

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0283.2
(142 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
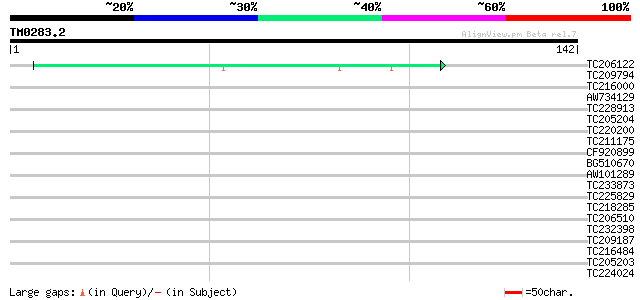
Score E
Sequences producing significant alignments: (bits) Value
TC206122 weakly similar to PIR|C86473|C86473 arabinogalactan-pro... 39 6e-04
TC209794 33 0.035
TC216000 similar to UP|O22253 (O22253) Photolyase/blue-light rec... 33 0.059
AW734129 32 0.078
TC228913 weakly similar to UP|Q8IP68 (Q8IP68) CG31813-PA, partia... 32 0.078
TC205204 homologue to UP|Q43558 (Q43558) Proline rich protein pr... 32 0.10
TC220200 weakly similar to UP|Q7M4A3 (Q7M4A3) Spermatid-specific... 32 0.10
TC211175 weakly similar to UP|Q9LIE8 (Q9LIE8) Similarity to cell... 31 0.23
CF920899 30 0.50
BG510670 30 0.50
AW101289 30 0.50
TC233873 weakly similar to UP|Q6ZJC8 (Q6ZJC8) BHLH protein famil... 30 0.50
TC225829 weakly similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like ara... 30 0.50
TC218285 weakly similar to UP|Q9SKM6 (Q9SKM6) En/Spm-like transp... 30 0.50
TC206510 similar to UP|Q84UV7 (Q84UV7) SGT1, partial (51%) 29 0.66
TC232398 29 0.66
TC209187 29 0.66
TC216484 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabin... 29 0.66
TC205203 similar to UP|Q39763 (Q39763) Proline-rich cell wall pr... 29 0.66
TC224024 weakly similar to UP|Q948Y6 (Q948Y6) VMP4 protein, part... 29 0.86
>TC206122 weakly similar to PIR|C86473|C86473 arabinogalactan-protein -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(40%)
Length = 797
Score = 39.3 bits (90), Expect = 6e-04
Identities = 33/109 (30%), Positives = 43/109 (39%), Gaps = 6/109 (5%)
Frame = +3
Query: 7 VLVVLCLAVCVGICRPDSPATSPSPSQKSDSPPSFASWAYDKFSHV-FGPYRNDEPVLPR 65
+L C+A G P+T+PSPS + +P A+ + P P P
Sbjct: 99 LLATACVAQAPGPAPTQPPSTTPSPSTSAPAPSPTANTPPPAAAPAPATPTPTPAPSSPT 278
Query: 66 TTENLQTGFGTGLGAP----VEAPAPGPFASAP-QPSSEAAQQAFSYGK 109
T G +P PAPGP AP QP+SE AFS K
Sbjct: 279 PTTAPSQAPGPTTSSPPAPGPSGPAPGPSGPAPDQPASEPPSAAFSISK 425
>TC209794
Length = 601
Score = 33.5 bits (75), Expect = 0.035
Identities = 18/47 (38%), Positives = 24/47 (50%)
Frame = +3
Query: 26 ATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQT 72
A +P+ + SFASWAYDK S G + +EPV E +T
Sbjct: 330 AAAPTIDAAAQKSGSFASWAYDKISRGLG-FGTEEPVAQTKYEETKT 467
>TC216000 similar to UP|O22253 (O22253) Photolyase/blue-light receptor
(Photolyase/blue light photoreceptor PHR2), partial
(84%)
Length = 1641
Score = 32.7 bits (73), Expect = 0.059
Identities = 26/85 (30%), Positives = 34/85 (39%), Gaps = 3/85 (3%)
Frame = +3
Query: 22 PDSPATSPSPSQKSDSPPSFASWAYDKFSHVFG---PYRNDEPVLPRTTENLQTGFGTGL 78
P +PAT P+P S +PPS S S + P P P T T
Sbjct: 216 PTAPATLPTP-PPSAAPPSSGSATTSASSTMSASPPPTTTPSPSSPSTASTPPTTASRHP 392
Query: 79 GAPVEAPAPGPFASAPQPSSEAAQQ 103
+ AP P +S P P+S AA +
Sbjct: 393 ASTRPAPTAPPSSSTPSPTSAAASR 467
>AW734129
Length = 276
Score = 32.3 bits (72), Expect = 0.078
Identities = 23/84 (27%), Positives = 29/84 (34%), Gaps = 35/84 (41%)
Frame = +2
Query: 3 NAKIVLVVLCLAVCVGI-----------------------------------CRPDSPAT 27
N V++V+CLAVCVG P A
Sbjct: 14 NVMFVVLVVCLAVCVGTSWGHDFIDDAKTKAGQAKDTAVDAAAPTVNSVIDAAAPTVDAP 193
Query: 28 SPSPSQKSDSPPSFASWAYDKFSH 51
+P+ + SFASWAY K SH
Sbjct: 194APTIDAAAQKSGSFASWAYGKISH 265
>TC228913 weakly similar to UP|Q8IP68 (Q8IP68) CG31813-PA, partial (19%)
Length = 692
Score = 32.3 bits (72), Expect = 0.078
Identities = 27/83 (32%), Positives = 41/83 (48%), Gaps = 1/83 (1%)
Frame = +2
Query: 22 PDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGAP 81
P SP+T+P+ S +++PPS S A + GP +D P+ T + AP
Sbjct: 107 PPSPSTTPTSSAPANAPPSSPS-ASTRLQTPSGP-SSDASTSPKPTST------SSRAAP 262
Query: 82 VEAPAPGPFAS-APQPSSEAAQQ 103
++P+ P AS A SS A+ Q
Sbjct: 263 SKSPSTWPSASLATSMSSPASPQ 331
>TC205204 homologue to UP|Q43558 (Q43558) Proline rich protein precursor,
partial (44%)
Length = 996
Score = 32.0 bits (71), Expect = 0.10
Identities = 23/92 (25%), Positives = 35/92 (38%)
Frame = +1
Query: 6 IVLVVLCLAVCVGICRPDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPR 65
++L+ L V G+ P+TSP+P SPP P ++ P +P
Sbjct: 130 VLLLALIFTVVAGV-GAQGPSTSPAPPTPQSSPP---------------PIQSSPPPVPS 261
Query: 66 TTENLQTGFGTGLGAPVEAPAPGPFASAPQPS 97
+ Q+ P P P P +S P S
Sbjct: 262 SPPPAQS------PPPASTPPPAPLSSPPPAS 339
>TC220200 weakly similar to UP|Q7M4A3 (Q7M4A3) Spermatid-specific protein T2
precursor, partial (40%)
Length = 506
Score = 32.0 bits (71), Expect = 0.10
Identities = 35/143 (24%), Positives = 51/143 (35%), Gaps = 6/143 (4%)
Frame = +2
Query: 3 NAKIVLVVLCLAVCVGICRP-----DSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYR 57
+A+ +++L +++ V I +SPA +P+P S P F S
Sbjct: 134 SARFSVLILLVSLLVNIAATAATAAESPAPTPTPDFPPSSSPPFPS-------------- 271
Query: 58 NDEPVLPRTTENLQTGFGTGLGAPVEAPAPGPFA-SAPQPSSEAAQQAFSYGKDKAYDIK 116
P +T N P PAP + AP PSS + D A+D
Sbjct: 272 ---PASNPSTANF---------PPSPTPAPSHHSPPAPSPSSNPSPSPAPAPVDDAHDGV 415
Query: 117 DKAEVAYEDAKDKAESAYQSAKK 139
A V D K + S KK
Sbjct: 416 SHAGVDGSDGKSSSSGGMSSGKK 484
>TC211175 weakly similar to UP|Q9LIE8 (Q9LIE8) Similarity to cell wall-plasma
membrane linker protein, partial (3%)
Length = 609
Score = 30.8 bits (68), Expect = 0.23
Identities = 23/74 (31%), Positives = 31/74 (41%)
Frame = +1
Query: 25 PATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGAPVEA 84
PAT+P + PPS A ++V P P T + + +GAP A
Sbjct: 295 PATNPPAKSPATQPPSSVPPASSPSNNVSSPPAPSPATPPPTVAPVSSP----VGAPPVA 462
Query: 85 PAPGPFASAPQPSS 98
AP P +S PSS
Sbjct: 463 EAPTPESSTGIPSS 504
Score = 27.3 bits (59), Expect = 2.5
Identities = 25/79 (31%), Positives = 31/79 (38%), Gaps = 2/79 (2%)
Frame = +1
Query: 21 RPDSPATSPSPSQKSD--SPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGL 78
+P SPAT P+P KS SPPS S A P N P T +
Sbjct: 190 KPPSPATHPAPPAKSPTLSPPS-QSPAVSPSGSTPPPATNPPAKSPATQPPSSVPPASSP 366
Query: 79 GAPVEAPAPGPFASAPQPS 97
V +P P P + P P+
Sbjct: 367 SNNVSSP-PAPSPATPPPT 420
>CF920899
Length = 743
Score = 29.6 bits (65), Expect = 0.50
Identities = 13/23 (56%), Positives = 16/23 (69%)
Frame = -3
Query: 80 APVEAPAPGPFASAPQPSSEAAQ 102
AP +APA GP ASAP P + A+
Sbjct: 687 APQKAPAEGPSASAPPPQNAPAE 619
Score = 27.7 bits (60), Expect = 1.9
Identities = 20/81 (24%), Positives = 31/81 (37%)
Frame = -3
Query: 26 ATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGAPVEAP 85
++SP+P + PS ++ P +N P + +G G+ GAP
Sbjct: 699 SSSPAPQKAPAEGPSASA----------PPPQNAPAEGPNSASPPASGSGSNEGAPSSQT 550
Query: 86 APGPFASAPQPSSEAAQQAFS 106
P P A P S+ FS
Sbjct: 549 EPAPIAPPPHGSATLLASTFS 487
>BG510670
Length = 440
Score = 29.6 bits (65), Expect = 0.50
Identities = 20/71 (28%), Positives = 28/71 (39%)
Frame = +3
Query: 22 PDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGAP 81
P P +P P Q +PP ++ ++ PY N P + + G G G AP
Sbjct: 114 PPHPYGAPQPPQSYGAPPPAQPYSAAPYAQPSAPY-NKPP--KNESHSHSGGSGGGYSAP 284
Query: 82 VEAPAPGPFAS 92
PFAS
Sbjct: 285 PATAYASPFAS 317
>AW101289
Length = 412
Score = 29.6 bits (65), Expect = 0.50
Identities = 22/83 (26%), Positives = 34/83 (40%), Gaps = 6/83 (7%)
Frame = +3
Query: 24 SPATSPSPSQKSDSPPS---FASWAYDKFSHVFGPYRNDEPVLPRTTENLQT---GFGTG 77
+P+T+P+ + + SPP+ ++W FS +N P TT + T
Sbjct: 42 APSTTPATTSPTSSPPASSKSSTWTPTSFSSTTS--QNSPPPPSETTTTPSSPPPNTATP 215
Query: 78 LGAPVEAPAPGPFASAPQPSSEA 100
AP GP +P PS A
Sbjct: 216 TSAPTSPHPSGPTPPSPSPSPAA 284
>TC233873 weakly similar to UP|Q6ZJC8 (Q6ZJC8) BHLH protein family-like,
partial (25%)
Length = 817
Score = 29.6 bits (65), Expect = 0.50
Identities = 23/77 (29%), Positives = 34/77 (43%), Gaps = 1/77 (1%)
Frame = -2
Query: 21 RPDSPATSPSPSQKS-DSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLG 79
+P P +P P +PP S ++ + S EP+ P T +L T G+
Sbjct: 627 QPSKPVPAPQPPLLPFPTPPPQHSSSFPQPSS-----SPREPLWPEPTGSLGTRPGSDRS 463
Query: 80 APVEAPAPGPFASAPQP 96
P P+P P AS P+P
Sbjct: 462 GPRSTPSPSP-ASVPRP 415
>TC225829 weakly similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like
arabinogalactan-protein 2, partial (57%)
Length = 1652
Score = 29.6 bits (65), Expect = 0.50
Identities = 23/85 (27%), Positives = 35/85 (41%)
Frame = +2
Query: 20 CRPDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLG 79
C P S +TSPSP K+ SP + +S S +T + +
Sbjct: 212 CPPSSTSTSPSPPSKTSSPSTSSSTTSAPKS---------------STRSTTAPPSSPPC 346
Query: 80 APVEAPAPGPFASAPQPSSEAAQQA 104
+ AP P P A++ P+S A+ A
Sbjct: 347 SRPPAPPPAPPATSTSPTSRPARSA 421
>TC218285 weakly similar to UP|Q9SKM6 (Q9SKM6) En/Spm-like transposon
protein, partial (18%)
Length = 765
Score = 29.6 bits (65), Expect = 0.50
Identities = 34/138 (24%), Positives = 51/138 (36%), Gaps = 1/138 (0%)
Frame = +1
Query: 3 NAKIVLVVLCLAVCVGICRPDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPV 62
+A+ ++L +++ V I +SPA +P+P K D PPS S P N P
Sbjct: 133 SARFSALLLLVSLLVNIAA-ESPAPAPTP--KPDLPPS--------SSPTSAPASNPSP- 276
Query: 63 LPRTTENLQTGFGTGLGAPVEAPAPGPFA-SAPQPSSEAAQQAFSYGKDKAYDIKDKAEV 121
P PAP + AP P + + D A+D + A +
Sbjct: 277 --------------ANSPPSPTPAPSHHSPPAPSPGTNPSPSPAPAPADDAHDGVNHAGM 414
Query: 122 AYEDAKDKAESAYQSAKK 139
D K + S KK
Sbjct: 415 GGVDEKSSSSGGMSSGKK 468
>TC206510 similar to UP|Q84UV7 (Q84UV7) SGT1, partial (51%)
Length = 885
Score = 29.3 bits (64), Expect = 0.66
Identities = 17/70 (24%), Positives = 29/70 (41%), Gaps = 1/70 (1%)
Frame = +3
Query: 4 AKIVLVVLCLAVCVGICRPDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYR-NDEPV 62
++ L L ++VC+ + P P + P + S S + W + P + N P
Sbjct: 150 SRTFLFPLSVSVCLSLSPPWLPISKQKPKRPSSKTTSNSPWTFFLRPFTSNPTKPNSTPT 329
Query: 63 LPRTTENLQT 72
P+ T N T
Sbjct: 330 EPKPTSNSTT 359
>TC232398
Length = 1054
Score = 29.3 bits (64), Expect = 0.66
Identities = 14/50 (28%), Positives = 23/50 (46%)
Frame = -1
Query: 8 LVVLCLAVCVGICRPDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYR 57
++++ +C I T P P K PS A+WA + F + G +R
Sbjct: 880 IIIISFNICGIIINYTKGVTPPPPLTKGSWTPSQATWARNAFGVITGCWR 731
>TC209187
Length = 1004
Score = 29.3 bits (64), Expect = 0.66
Identities = 21/73 (28%), Positives = 29/73 (38%)
Frame = -3
Query: 25 PATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGAPVEA 84
P + P P Q S +PP A + + P + P P++ P A
Sbjct: 228 PNSPPPPPQSSTTPPEHAQ--HPPLHPPYPPLHHSPPTPPKSYS------WPPPSPPRAA 73
Query: 85 PAPGPFASAPQPS 97
PAP P +SAP S
Sbjct: 72 PAPAPKSSAPTGS 34
>TC216484 similar to UP|FLA1_ARATH (Q9FM65) Fasciclin-like arabinogalactan
protein 1 precursor, partial (58%)
Length = 1213
Score = 29.3 bits (64), Expect = 0.66
Identities = 25/87 (28%), Positives = 33/87 (37%), Gaps = 8/87 (9%)
Frame = +1
Query: 22 PDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTENLQTGFGTGLGAP 81
P S A PSPS S +PP + + + P R P +T + AP
Sbjct: 190 PKSTAKPPSPSAPSTTPP---CRTFSRSTPPSTPSRTSSPSTSSSTTSAPRSSTRSPTAP 360
Query: 82 VE--------APAPGPFASAPQPSSEA 100
AP P P AS+ P+S A
Sbjct: 361 PSPPQCTKPPAPPPDPPASSTSPTSTA 441
>TC205203 similar to UP|Q39763 (Q39763) Proline-rich cell wall protein,
partial (48%)
Length = 1076
Score = 29.3 bits (64), Expect = 0.66
Identities = 24/94 (25%), Positives = 37/94 (38%), Gaps = 1/94 (1%)
Frame = +3
Query: 6 IVLVVLCLAVCVGI-CRPDSPATSPSPSQKSDSPPSFASWAYDKFSHVFGPYRNDEPVLP 64
+ L+ L + G+ + P+TSP+P SPP P ++ P +P
Sbjct: 216 VFLLGLIFTLVAGVGAQAQGPSTSPAPPTPQASPP---------------PVQSSPPPVP 350
Query: 65 RTTENLQTGFGTGLGAPVEAPAPGPFASAPQPSS 98
+ Q+ P P P P S+P PSS
Sbjct: 351 SSPPPSQS------PPPASTPPPSPL-SSPPPSS 431
>TC224024 weakly similar to UP|Q948Y6 (Q948Y6) VMP4 protein, partial (3%)
Length = 425
Score = 28.9 bits (63), Expect = 0.86
Identities = 22/93 (23%), Positives = 39/93 (41%), Gaps = 7/93 (7%)
Frame = +3
Query: 16 CVGICRPDSPATSPSPSQ-------KSDSPPSFASWAYDKFSHVFGPYRNDEPVLPRTTE 68
C G+ + ++P SQ K + PS ++ F + P + P P ++
Sbjct: 24 CAGVRYSGTKLSNPPQSQIRTQLCAKKCASPSETGSSFSPFGNTRSPSSSTSPPPPLSSP 203
Query: 69 NLQTGFGTGLGAPVEAPAPGPFASAPQPSSEAA 101
+ T +P +P+P P S P PSS ++
Sbjct: 204 SSPTP-----RSPSPSPSPTPPPSPPPPSSASS 287
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.311 0.128 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,955,120
Number of Sequences: 63676
Number of extensions: 97419
Number of successful extensions: 1278
Number of sequences better than 10.0: 140
Number of HSP's better than 10.0 without gapping: 1127
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1252
length of query: 142
length of database: 12,639,632
effective HSP length: 88
effective length of query: 54
effective length of database: 7,036,144
effective search space: 379951776
effective search space used: 379951776
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0283.2


