

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
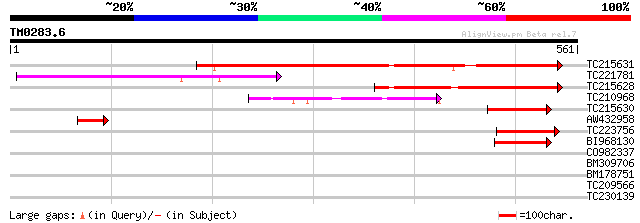
Query= TM0283.6
(561 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC215631 397 e-111
TC221781 223 2e-58
TC215628 similar to UP|Q8RY59 (Q8RY59) At1g32230/F3C3_1 (Radical... 192 4e-49
TC210968 weakly similar to UP|Q9LQC4 (Q9LQC4) F28C11.18, partial... 103 3e-22
TC215630 81 1e-15
AW432958 weakly similar to GP|552132|gb|AAA Bkm-like protein {Dr... 53 4e-07
TC223756 weakly similar to UP|Q9FJJ3 (Q9FJJ3) Arabidopsis thalia... 47 2e-05
BI968130 similar to GP|15450639|gb| AT5g62520/K19B1_13 {Arabidop... 47 3e-05
CO982337 33 0.44
BM309706 32 0.58
BM178751 28 8.3
TC209566 28 8.3
TC230139 UP|Q95RL3 (Q95RL3) LD21089p, partial (8%) 28 8.3
>TC215631
Length = 1495
Score = 397 bits (1020), Expect = e-111
Identities = 206/378 (54%), Positives = 265/378 (69%), Gaps = 16/378 (4%)
Frame = +3
Query: 186 LRECIGESNALVKDAQ-----------VKVRNMMSIMDCGYIGEEIAKNEELGLSAYTGY 234
L EC GESNALVK Q V+V + ++ DCG +GE+I +++++GL AYT
Sbjct: 3 LSECSGESNALVKGIQIDTKQNCCQYDVEVEDSINKKDCGNVGEDIQQHQDIGLDAYTES 182
Query: 235 VHGNLVLDSIRDLFCNGMSIIGNNDFEIVEIYRCSCASMQVQSKLFLTQAEITKELHGDA 294
V+G L L+S++ +F GMS G+ D +IVEIY+CS ASMQ + +LF QAE+TK+ HG+A
Sbjct: 183 VYGKLDLNSVQKMFLKGMSSFGSTDSDIVEIYQCSGASMQARWELFQKQAELTKKNHGEA 362
Query: 295 NVRYAWLSFSKGELSTMMEHGLGHCELFTTKCTYGVGVHLAAISCPYASARYCDIDGNGV 354
N+RYAWL+ SKGELSTMM +GL H L +KCTYG+GVHLAA++CP AS RYCD+D NGV
Sbjct: 363 NIRYAWLASSKGELSTMMNYGLSHYGLSGSKCTYGIGVHLAAVTCPDASVRYCDVDENGV 542
Query: 355 RHLILCRIIMGNMELLCPSIGTSNVQFQPSSSEYDNGVDDIQCPRYYTIWNMNISTYICP 414
RHL LCR+IMGNME+L P G QF PSS EYDNGVD I+CP+YY +WNMN++T+I P
Sbjct: 543 RHLALCRVIMGNMEILQPGTG----QFHPSSCEYDNGVDAIECPQYYVVWNMNMNTHIYP 710
Query: 415 AFVVSFKVSRGVEGYFCGTVDKN----NVSRDNSSG-ANSDSHCSGDPVQSASSAYNEIA 469
FVVSFKVS EG+FCG+ KN N + D G NS+S S+ N A
Sbjct: 711 EFVVSFKVSSDAEGHFCGSEGKNVSGVNTACDGPHGLLNSES----------STVDNGKA 860
Query: 470 GNGVACTQKIPKSPWMPLHMLIAAINDKVPPNDMSLIKENFELYKAKQISRDDFVKEMRL 529
+ V+ T K+PKSPWMP +L+ AI D+VPP M +IK +E +++K ISRDDFVK +RL
Sbjct: 861 PSMVSSTPKVPKSPWMPFPVLLDAIRDQVPPTGMDVIKTYYEQFRSKHISRDDFVKMLRL 1040
Query: 530 IVGDTLLRDTITTLQFMV 547
IVGD LLR TIT LQ+ +
Sbjct: 1041IVGDGLLRTTITNLQYKI 1094
>TC221781
Length = 1116
Score = 223 bits (568), Expect = 2e-58
Identities = 124/286 (43%), Positives = 166/286 (57%), Gaps = 23/286 (8%)
Frame = +1
Query: 7 LKRNETTRCGVHIRGASQQELCQQHFCASPTINFVNRMRLGDYKRKMNILGSGSGQSYLR 66
LKR TR H+ GASQ L S T V RMRLG Y+ ++ G G+S R
Sbjct: 259 LKRKRATRYAAHLSGASQPMLGHWSSLTSQTSRAVKRMRLGGYRNRLTNAGPHIGRSLAR 438
Query: 67 FYLNYKKSARPERMMFYNHGEWVDYPKDIVDLVKKDFEIKNAAIGIRLNGRDLMLDFLHA 126
+LNYKKS R ER+MFY +GEW+D+PKD+VDLVKKD E+K AA+ I NG L+ DFLH
Sbjct: 439 RFLNYKKSGRLERLMFYENGEWLDFPKDVVDLVKKDLEVKKAAVEIESNGYHLLFDFLHL 618
Query: 127 CQMDLKTGFIQPIAWIDKVGGYFFPEVDVASGEESYNFGEQE-----------GMKFHVE 175
++DLKTG QPIA ID+ G FFPE+ AS E+ YN +QE +K H+E
Sbjct: 619 HKVDLKTGLQQPIALIDEAGCCFFPEIYAASDEKPYNLSKQECGRSPDSYASNEIKLHLE 798
Query: 176 IKMNMVDEFQLRECIGESNALVKDAQVKVRN-----------MMSIMDCGYIGEEIAKNE 224
++N VD+ L EC G+SNALVK Q+ + ++ DCG +GE I +++
Sbjct: 799 XEINGVDQSXLXECSGKSNALVKGIQIDTKQNCCQYDVXXEXSINKKDCGNVGENIQQHQ 978
Query: 225 ELGLSAYTGY-VHGNLVLDSIRDLFCNGMSIIGNNDFEIVEIYRCS 269
++GL AYTG + + + F GM G+ D +I IY+CS
Sbjct: 979 DIGLDAYTGISIWEYWI*ILXKRXFLRGMXXFGSTDSDIXXIYQCS 1116
>TC215628 similar to UP|Q8RY59 (Q8RY59) At1g32230/F3C3_1 (Radical-induced
cell death 1-1), partial (9%)
Length = 930
Score = 192 bits (487), Expect = 4e-49
Identities = 103/187 (55%), Positives = 126/187 (67%), Gaps = 1/187 (0%)
Frame = +2
Query: 362 IIMGNMELLCPSIGTSNVQFQPSSSEYDNGVDDIQCPRYYTIWNMNISTYICPAFVVSFK 421
+IMGNME+L P G QF PSS EYDNGVD I+CPRYY +WNMN++T+I P FVVSFK
Sbjct: 8 VIMGNMEILQPGTG----QFHPSSCEYDNGVDSIECPRYYVVWNMNMNTHIYPEFVVSFK 175
Query: 422 VSRGVEGYFCGTVDKNNVSRDNSSGANSDSHCSGDPVQSASSAY-NEIAGNGVACTQKIP 480
VS EG+FCG+ KN SG NS + S SS N A + VA T K+P
Sbjct: 176 VSSDAEGHFCGSEGKN------VSGVNSACQGPHGLLHSESSTVDNGKAPSMVASTPKVP 337
Query: 481 KSPWMPLHMLIAAINDKVPPNDMSLIKENFELYKAKQISRDDFVKEMRLIVGDTLLRDTI 540
KSPWMP +L+ AI ++VPP M +IK +E +++K ISRDDFVK +RLIVGD LLR TI
Sbjct: 338 KSPWMPFPLLLDAIRNQVPPKGMDVIKIYYEQFRSKHISRDDFVKMLRLIVGDGLLRTTI 517
Query: 541 TTLQFMV 547
T LQF +
Sbjct: 518 TNLQFKI 538
>TC210968 weakly similar to UP|Q9LQC4 (Q9LQC4) F28C11.18, partial (17%)
Length = 889
Score = 103 bits (256), Expect = 3e-22
Identities = 65/199 (32%), Positives = 107/199 (53%), Gaps = 8/199 (4%)
Frame = +3
Query: 237 GNLVLDSIRDLFCNGMSIIGNNDFEIVEIYRCSCASMQVQSKL--FLTQAEITKELHGD- 293
G++V D I+ F G+ ++G E+V + R +C+ + Q++L F A L G
Sbjct: 288 GDVVHDLIKTRFIRGLGMLGPKT-EVVSVRRNACSDVVSQARLHSFHAHARAVARLRGGG 464
Query: 294 --ANVRYAWLSFS-KGELSTMMEHGLGHCELFTTKCTYGVGVHLAAISCPYASARYCDID 350
ANV+YAW + + +++ ++ G G +G + L+ P SAR C +
Sbjct: 465 NHANVKYAWYRTNGEDDVNDIVSQGFGFA--------HGPKLVLSPDDAPLQSARGCGVG 620
Query: 351 GNGVRHLILCRIIMGNMELLCPSIGTSNVQFQPSSSEYDNGVDDIQCPRYYTIWNMNIST 410
+GVRH +LCR+I+G E+ + + PS EYD+GVD P Y IW+ ++T
Sbjct: 621 KDGVRHALLCRVILGRSEI----VRDNTEHCYPSCEEYDSGVDSFSGPTKYIIWSNRMNT 788
Query: 411 YICPAFVVSFKVS--RGVE 427
++ PA+VVSF+VS +G+E
Sbjct: 789 HVLPAYVVSFRVSSFKGME 845
>TC215630
Length = 518
Score = 81.3 bits (199), Expect = 1e-15
Identities = 38/64 (59%), Positives = 49/64 (76%)
Frame = +1
Query: 473 VACTQKIPKSPWMPLHMLIAAINDKVPPNDMSLIKENFELYKAKQISRDDFVKEMRLIVG 532
VA T K+PKSPWMP +L+ AI ++VPP M +IK +E +++K ISRDDFVK +RLIVG
Sbjct: 325 VASTPKVPKSPWMPFPVLLDAIRNQVPPKGMDVIKTYYEQFRSKHISRDDFVKMLRLIVG 504
Query: 533 DTLL 536
D LL
Sbjct: 505 DGLL 516
>AW432958 weakly similar to GP|552132|gb|AAA Bkm-like protein {Drosophila
melanogaster}, partial (35%)
Length = 199
Score = 52.8 bits (125), Expect = 4e-07
Identities = 20/30 (66%), Positives = 26/30 (86%)
Frame = -1
Query: 68 YLNYKKSARPERMMFYNHGEWVDYPKDIVD 97
+LNYKKS R ER+MFY +G W+D+PKD+VD
Sbjct: 103 FLNYKKSGRLERLMFYENGSWLDFPKDVVD 14
>TC223756 weakly similar to UP|Q9FJJ3 (Q9FJJ3) Arabidopsis thaliana genomic
DNA, chromosome 5, TAC clone:K19B1 (AT5g62520/K19B1_13),
partial (19%)
Length = 532
Score = 47.4 bits (111), Expect = 2e-05
Identities = 21/63 (33%), Positives = 41/63 (64%)
Frame = +1
Query: 482 SPWMPLHMLIAAINDKVPPNDMSLIKENFELYKAKQISRDDFVKEMRLIVGDTLLRDTIT 541
SPWM LI+ ++ +PP++++ I + + Y+ K+ISR + ++++R+I GD LL I
Sbjct: 1 SPWMAFPALISMLSKILPPSEVASIAKFHKDYREKRISRHELIQKVRVIAGDKLLLSVIK 180
Query: 542 TLQ 544
+ +
Sbjct: 181 SFR 189
>BI968130 similar to GP|15450639|gb| AT5g62520/K19B1_13 {Arabidopsis
thaliana}, partial (6%)
Length = 688
Score = 46.6 bits (109), Expect = 3e-05
Identities = 22/57 (38%), Positives = 35/57 (60%)
Frame = -2
Query: 480 PKSPWMPLHMLIAAINDKVPPNDMSLIKENFELYKAKQISRDDFVKEMRLIVGDTLL 536
P SPWMP LI A++ +PP D++LI + ++ K K+ + ++ +R I GD LL
Sbjct: 606 PTSPWMPFPALICALSKVLPPCDIALISKFYKDKKDKKXXXXELMQRVREIAGDKLL 436
>CO982337
Length = 782
Score = 32.7 bits (73), Expect = 0.44
Identities = 31/102 (30%), Positives = 45/102 (43%), Gaps = 16/102 (15%)
Frame = -2
Query: 210 IMDCGYIGEEIAKNEELGLSAYTGYVHGNLVLDSIRDL-----FCNGMSIIGNNDFEIV- 263
I DCG EEI N+++G+ A G+V LV +L FC G N F ++
Sbjct: 778 ISDCGV--EEIIVNDQVGVEAALGFVFPKLVSIKFLNLAELRYFCTGNH---NLRFPLLN 614
Query: 264 EIYRCSCASMQVQS----------KLFLTQAEITKELHGDAN 295
++Y C +M+ S K++LTQ GD N
Sbjct: 613 KLYTVECPAMETFSQGILRASILRKIYLTQEGDQVYWEGDLN 488
>BM309706
Length = 424
Score = 32.3 bits (72), Expect = 0.58
Identities = 19/71 (26%), Positives = 37/71 (51%), Gaps = 3/71 (4%)
Frame = +3
Query: 233 GYVHGNLVLDSIRDLFCNGMSIIGNNDFEIVEIYRCSCASMQVQSK---LFLTQAEITKE 289
G+V + D +R F G++ G E++ + R +C+S+ Q++ + + K
Sbjct: 213 GFVPADENADLVRTRFVRGLAAHGLKP-EVLAVGRNACSSVMAQARQHSFHVFARAVAKL 389
Query: 290 LHGDANVRYAW 300
G+ANV++AW
Sbjct: 390 RDGNANVKFAW 422
>BM178751
Length = 421
Score = 28.5 bits (62), Expect = 8.3
Identities = 16/40 (40%), Positives = 20/40 (50%)
Frame = +2
Query: 268 CSCASMQVQSKLFLTQAEITKELHGDANVRYAWLSFSKGE 307
CS S Q+ SKL +T T L GD N+ SKG+
Sbjct: 179 CSQGSYQLLSKLSITSQLQTSSLDGDTNIAVFTWD*SKGQ 298
>TC209566
Length = 793
Score = 28.5 bits (62), Expect = 8.3
Identities = 9/19 (47%), Positives = 13/19 (68%)
Frame = -3
Query: 319 CELFTTKCTYGVGVHLAAI 337
C++FT C YG+ VH A +
Sbjct: 563 CKIFTPSCNYGINVHWAIL 507
>TC230139 UP|Q95RL3 (Q95RL3) LD21089p, partial (8%)
Length = 731
Score = 28.5 bits (62), Expect = 8.3
Identities = 17/54 (31%), Positives = 25/54 (45%)
Frame = +2
Query: 7 LKRNETTRCGVHIRGASQQELCQQHFCASPTINFVNRMRLGDYKRKMNILGSGS 60
++RN V ++ A QQ+ QQH C S N M L + + G+GS
Sbjct: 512 VRRNGVVNISVRVKVAQQQQQQQQHSCNS------NSMSLSSAVTGVPVAGNGS 655
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.138 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,712,695
Number of Sequences: 63676
Number of extensions: 361635
Number of successful extensions: 2317
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 2286
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2306
length of query: 561
length of database: 12,639,632
effective HSP length: 102
effective length of query: 459
effective length of database: 6,144,680
effective search space: 2820408120
effective search space used: 2820408120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0283.6


