

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0284.4
(247 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
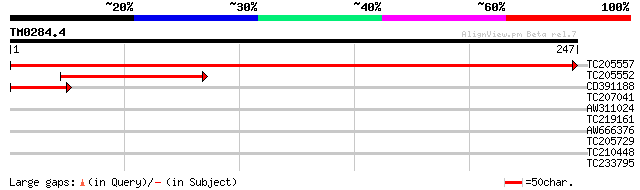
Score E
Sequences producing significant alignments: (bits) Value
TC205557 similar to UP|PMM_ARATH (O80840) Probable phosphomannom... 473 e-134
TC205552 similar to UP|PMM_ARATH (O80840) Probable phosphomannom... 112 2e-25
CD391188 44 9e-05
TC207041 29 1.7
AW311024 29 1.7
TC219161 homologue to GB|AAM10488.1|29465725|AY089970 uracil pho... 28 3.0
AW666376 28 5.0
TC205729 similar to UP|Q8GZV0 (Q8GZV0) Obtusifoliol-14-demethyla... 28 5.0
TC210448 similar to UP|CHI1_TULBA (Q9SLP4) Chitinase 1 precursor... 27 8.6
TC233795 27 8.6
>TC205557 similar to UP|PMM_ARATH (O80840) Probable phosphomannomutase (PMM)
, complete
Length = 1699
Score = 473 bits (1216), Expect = e-134
Identities = 229/247 (92%), Positives = 240/247 (96%)
Frame = +3
Query: 1 MAVAKPGVIALFDVDGTLTAPRKVANSEMLGFMQELRKVVTVGVVGGSDLIKISEQLGST 60
MA +PG+IALFDVDGTLTAPRKV EML FMQELRKVVTVGVVGGSDLIKISEQLGST
Sbjct: 75 MAARRPGLIALFDVDGTLTAPRKVVTPEMLTFMQELRKVVTVGVVGGSDLIKISEQLGST 254
Query: 61 VTHDYDYVFSENGLVAHKQGKLIGTESLKTFLGDEKLKEFINFTLHYIADLDIPIKRGTF 120
VT+DYDYVFSENGLVAHK+GKLIGT+SLK+FLG+EKLKEFINFTLHYIADLDIPIKRGTF
Sbjct: 255 VTNDYDYVFSENGLVAHKEGKLIGTQSLKSFLGEEKLKEFINFTLHYIADLDIPIKRGTF 434
Query: 121 IEFRSGMLNVSPIGRNCSQEERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFSIGGQIS 180
IEFRSGMLNVSPIGRNCSQEERDEFEKYDKV NIRPKMVSVLREKFAHLNLTFSIGGQIS
Sbjct: 435 IEFRSGMLNVSPIGRNCSQEERDEFEKYDKVHNIRPKMVSVLREKFAHLNLTFSIGGQIS 614
Query: 181 FDVFPQGWDKTYCLRYLDGFNEIHFFGDKTYKGGNDHEIYESERTIGHTVTSPEDTIKQC 240
FDVFPQGWDKTYCLRYLDGFNEIHFFGDKTYKGGNDHEIYESERT+GHTVTSP+DT+KQC
Sbjct: 615 FDVFPQGWDKTYCLRYLDGFNEIHFFGDKTYKGGNDHEIYESERTVGHTVTSPDDTVKQC 794
Query: 241 KSLFLES 247
KSLFLE+
Sbjct: 795 KSLFLEN 815
>TC205552 similar to UP|PMM_ARATH (O80840) Probable phosphomannomutase (PMM) ,
partial (25%)
Length = 1033
Score = 112 bits (280), Expect = 2e-25
Identities = 56/64 (87%), Positives = 60/64 (93%)
Frame = -3
Query: 23 KVANSEMLGFMQELRKVVTVGVVGGSDLIKISEQLGSTVTHDYDYVFSENGLVAHKQGKL 82
KV EML FMQELRKVVTVGVVGGSDLIKISEQLGSTVT++YDYVFSENGLVAHK+GKL
Sbjct: 1031 KVVTPEMLTFMQELRKVVTVGVVGGSDLIKISEQLGSTVTNEYDYVFSENGLVAHKEGKL 852
Query: 83 IGTE 86
IGT+
Sbjct: 851 IGTQ 840
>CD391188
Length = 621
Score = 43.5 bits (101), Expect = 9e-05
Identities = 20/27 (74%), Positives = 24/27 (88%)
Frame = -3
Query: 1 MAVAKPGVIALFDVDGTLTAPRKVANS 27
MA +PG+IALFDVDGTL APRKV++S
Sbjct: 547 MASRRPGLIALFDVDGTLPAPRKVSDS 467
>TC207041
Length = 868
Score = 29.3 bits (64), Expect = 1.7
Identities = 16/45 (35%), Positives = 20/45 (43%)
Frame = -1
Query: 163 REKFAHLNLTFSIGGQISFDVFPQGWDKTYCLRYLDGFNEIHFFG 207
R KF +LN F + +GWDKTY R N H +G
Sbjct: 544 RPKFKNLNFIF---------LHMKGWDKTYFCRKFQNDNGFHEYG 437
>AW311024
Length = 204
Score = 29.3 bits (64), Expect = 1.7
Identities = 11/14 (78%), Positives = 14/14 (99%)
Frame = -1
Query: 234 EDTIKQCKSLFLES 247
+DT+KQCKSLFLE+
Sbjct: 204 DDTVKQCKSLFLEN 163
>TC219161 homologue to GB|AAM10488.1|29465725|AY089970 uracil
phosphoribosyltransferase {Arabidopsis thaliana;} ,
partial (25%)
Length = 642
Score = 28.5 bits (62), Expect = 3.0
Identities = 11/27 (40%), Positives = 14/27 (51%)
Frame = -1
Query: 172 TFSIGGQISFDVFPQGWDKTYCLRYLD 198
T IG ++S FPQ WD T Y +
Sbjct: 378 TLIIGAEVSITKFPQSWDNTILFIYFN 298
>AW666376
Length = 425
Score = 27.7 bits (60), Expect = 5.0
Identities = 12/38 (31%), Positives = 24/38 (62%)
Frame = +3
Query: 86 ESLKTFLGDEKLKEFINFTLHYIADLDIPIKRGTFIEF 123
+SL+ L KL+ F+N + ++ ++D+ + GT +EF
Sbjct: 9 DSLQNKLFLAKLEGFVNQLVRFLHNVDVGVTAGTSVEF 122
>TC205729 similar to UP|Q8GZV0 (Q8GZV0) Obtusifoliol-14-demethylase, partial
(96%)
Length = 1666
Score = 27.7 bits (60), Expect = 5.0
Identities = 13/29 (44%), Positives = 17/29 (57%)
Frame = +1
Query: 156 PKMVSVLREKFAHLNLTFSIGGQISFDVF 184
PK+ SV K H N+TF IG ++S F
Sbjct: 433 PKLGSVFTLKLFHKNITFLIGPEVSAHFF 519
>TC210448 similar to UP|CHI1_TULBA (Q9SLP4) Chitinase 1 precursor (Tulip
bulb chitinase-1) (TBC-1) , partial (70%)
Length = 1157
Score = 26.9 bits (58), Expect = 8.6
Identities = 20/104 (19%), Positives = 38/104 (36%), Gaps = 1/104 (0%)
Frame = +1
Query: 82 LIGTESLKTFLGDEKL-KEFINFTLHYIADLDIPIKRGTFIEFRSGMLNVSPIGRNCSQE 140
L+ + ++ D L +E+I + + D+PI F
Sbjct: 154 LLASTQVRAATSDSSLFREYIGALFNGVKFSDVPINPNVNFHFILSFAIDYDTSSGSPSP 333
Query: 141 ERDEFEKYDKVQNIRPKMVSVLREKFAHLNLTFSIGGQISFDVF 184
+F + QN+ P VS ++ K ++ + S+GG F
Sbjct: 334 TNGKFNIFWDNQNLTPTQVSSIKAKNPNVKVALSLGGDTVSSAF 465
>TC233795
Length = 713
Score = 26.9 bits (58), Expect = 8.6
Identities = 10/15 (66%), Positives = 11/15 (72%)
Frame = +3
Query: 207 GDKTYKGGNDHEIYE 221
GD+ YK G DHE YE
Sbjct: 243 GDEDYKNGRDHEDYE 287
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.140 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,147,365
Number of Sequences: 63676
Number of extensions: 110539
Number of successful extensions: 410
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 409
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 410
length of query: 247
length of database: 12,639,632
effective HSP length: 95
effective length of query: 152
effective length of database: 6,590,412
effective search space: 1001742624
effective search space used: 1001742624
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0284.4


