

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0290.7
(47 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
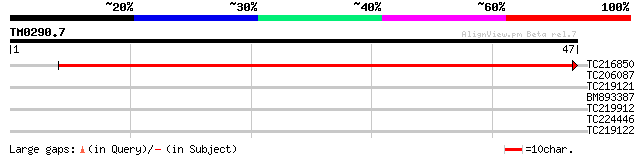
TC216850 homologue to UP|Q42793 (Q42793) Acetyl CoA carboxylase ... 77 2e-15
TC206087 similar to UP|Q38931 (Q38931) Rof1, partial (50%) 27 2.3
TC219121 similar to UP|Q9M9U3 (Q9M9U3) F6A14.17 protein (At1g187... 27 2.3
BM893387 weakly similar to PIR|A86321|A86 F6A14.17 protein - Ara... 27 2.3
TC219912 similar to UP|Q7XZU1 (Q7XZU1) SAC domain protein 4, par... 25 6.7
TC224446 25 8.7
TC219122 similar to UP|Q9M9U3 (Q9M9U3) F6A14.17 protein (At1g187... 25 8.7
>TC216850 homologue to UP|Q42793 (Q42793) Acetyl CoA carboxylase , complete
Length = 7100
Score = 77.0 bits (188), Expect = 2e-15
Identities = 35/43 (81%), Positives = 37/43 (85%)
Frame = +1
Query: 5 NPDYGFNPTGWEVQELGFKSKPNVWAYFSVKSGGGIHEFLDSQ 47
+PD GF PT +VQEL FKSKPNVWAYFSVKSGGGIHEF DSQ
Sbjct: 1321 DPDDGFKPTSGKVQELNFKSKPNVWAYFSVKSGGGIHEFSDSQ 1449
>TC206087 similar to UP|Q38931 (Q38931) Rof1, partial (50%)
Length = 1183
Score = 26.9 bits (58), Expect = 2.3
Identities = 15/35 (42%), Positives = 20/35 (56%)
Frame = +3
Query: 1 VLIINPDYGFNPTGWEVQELGFKSKPNVWAYFSVK 35
+LII P+Y F P+G QEL PN Y+ V+
Sbjct: 237 LLIIQPEYAFGPSG-SSQELA-NVPPNSTVYYEVE 335
>TC219121 similar to UP|Q9M9U3 (Q9M9U3) F6A14.17 protein
(At1g18720/F6A14_17), partial (82%)
Length = 856
Score = 26.9 bits (58), Expect = 2.3
Identities = 13/37 (35%), Positives = 17/37 (45%), Gaps = 1/37 (2%)
Frame = -1
Query: 2 LIINPDYGFNPTGWEVQELGFKSK-PNVWAYFSVKSG 37
L++NP YG PT W K WA S ++G
Sbjct: 652 LVLNPGYGSYPTNWRTSSNTKKGAIRKAWARLSNRAG 542
>BM893387 weakly similar to PIR|A86321|A86 F6A14.17 protein - Arabidopsis
thaliana, partial (50%)
Length = 399
Score = 26.9 bits (58), Expect = 2.3
Identities = 13/37 (35%), Positives = 17/37 (45%), Gaps = 1/37 (2%)
Frame = -3
Query: 2 LIINPDYGFNPTGWEVQELGFKSK-PNVWAYFSVKSG 37
L++NP YG PT W K WA S ++G
Sbjct: 397 LVLNPGYGSYPTNWRTSSNTKKGAIRKAWARLSNRAG 287
>TC219912 similar to UP|Q7XZU1 (Q7XZU1) SAC domain protein 4, partial (24%)
Length = 823
Score = 25.4 bits (54), Expect = 6.7
Identities = 9/21 (42%), Positives = 13/21 (61%)
Frame = +3
Query: 8 YGFNPTGWEVQELGFKSKPNV 28
YG G+++Q LGF PN+
Sbjct: 702 YGLAALGYQLQALGFTETPNI 764
>TC224446
Length = 454
Score = 25.0 bits (53), Expect = 8.7
Identities = 13/33 (39%), Positives = 18/33 (54%)
Frame = +1
Query: 1 VLIINPDYGFNPTGWEVQELGFKSKPNVWAYFS 33
V+I + +GF T W L F P+ +AYFS
Sbjct: 22 VIIFSLFFGFLFTLWLSGSLSFSLPPSKFAYFS 120
>TC219122 similar to UP|Q9M9U3 (Q9M9U3) F6A14.17 protein
(At1g18720/F6A14_17), partial (39%)
Length = 577
Score = 25.0 bits (53), Expect = 8.7
Identities = 8/14 (57%), Positives = 10/14 (71%)
Frame = -3
Query: 2 LIINPDYGFNPTGW 15
L++NP YG PT W
Sbjct: 347 LVLNPGYGSYPTNW 306
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.142 0.463
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,488,415
Number of Sequences: 63676
Number of extensions: 23891
Number of successful extensions: 109
Number of sequences better than 10.0: 15
Number of HSP's better than 10.0 without gapping: 109
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 109
length of query: 47
length of database: 12,639,632
effective HSP length: 23
effective length of query: 24
effective length of database: 11,175,084
effective search space: 268202016
effective search space used: 268202016
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0290.7


