

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0288.8
(846 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
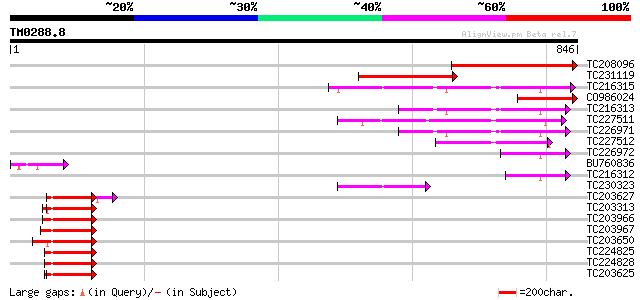
Score E
Sequences producing significant alignments: (bits) Value
TC208096 weakly similar to UP|Q9SU29 (Q9SU29) Polyubiquitin-like... 359 2e-99
TC231119 similar to UP|Q9SU29 (Q9SU29) Polyubiquitin-like protei... 268 6e-72
TC216315 similar to GB|AAF36454.1|7108521|AF127564 ubiquitin-pro... 183 2e-46
CO986024 166 4e-41
TC216313 similar to GB|AAF36454.1|7108521|AF127564 ubiquitin-pro... 137 3e-32
TC227511 similar to GB|AAQ89615.1|37202000|BT010593 At3g17205 {A... 134 2e-31
TC226971 homologue to GB|AAF36454.1|7108521|AF127564 ubiquitin-p... 131 1e-30
TC227512 similar to GB|AAQ89615.1|37202000|BT010593 At3g17205 {A... 84 3e-16
TC226972 similar to GB|AAF36454.1|7108521|AF127564 ubiquitin-pro... 81 2e-15
BU760836 80 4e-15
TC216312 similar to GB|AAF36454.1|7108521|AF127564 ubiquitin-pro... 75 2e-13
TC230323 homologue to GB|AAF36454.1|7108521|AF127564 ubiquitin-p... 72 1e-12
TC203627 UP|Q41067 (Q41067) Polyubiquitin, partial (47%) 67 3e-11
TC203313 PIR|S01425|UQMUM polyubiquitin 5 - Arabidopsis thaliana... 66 7e-11
TC203966 homologue to UP|Q6UCJ3 (Q6UCJ3) Ubiquitin extension pro... 65 2e-10
TC203967 homologue to UP|Q6UCJ3 (Q6UCJ3) Ubiquitin extension pro... 65 2e-10
TC203650 GB|AAA34124.1|170354|TOBUBI4A pentameric polyubiquitin ... 64 4e-10
TC224825 UP|Q9AR09 (Q9AR09) Ubiquitin fused to ribosomal protein... 63 6e-10
TC224828 UP|Q9AR09 (Q9AR09) Ubiquitin fused to ribosomal protein... 63 6e-10
TC203625 UP|Q39257 (Q39257) Ubiquitin, partial (83%) 62 8e-10
>TC208096 weakly similar to UP|Q9SU29 (Q9SU29) Polyubiquitin-like protein,
partial (19%)
Length = 789
Score = 359 bits (922), Expect = 2e-99
Identities = 167/188 (88%), Positives = 181/188 (95%)
Frame = +2
Query: 659 LCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLE 718
LCPGGKNL+VNSKNR+KY+DLLI+DRFVTSISEQVSHF KGF DILS+SK QQ+FFQSL+
Sbjct: 2 LCPGGKNLVVNSKNRDKYVDLLIQDRFVTSISEQVSHFVKGFADILSNSKLQQYFFQSLD 181
Query: 719 LEDLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTS 778
LEDLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQI+WFWEIVGRMTA+QRKVLLFFWTS
Sbjct: 182 LEDLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQISWFWEIVGRMTADQRKVLLFFWTS 361
Query: 779 VKYLPVEGFCGLASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQEH 838
VKYLPVEGF GLASRLYIY+S EP DRLPSSHTCF+R+CFPAYSSM VM++RLEVITQEH
Sbjct: 362 VKYLPVEGFRGLASRLYIYRSLEPGDRLPSSHTCFFRLCFPAYSSMAVMKDRLEVITQEH 541
Query: 839 IGCSFGTW 846
IGCSFGTW
Sbjct: 542 IGCSFGTW 565
>TC231119 similar to UP|Q9SU29 (Q9SU29) Polyubiquitin-like protein, partial
(16%)
Length = 442
Score = 268 bits (686), Expect = 6e-72
Identities = 126/147 (85%), Positives = 139/147 (93%)
Frame = +2
Query: 521 EEATGPGVLREWFLLVCQALFNPQNALFVACPNDRRRFYPNHASKVHPLHLEYFGFSGRV 580
EEATGPGVLREWFLLVCQA+FNPQNALFVACPNDRRRF+PN ASKVHPLHLEYF F+GRV
Sbjct: 2 EEATGPGVLREWFLLVCQAIFNPQNALFVACPNDRRRFFPNPASKVHPLHLEYFSFAGRV 181
Query: 581 IALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDAL 640
IALALMH+VQVGIV DRVFFLQLAG +I +ED++DADP+LY+SCKQIL+MD+DFIDSDAL
Sbjct: 182 IALALMHRVQVGIVFDRVFFLQLAGNYIAIEDIRDADPYLYTSCKQILDMDADFIDSDAL 361
Query: 641 GLTFVREVEELGDTKVVELCPGGKNLI 667
GLTFVREVEELG KVVELCP GKNL+
Sbjct: 362 GLTFVREVEELGQRKVVELCPXGKNLV 442
>TC216315 similar to GB|AAF36454.1|7108521|AF127564 ubiquitin-protein ligase 1
{Arabidopsis thaliana;} , partial (12%)
Length = 1938
Score = 183 bits (465), Expect = 2e-46
Identities = 124/386 (32%), Positives = 195/386 (50%), Gaps = 17/386 (4%)
Frame = +1
Query: 476 EVKEDYEELHEML---IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPGVL-RE 531
++K ++ H L + R+ +L +S+ + T L L + F+ EE G L RE
Sbjct: 634 KIKHQHDHHHSPLRISVRRAYVLEDSYNQLRMRSTQDLKGRLTVHFQGEEGIDAGGLTRE 813
Query: 532 WFLLVCQALFNPQNALFVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQV 591
W+ L+ + +F+ LF N+ F PN S HL YF F GRV+ AL +
Sbjct: 814 WYQLLSRVIFDKGALLFTTVGNEST-FQPNPNSVYQTEHLSYFKFVGRVVGKALFDGQLL 990
Query: 592 GIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEE- 650
+ R F+ + G +T D++ DP + + K +LE D S+ L LTF + +E
Sbjct: 991 DVHFTRSFYKHVLGAKVTYHDIEAIDPDYFRNLKWMLENDI----SEILDLTFSIDADEE 1158
Query: 651 ----LGDTKVV--ELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDIL 704
T+V EL PGG+N V +N+ +Y+DL+ + R T+I Q++ F +GF +++
Sbjct: 1159 KLILYERTEVTDYELIPGGRNTKVTEENKHQYVDLVAEHRLTTAIRPQINAFLEGFNELI 1338
Query: 705 SSSKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQITWFWEIVGRM 764
F + LEL ++ G + I ++D +A+TEY+GY I WFWE+V
Sbjct: 1339 PRELISIFNDKELEL-----LISGLPE-IDLDDLRANTEYSGYSGASPVIQWFWEVVQGF 1500
Query: 765 TAEQRKVLLFFWTSVKYLPVEGFCGL-----ASRLYIYKSPEPEDRLPSSHTCFYRMCFP 819
+ E + LL F T +P+EGF L A R I+K+ D LPS+HTCF ++ P
Sbjct: 1501 SKEDKARLLQFVTGTSKVPLEGFSALQGISGAQRFQIHKAYGSSDHLPSAHTCFNQLDLP 1680
Query: 820 AYSSMTVMQERLEV-ITQEHIGCSFG 844
Y S ++ERL + I + + G FG
Sbjct: 1681 EYPSKQHLEERLLLAIHEANEGFGFG 1758
>CO986024
Length = 791
Score = 166 bits (420), Expect = 4e-41
Identities = 76/89 (85%), Positives = 84/89 (93%)
Frame = -2
Query: 758 WEIVGRMTAEQRKVLLFFWTSVKYLPVEGFCGLASRLYIYKSPEPEDRLPSSHTCFYRMC 817
W+IV RMTA+QRKVLLFFWTSVKYLPVEGF GLASRLYIY+S EP DRLPSSHTCF+R+C
Sbjct: 424 WQIVERMTADQRKVLLFFWTSVKYLPVEGFRGLASRLYIYRSLEPGDRLPSSHTCFFRLC 245
Query: 818 FPAYSSMTVMQERLEVITQEHIGCSFGTW 846
FPAYSS+ VM++RLEVITQEHIGCSFGTW
Sbjct: 244 FPAYSSIAVMKDRLEVITQEHIGCSFGTW 158
>TC216313 similar to GB|AAF36454.1|7108521|AF127564 ubiquitin-protein ligase
1 {Arabidopsis thaliana;} , partial (7%)
Length = 785
Score = 137 bits (344), Expect = 3e-32
Identities = 86/269 (31%), Positives = 141/269 (51%), Gaps = 12/269 (4%)
Frame = +1
Query: 581 IALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDAL 640
+ AL + + R F+ + G +T D++ DP + + K +LE D SD L
Sbjct: 1 VGKALFDGQLLDVHFTRSFYKHILGVKVTYHDIEAIDPDYFKNLKWMLENDI----SDVL 168
Query: 641 GLTFVREVEE-----LGDTKVV--ELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQV 693
LTF + +E T+V EL PGG+N+ V +N+ +Y+DL+ + R T+I Q+
Sbjct: 169 DLTFSIDADEEKLILYERTEVTDYELIPGGRNIKVTEENKHQYVDLVAEHRLTTAIRPQI 348
Query: 694 SHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQ 753
++F +GF +++ F + LEL ++ G D I ++D +A+TEY+GY
Sbjct: 349 NYFLEGFNEMIPRELISIFNDKELEL-----LISGLPD-IDLDDLRANTEYSGYSAASPV 510
Query: 754 ITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGFCGL-----ASRLYIYKSPEPEDRLPS 808
I WFWE+V ++ E + LL F T +P+EGF L + + I+K+ + LPS
Sbjct: 511 IQWFWEVVQGLSKEDKARLLQFVTGTSKVPLEGFSALQGISGSQKFQIHKA*GSPEHLPS 690
Query: 809 SHTCFYRMCFPAYSSMTVMQERLEVITQE 837
+HTCF ++ P Y S ++ERL + E
Sbjct: 691 AHTCFNQLDLPEYPSKHHLEERLLLAIHE 777
>TC227511 similar to GB|AAQ89615.1|37202000|BT010593 At3g17205 {Arabidopsis
thaliana;} , partial (44%)
Length = 1712
Score = 134 bits (337), Expect = 2e-31
Identities = 101/356 (28%), Positives = 162/356 (45%), Gaps = 13/356 (3%)
Frame = +2
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEAT------GPGVLREWFLLVCQALFN 542
I R +L +++ +++ SL + + F NE G G+ +++ + +A F+
Sbjct: 389 IKRDHILEDAYNQMSQLTEDSLRGSIRVTFVNEFGVEEAGIDGGGIFKDFMENITRAAFD 568
Query: 543 PQNALFVACPNDRRRFYPNHAS-KVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFL 601
Q LF + Y N S +H H ++F F G ++A A+ + V I F
Sbjct: 569 VQYGLFKETAD--HLLYANPGSGMIHEQHFQFFHFLGTLLAKAMFEGILVDIPFATFFLS 742
Query: 602 QLAGKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDALGLTFVREVEELGDTKVVELCP 661
+L KH L D+ DP LY + ++ + D L L FV E G+ EL P
Sbjct: 743 KLKQKHNYLNDLPSLDPELY---RHLIFLKHYKGDISELELYFVIVNNEYGEQTEEELLP 913
Query: 662 GGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSHFSKGFTDILSSSKFQQFFFQSLELED 721
GG+NL V ++N +I L+ R I +Q SHF +GF ++ F L+L
Sbjct: 914 GGRNLRVTNENVITFIHLVANHRLNFQIRQQSSHFLRGFQQLMQKDWIDMFNEHELQL-- 1087
Query: 722 LDWMLHGSEDTISVEDWKAHTEY-NGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVK 780
++ GS D++ ++D + HT Y GY + FWE++ + E RK L F T
Sbjct: 1088---LISGSLDSLDIDDLRLHTNYAGGYHGEHYVMEMFWEVLKGFSLENRKKFLKFVTGCS 1258
Query: 781 YLPVEGFCGLASRLYIYK-----SPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERL 831
P+ GF L I + + E DRLP+S TC + P Y+S ++ +L
Sbjct: 1259RGPLLGFRYLEPMFCIQRASGNAAEESLDRLPTSATCMNLLKLPPYTSKEQLETKL 1426
>TC226971 homologue to GB|AAF36454.1|7108521|AF127564 ubiquitin-protein
ligase 1 {Arabidopsis thaliana;} , partial (7%)
Length = 1359
Score = 131 bits (329), Expect = 1e-30
Identities = 84/269 (31%), Positives = 131/269 (48%), Gaps = 12/269 (4%)
Frame = +1
Query: 581 IALALMHKVQVGIVLDRVFFLQLAGKHITLEDVKDADPFLYSSCKQILEMDSDFIDSDAL 640
+ AL + + R F+ + G +T D++ DP Y + K +LE D SD
Sbjct: 151 VGKALFDGQLLDVYFTRSFYKHILGVKVTYHDIEAVDPDYYKNLKWMLENDV----SDIP 318
Query: 641 GLTFVREVEE-------LGDTKVVELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQV 693
LTF + +E + EL PGG+N+ V + + +Y+DL+ + +I Q+
Sbjct: 319 DLTFSMDADEEKHILYEKNEVTDYELKPGGRNIRVTEETKHEYVDLVAEHLLTNAIRPQI 498
Query: 694 SHFSKGFTDILSSSKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEYNGYKETDIQ 753
+ F +GF +++ F + LEL ++ G + I ++D KA+TEY GY
Sbjct: 499 NSFLEGFNELVPRELISIFNDKELEL-----LISGLPE-IDLDDLKANTEYTGYTVASNV 660
Query: 754 ITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGFCGL-----ASRLYIYKSPEPEDRLPS 808
+ WFWE+V E LL F T +P+EGF L R I+K+ DRLPS
Sbjct: 661 VQWFWEVVKAFNKEDMARLLQFVTGTSKVPLEGFKALQGISGPQRFQIHKAYGAPDRLPS 840
Query: 809 SHTCFYRMCFPAYSSMTVMQERLEVITQE 837
+HTCF ++ P Y+S +QERL + E
Sbjct: 841 AHTCFNQLDLPEYTSKEQLQERLLLAIHE 927
>TC227512 similar to GB|AAQ89615.1|37202000|BT010593 At3g17205 {Arabidopsis
thaliana;} , partial (17%)
Length = 541
Score = 84.0 bits (206), Expect = 3e-16
Identities = 61/180 (33%), Positives = 86/180 (46%), Gaps = 6/180 (3%)
Frame = +2
Query: 636 DSDALGLTFVREVEELGDTKVVELCPGGKNLIVNSKNREKYIDLLIKDRFVTSISEQVSH 695
D L L FV E G+ EL PGGKNL V ++N +I L+ R I +Q SH
Sbjct: 17 DISELELYFVIVNNEYGEQTEEELLPGGKNLRVTNENVITFIHLVANHRLNFQIRQQSSH 196
Query: 696 FSKGFTDILSSSKFQQFFFQSLELEDLDWMLHGSEDTISVEDWKAHTEY-NGYKETDIQI 754
F +GF ++ F L+L ++ GS D++ V+D + HT Y GY I
Sbjct: 197 FLRGFQQLIQKDWIDMFNEHELQL-----LISGSLDSLDVDDLRQHTNYAGGYHSDHHVI 361
Query: 755 TWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGFCGLASRLYIYK--SPEPE---DRLPSS 809
FWE++ + E +K L F T P+ GF L I + S +P+ DRLP+S
Sbjct: 362 EMFWEVLKGFSLENKKKFLKFVTGCSRGPLLGFQYLEPLFCIQRAGSNDPDEALDRLPTS 541
>TC226972 similar to GB|AAF36454.1|7108521|AF127564 ubiquitin-protein ligase
1 {Arabidopsis thaliana;} , partial (3%)
Length = 957
Score = 81.3 bits (199), Expect = 2e-15
Identities = 43/110 (39%), Positives = 59/110 (53%), Gaps = 5/110 (4%)
Frame = +1
Query: 733 ISVEDWKAHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGFCGL-- 790
I ++D KA+TEY GY + WFWE+V E LL F T +P+EGF L
Sbjct: 37 IDLDDLKANTEYTGYTVASNVVQWFWEVVKAFNKEDMARLLQFVTGTSKVPLEGFKALQG 216
Query: 791 ---ASRLYIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQE 837
R ++K+ DRLPS+HTCF ++ P Y+S +QERL + E
Sbjct: 217 ISGPQRFQVHKAYGAPDRLPSAHTCFNQLDLPEYTSKEQLQERLLLAIHE 366
>BU760836
Length = 444
Score = 80.1 bits (196), Expect = 4e-15
Identities = 52/113 (46%), Positives = 66/113 (58%), Gaps = 26/113 (23%)
Frame = +1
Query: 1 MSVIETPSVHH-------HR---KRKHDEIDDDEGGVSDIDPIRMKKDDA---------- 40
MSVI+TP+VHH HR KRK D+ DD++ SD+ +RM+KD+A
Sbjct: 112 MSVIDTPAVHHRSGGATDHRHPSKRKFDDEDDED--FSDLVCVRMRKDEAKAVNSWSASS 285
Query: 41 ------AKAVYSAVHHRSIMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQA 87
A S + +QFFVRMM GNTIVMQA P+DTV+SIHERIQ+
Sbjct: 286 SSSSSDAGGCSSLQQQQRSHIQFFVRMMSAGNTIVMQAFPEDTVKSIHERIQS 444
>TC216312 similar to GB|AAF36454.1|7108521|AF127564 ubiquitin-protein ligase
1 {Arabidopsis thaliana;} , partial (4%)
Length = 621
Score = 74.7 bits (182), Expect = 2e-13
Identities = 39/103 (37%), Positives = 57/103 (54%), Gaps = 5/103 (4%)
Frame = +2
Query: 740 AHTEYNGYKETDIQITWFWEIVGRMTAEQRKVLLFFWTSVKYLPVEGFCGL-----ASRL 794
A+TEY+GY I WFWE+V ++ E + LL F T +P+EGF L + +
Sbjct: 113 ANTEYSGYSXASPVIQWFWEVVQGLSKEDKARLLQFVTGTSKVPLEGFSALQGISGSQKF 292
Query: 795 YIYKSPEPEDRLPSSHTCFYRMCFPAYSSMTVMQERLEVITQE 837
I+K+ D LPS+HTCF ++ P Y S ++ERL + E
Sbjct: 293 QIHKAYGSPDHLPSAHTCFNQLDLPEYPSKHHLEERLLLAIHE 421
>TC230323 homologue to GB|AAF36454.1|7108521|AF127564 ubiquitin-protein
ligase 1 {Arabidopsis thaliana;} , partial (6%)
Length = 763
Score = 72.0 bits (175), Expect = 1e-12
Identities = 43/141 (30%), Positives = 68/141 (47%), Gaps = 1/141 (0%)
Frame = +1
Query: 489 IDRSQLLSESFEYIARADTASLHAGLFMEFKNEEATGPGVL-REWFLLVCQALFNPQNAL 547
+ R+ +L +S+ + T L L ++F+ EE G L REW+ L+ + +F+ L
Sbjct: 343 VRRAYILEDSYNQLRMRPTQDLKGRLNVQFQGEEGIDAGGLTREWYQLLSRVIFDKGALL 522
Query: 548 FVACPNDRRRFYPNHASKVHPLHLEYFGFSGRVIALALMHKVQVGIVLDRVFFLQLAGKH 607
F N+ F PN S HL YF F GRV+ AL + + R F+ + G
Sbjct: 523 FTTVGNNAT-FQPNPNSVYQTEHLSYFKFVGRVVGKALFDGQLLDVYFTRSFYKHILGVK 699
Query: 608 ITLEDVKDADPFLYSSCKQIL 628
+T D++ DP Y + K +L
Sbjct: 700 VTYHDIEAVDPDYYKNLKWML 762
>TC203627 UP|Q41067 (Q41067) Polyubiquitin, partial (47%)
Length = 1191
Score = 67.4 bits (163), Expect = 3e-11
Identities = 37/119 (31%), Positives = 66/119 (55%), Gaps = 12/119 (10%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 771 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 947
Query: 115 SIGKDASLQLVGRMR------------STGHPHAWQVINDMVSLVIRLCHNEPVHDANK 161
+I K+++L LV R+R HP W+ ++ + ++ ++ E H N+
Sbjct: 948 NIQKESTLHLVLRLRGGMQIFVKDPHWKDHHPLRWRALDTIDNVKAKIQDKEGFHQTNR 1124
Score = 62.4 bits (150), Expect = 8e-10
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 87 MQIFVKTL-TGKTITLEVESSDTIDNVKSKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 263
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 264 NIQKESTLHLVLRLR 308
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 543 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 719
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 720 NIQKESTLHLVLRLR 764
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 315 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 491
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 492 NIQKESTLHLVLRLR 536
>TC203313 PIR|S01425|UQMUM polyubiquitin 5 - Arabidopsis thaliana
{Arabidopsis thaliana;} , complete
Length = 1377
Score = 65.9 bits (159), Expect = 7e-11
Identities = 36/84 (42%), Positives = 56/84 (65%), Gaps = 4/84 (4%)
Frame = +1
Query: 50 HRSIML----QFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQL 105
HRSI+L Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL
Sbjct: 34 HRSILLLLKMQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQL 210
Query: 106 QSEQTLAECSIGKDASLQLVGRMR 129
+ +TLA+ +I K+++L LV R+R
Sbjct: 211 EDGRTLADYNIQKESTLHLVLRLR 282
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +1
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 517 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 693
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 694 NIQKESTLHLVLRLR 738
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +1
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 745 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 921
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 922 NIQKESTLHLVLRLR 966
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +1
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 973 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 1149
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 1150 NIQKESTLHLVLRLR 1194
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +1
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 289 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 465
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 466 NIQKESTLHLVLRLR 510
>TC203966 homologue to UP|Q6UCJ3 (Q6UCJ3) Ubiquitin extension protein,
partial (98%)
Length = 715
Score = 64.7 bits (156), Expect = 2e-10
Identities = 32/81 (39%), Positives = 53/81 (64%)
Frame = +2
Query: 49 HHRSIMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSE 108
H + +Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+
Sbjct: 107 HSPGVTMQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDG 283
Query: 109 QTLAECSIGKDASLQLVGRMR 129
+TLA+ +I K+++L LV R+R
Sbjct: 284 RTLADYNIQKESTLHLVLRLR 346
>TC203967 homologue to UP|Q6UCJ3 (Q6UCJ3) Ubiquitin extension protein,
partial (98%)
Length = 746
Score = 64.7 bits (156), Expect = 2e-10
Identities = 33/83 (39%), Positives = 54/83 (64%)
Frame = +3
Query: 47 AVHHRSIMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQ 106
A H + +Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+
Sbjct: 45 ASHPPCVTMQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLE 221
Query: 107 SEQTLAECSIGKDASLQLVGRMR 129
+TLA+ +I K+++L LV R+R
Sbjct: 222 DGRTLADYNIQKESTLHLVLRLR 290
>TC203650 GB|AAA34124.1|170354|TOBUBI4A pentameric polyubiquitin {Nicotiana
sylvestris;} , partial (23%)
Length = 692
Score = 63.5 bits (153), Expect = 4e-10
Identities = 36/100 (36%), Positives = 61/100 (61%), Gaps = 4/100 (4%)
Frame = +2
Query: 34 RMKKDDAAKAVYSAVHHRSIML----QFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALK 89
+ KK ++ +A + H + L Q FV+ + G TI ++ DT+ ++ +IQ +
Sbjct: 188 KKKKKNSPRAEFGTRLHLVLRLRGGMQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKE 364
Query: 90 GIPVFEQRLIYNGKQLQSEQTLAECSIGKDASLQLVGRMR 129
GIP +QRLI+ GKQL+ +TLA+ +I K+++L LV R+R
Sbjct: 365 GIPPDQQRLIFAGKQLEDGRTLADYNIQKESTLHLVLRLR 484
>TC224825 UP|Q9AR09 (Q9AR09) Ubiquitin fused to ribosomal protein L40,
complete
Length = 651
Score = 62.8 bits (151), Expect = 6e-10
Identities = 31/78 (39%), Positives = 53/78 (67%)
Frame = +3
Query: 52 SIMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTL 111
++ +Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TL
Sbjct: 36 ALKMQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTL 212
Query: 112 AECSIGKDASLQLVGRMR 129
A+ +I K+++L LV R+R
Sbjct: 213 ADYNIQKESTLHLVLRLR 266
>TC224828 UP|Q9AR09 (Q9AR09) Ubiquitin fused to ribosomal protein L40,
complete
Length = 1196
Score = 62.8 bits (151), Expect = 6e-10
Identities = 31/78 (39%), Positives = 53/78 (67%)
Frame = +3
Query: 52 SIMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTL 111
++ +Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TL
Sbjct: 519 ALKMQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTL 695
Query: 112 AECSIGKDASLQLVGRMR 129
A+ +I K+++L LV R+R
Sbjct: 696 ADYNIQKESTLHLVLRLR 749
>TC203625 UP|Q39257 (Q39257) Ubiquitin, partial (83%)
Length = 1427
Score = 62.4 bits (150), Expect = 8e-10
Identities = 31/77 (40%), Positives = 52/77 (67%)
Frame = +3
Query: 53 IMLQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLA 112
+ +Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA
Sbjct: 78 LKMQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLA 254
Query: 113 ECSIGKDASLQLVGRMR 129
+ +I K+++L LV R+R
Sbjct: 255 DYNIQKESTLHLVLRLR 305
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 540 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 716
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 717 NIQKESTLHLVLRLR 761
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 768 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 944
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 945 NIQKESTLHLVLRLR 989
Score = 62.0 bits (149), Expect = 1e-09
Identities = 31/75 (41%), Positives = 51/75 (67%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G TI ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 312 MQIFVKTL-TGKTITLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 488
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 489 NIQKESTLHLVLRLR 533
Score = 61.6 bits (148), Expect = 1e-09
Identities = 30/75 (40%), Positives = 51/75 (68%)
Frame = +3
Query: 55 LQFFVRMMGNGNTIVMQAHPDDTVQSIHERIQALKGIPVFEQRLIYNGKQLQSEQTLAEC 114
+Q FV+ + G T+ ++ DT+ ++ +IQ +GIP +QRLI+ GKQL+ +TLA+
Sbjct: 996 MQIFVKTL-TGKTVTLEVESSDTIDNVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLADY 1172
Query: 115 SIGKDASLQLVGRMR 129
+I K+++L LV R+R
Sbjct: 1173 NIQKESTLHLVLRLR 1217
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.137 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,504,318
Number of Sequences: 63676
Number of extensions: 582032
Number of successful extensions: 2994
Number of sequences better than 10.0: 148
Number of HSP's better than 10.0 without gapping: 2853
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2971
length of query: 846
length of database: 12,639,632
effective HSP length: 105
effective length of query: 741
effective length of database: 5,953,652
effective search space: 4411656132
effective search space used: 4411656132
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0288.8


