LML N+ + W AV+ + L+N +P+ L P +
Sbjct: 6 TPQQNGIAERKNRHLLETARSLMLNLNVPIHHWGDAVLTSCFLINRMPSSSLENQIPHSL 185
Query: 656 LHNLKPDIEHL--KVFGSLCYATSLTAHRTKFPLRARKCIVLGQKEGGVKGYLMFDLKTR 713
+ P + H+ KVFG C+ L+ K R+ KC+ LG KGY + R
Sbjct: 186 VFPHDP-LFHVSPKVFGCTCFVHDLSPGLDKLSARSVKCVFLGYSR-LQKGYKCYSPTMR 359
Query: 714 EVFLSRNVIFYE--TIFPFQATDKVSLSSLFPINS 746
++S +V F+E F SL + PI S
Sbjct: 360 RYYMSADVTFFEDTPFFSPSVDHSSSLQEVLPIPS 464
>CO981347
Length = 624
Score = 58.2 bits (139), Expect(3) = 1e-14
Identities = 31/96 (32%), Positives = 50/96 (51%), Gaps = 3/96 (3%)
Frame = +2
Query: 560 QYGSLVKIVRLDNGVEFAM---TDFYSQHGVQHQLSCVKTPQQNGVVERKHQHILNIARA 616
Q G+ +K++R DNG+EF + +F + G++ TP QNG+ ER + IL R
Sbjct: 146 QLGTKLKVLRTDNGLEFVLEQFNEFCRKIGIKRHKIVPHTP*QNGLAERMNMTILERVRC 325
Query: 617 LMLQSNLTLEFWSFAVIHAVCLMNLLPTKVLSYSSP 652
++L + L FW A L+N P+ L + +P
Sbjct: 326 MLLSARLPKTFWGEAANTTSYLINRCPSSTLGFKTP 433
Score = 30.8 bits (68), Expect(3) = 1e-14
Identities = 20/54 (37%), Positives = 29/54 (53%)
Frame = +3
Query: 661 PDIEHLKVFGSLCYATSLTAHRTKFPLRARKCIVLGQKEGGVKGYLMFDLKTRE 714
P+ LKVFGSL + + K RA KC+ +G + GVK Y ++ L+ E
Sbjct: 459 PNYSGLKVFGSLAFD---HVKQGKLDARAVKCVFIGYPK-GVKRYKLWKLEPGE 608
Score = 29.3 bits (64), Expect(3) = 1e-14
Identities = 14/35 (40%), Positives = 21/35 (60%)
Frame = +3
Query: 512 PYSVASITGAKYFHIIVYDKSRFTWIKLMKAKSEA 546
P V + G+ YF I+ D SR W+ ++K KSE+
Sbjct: 3 PSRVKTHGGSSYFLTIIDDFSRRVWLYVLKNKSES 107
>CO982898
Length = 816
Score = 50.8 bits (120), Expect(2) = 3e-12
Identities = 31/74 (41%), Positives = 44/74 (58%), Gaps = 1/74 (1%)
Frame = -3
Query: 571 DNGVEF-AMTDFYSQHGVQHQLSCVKTPQQNGVVERKHQHILNIARALMLQSNLTLEFWS 629
D G EF +T+F + G+ H+L T QN VVERK +HI+ +L+ ++L L+FW
Sbjct: 814 DWGGEF*PLTEFLND*GIIHKLIFPHTQYQN-VVERKQRHIVKFGLSLLSHASLPLKFWG 638
Query: 630 FAVIHAVCLMNLLP 643
A I V L+N LP
Sbjct: 637 HAFITPVFLINRLP 596
Score = 39.3 bits (90), Expect(2) = 3e-12
Identities = 28/102 (27%), Positives = 46/102 (44%)
Frame = -1
Query: 643 PTKVLSYSSPFIILHNLKPDIEHLKVFGSLCYATSLTAHRTKFPLRARKCIVLGQKEGGV 702
PT ++++ P+ L N D +VFG C+ + KF ++ +CI LG
Sbjct: 597 PTVSINFAVPYTQLFNQMLDYSFPRVFGCACFPLIRPYNIQKFNFKSHECIFLGNYTSH- 421
Query: 703 KGYLMFDLKTREVFLSRNVIFYETIFPFQATDKVSLSSLFPI 744
K Y L +S++VIF E FP++ S+ P+
Sbjct: 420 KNYKC--LAPAGKLVSKDVIFNEI*FPYKELFLPDSSTCIPV 301
>BI784757
Length = 430
Score = 68.6 bits (166), Expect = 1e-11
Identities = 39/128 (30%), Positives = 67/128 (51%), Gaps = 4/128 (3%)
Frame = +1
Query: 469 PITTSPCKNCDTCHFAKQKRLSFSLRNTIYGTV-LELLHVDIWGPYSVASITGAKYFHII 527
P P + CD C KQ R +F I LE+++ D+ GP S+ G +YF
Sbjct: 40 PQIKPPSEVCDGCLQCKQSRSTFKQNVPIRAKEKLEVIYSDVCGPMQTESLGGNRYFISF 219
Query: 528 VYDKSRFTWIKLMKAKSEARAALQTFILYSQRQYGSLVKIVRLDNGVEFAMTDFY---SQ 584
+ + +R W+ L++ KS+ + F +++Q GSL+KI+R + G E+ T+F Q
Sbjct: 220 IDELTRKVWVYLIRRKSDFFEVFEKFKNMAKKQSGSLIKILRTNGGGEYVSTEFQEFCDQ 399
Query: 585 HGVQHQLS 592
G+ H+++
Sbjct: 400 QGIIHEVT 423
>BI701169
Length = 407
Score = 67.0 bits (162), Expect = 3e-11
Identities = 35/98 (35%), Positives = 56/98 (56%), Gaps = 3/98 (3%)
Frame = +2
Query: 524 FHIIVYDKSRFTWIKLMKAKSEARAALQTFILYSQRQYGSLVKIVRLDNGVEFAMTDFYS 583
F + + D SR TW+ K K E + F +++ G +K +R D G EF +F
Sbjct: 113 FFLFIDDFSRKTWVYFFKHKLEVFENFKKFKAIVEKESGFKIKAMRSDRGGEF*SNEFQK 292
Query: 584 ---QHGVQHQLSCVKTPQQNGVVERKHQHILNIARALM 618
HG++ L +++PQQNGV ERK++ ILN+AR+++
Sbjct: 293 YCDDHGIRRPLMVLRSPQQNGVAERKNRTILNMARSML 406
>BE659348 weakly similar to PIR|JC7809|JC78 sulfakinin receptor protein
DSK-R1 - fruit fly (Drosophila melanogaster), partial
(6%)
Length = 770
Score = 63.9 bits (154), Expect = 2e-10
Identities = 58/215 (26%), Positives = 93/215 (42%), Gaps = 6/215 (2%)
Frame = -1
Query: 119 LRGLNDNFAAARS*ILLMEPLPTLSKICSMIIQQERHLGIGVLHEPAVMAVHTGNTQGTH 178
LRG++ +F R +L + +P+L + + +++ + P + N + +
Sbjct: 611 LRGMHPDFDHIRDQVLTSQEVPSLENLITRLLR---------VPSPKIGGNSVDNIETSV 459
Query: 179 SNSNRG----RSNSQYSSSKSNV*RLCTHCGRQNHTVETCFLKHGYPSNFPNGYKPEKAY 234
SNRG R N + + C++C R HT +TC+ HG+P N K E +
Sbjct: 458 MVSNRGGQGGRGNQGGRAGRGGR-P*CSYCKRVGHTQDTCYSIHGFPGKSVNISKSETSE 282
Query: 235 IPTSGGESHSSSDQDEPPSSSLELTREYLQGLLALLPQSKPTAVAQTPNLKPISTSNLVS 294
I + +EYLQ K T +QT S+++S
Sbjct: 281 IKFLEADY-----------------QEYLQ--------LKATKESQT--------SSVIS 201
Query: 295 SHNVIA--NNIGMVNSQWIMDSGATDHIASSLSFF 327
HN A + +G S WI+DSGA+DHIAS+ S F
Sbjct: 200 GHNSTACISQVGNNQSPWIIDSGASDHIASNSSLF 96
>CO980323
Length = 769
Score = 53.1 bits (126), Expect = 4e-07
Identities = 35/88 (39%), Positives = 49/88 (54%)
Frame = -2
Query: 654 IILHNLKPDIEHLKVFGSLCYATSLTAHRTKFPLRARKCIVLGQKEGGVKGYLMFDLKTR 713
++ + L PD E + GS+ + L KF RA KC G K GVKGYL + + ++
Sbjct: 303 LVFYTLYPDTEGFWLTGSVM-SLHLLLPAGKFDGRA*KCAFWGYK-AGVKGYLAYGIHSK 130
Query: 714 EVFLSRNVIFYETIFPFQATDKVSLSSL 741
E+ +SRN FYE IFP + +S SSL
Sbjct: 129 EILISRNAQFYEHIFPCLSL-SISQSSL 49
>TC213920
Length = 428
Score = 53.1 bits (126), Expect = 4e-07
Identities = 25/77 (32%), Positives = 50/77 (64%)
Frame = -3
Query: 590 QLSCVKTPQQNGVVERKHQHILNIARALMLQSNLTLEFWSFAVIHAVCLMNLLPTKVLSY 649
Q + + TPQQNGV E++++ ++++ R++++ L++ W + + A+ L+N +P+KV+
Sbjct: 345 QYTILGTPQQNGVSEKRNRTLMDMVRSMLIN*TLSISLWMYTLKTAM*LLNRVPSKVVP- 169
Query: 650 SSPFIILHNLKPDIEHL 666
+PF + N P I HL
Sbjct: 168 KTPFEL*TNRIPSIRHL 118
>CO983960
Length = 875
Score = 38.1 bits (87), Expect(2) = 5e-07
Identities = 16/28 (57%), Positives = 18/28 (64%)
Frame = +2
Query: 113 DCVLCFLRGLNDNFAAARS*ILLMEPLP 140
DCVLCFL GLNDN+ I L +P P
Sbjct: 488 DCVLCFLHGLNDNYVIVPFMIFLFKPFP 571
Score = 34.3 bits (77), Expect(2) = 5e-07
Identities = 16/32 (50%), Positives = 21/32 (65%)
Frame = +3
Query: 140 PTLSKICSMIIQQERHLGIGVLHEPAVMAVHT 171
P L K SMI+ QER L IG+ +PA +A+ T
Sbjct: 564 PFLFKGLSMIVHQERQLNIGMCPDPAALAIQT 659
>BG359773
Length = 382
Score = 47.4 bits (111), Expect = 2e-05
Identities = 23/87 (26%), Positives = 50/87 (57%)
Frame = -1
Query: 584 QHGVQHQLSCVKTPQQNGVVERKHQHILNIARALMLQSNLTLEFWSFAVIHAVCLMNLLP 643
+ G+ Q + + PQ NGV E++++ ++++ ++++ L + W +A+ + L+N +P
Sbjct: 304 KRGICAQYTMLGIPQ*NGVSEKRNRILMDMVTSMLINLTLPISLWMYALKIVMYLLNRVP 125
Query: 644 TKVLSYSSPFIILHNLKPDIEHLKVFG 670
+K + + F + N P I HL V+G
Sbjct: 124 SKAVP-KTHFELWTNRTPSIRHLDVWG 47
>BI425191 weakly similar to GP|14586969|gb| pol polyprotein {Citrus x
paradisi}, partial (7%)
Length = 423
Score = 46.6 bits (109), Expect = 4e-05
Identities = 25/75 (33%), Positives = 40/75 (53%)
Frame = +3
Query: 501 VLELLHVDIWGPYSVASITGAKYFHIIVYDKSRFTWIKLMKAKSEARAALQTFILYSQRQ 560
+LE++H DI P+ V S +YF + D S + ++ L+ K +A L+ + +RQ
Sbjct: 99 LLEIMHTDICRPFDVNSFGKERYFITFIDDYSHYGYVYLLHEKFQAVDVLEIHLNEVERQ 278
Query: 561 YGSLVKIVRLDNGVE 575
VK+VR D G E
Sbjct: 279 LDRKVKVVRSDRGGE 323
>BQ296988 similar to GP|21740616|em OSJNBb0089K24.12 {Oryza sativa (japonica
cultivar-group)}, partial (1%)
Length = 408
Score = 46.2 bits (108), Expect = 5e-05
Identities = 20/46 (43%), Positives = 31/46 (66%)
Frame = -1
Query: 96 HACCCEVVKIFKQYRENDCVLCFLRGLNDNFAAARS*ILLMEPLPT 141
H C C+ +++ +++ V+ FL GLND F A +S ILL+EPLP+
Sbjct: 138 HRCSCDAMRLARRHHHTLHVMRFLTGLNDEFNAVKSQILLIEPLPS 1
>BM892610
Length = 421
Score = 45.4 bits (106), Expect = 9e-05
Identities = 29/95 (30%), Positives = 48/95 (50%)
Frame = -3
Query: 636 VCLMNLLPTKVLSYSSPFIILHNLKPDIEHLKVFGSLCYATSLTAHRTKFPLRARKCIVL 695
V L+N LPT L + P + + L P+ L+ FG CY ++ K R+ +C+ L
Sbjct: 416 VYLINRLPTTSLKFQVPCTVFNKL-PNYNFLRNFGCSCYPFLRPYNKHKLDFRSHECLFL 240
Query: 696 GQKEGGVKGYLMFDLKTREVFLSRNVIFYETIFPF 730
G KGY ++++S++VIF E +P+
Sbjct: 239 GYSTPH-KGYKCLS-PNGKLYVSKDVIFNELKYPY 141
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.337 0.145 0.465
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,236,044
Number of Sequences: 63676
Number of extensions: 644345
Number of successful extensions: 6122
Number of sequences better than 10.0: 74
Number of HSP's better than 10.0 without gapping: 6002
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 6098
length of query: 754
length of database: 12,639,632
effective HSP length: 104
effective length of query: 650
effective length of database: 6,017,328
effective search space: 3911263200
effective search space used: 3911263200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0296a.5


