

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0283.14.1
(63 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
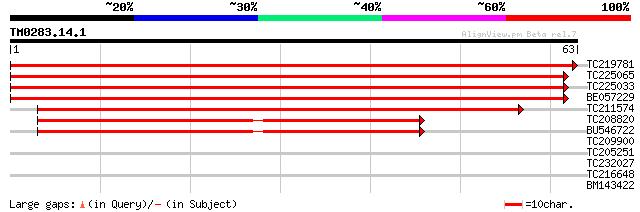
TC219781 weakly similar to UP|Q9ASW5 (Q9ASW5) At2g18360/T30D6.13... 113 2e-26
TC225065 weakly similar to UP|Q9ASW5 (Q9ASW5) At2g18360/T30D6.13... 104 8e-24
TC225033 similar to UP|O65758 (O65758) Histone H1, partial (81%) 104 8e-24
BE057229 weakly similar to GP|13605611|gb| At2g18360/T30D6.13 {A... 104 8e-24
TC211574 48 1e-06
TC208820 similar to UP|Q6NL07 (Q6NL07) At1g13820, partial (83%) 40 3e-04
BU546722 39 5e-04
TC209900 similar to UP|Q8LSY8 (Q8LSY8) Phosphoribosyltransferase... 26 3.6
TC205251 similar to UP|Q9ZP50 (Q9ZP50) FtsH-like protein Pftf pr... 26 3.6
TC232027 homologue to UP|Q7PDP0 (Q7PDP0) Erythrocyte membrane pr... 26 4.7
TC216648 weakly similar to UP|Q9FFQ8 (Q9FFQ8) Emb|CAB62360.1, pa... 25 7.9
BM143422 25 7.9
>TC219781 weakly similar to UP|Q9ASW5 (Q9ASW5) At2g18360/T30D6.13 (Expressed
protein), partial (76%)
Length = 1144
Score = 113 bits (282), Expect = 2e-26
Identities = 52/63 (82%), Positives = 57/63 (89%)
Frame = +2
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFIASF 60
RIHLLWGEND+IFK ELA++MKEQLG+G TF+GIKKAGHLVHLERPCVYNRCLK IASF
Sbjct: 758 RIHLLWGENDRIFKLELAQSMKEQLGNGTTFEGIKKAGHLVHLERPCVYNRCLKHIIASF 937
Query: 61 FAS 63
S
Sbjct: 938 LDS 946
>TC225065 weakly similar to UP|Q9ASW5 (Q9ASW5) At2g18360/T30D6.13 (Expressed
protein), partial (92%)
Length = 1046
Score = 104 bits (260), Expect = 8e-24
Identities = 47/62 (75%), Positives = 56/62 (89%)
Frame = +2
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFIASF 60
RIHLLWGEND+IF ELA+NMKEQLGDG TF+ IKKAGH+V++ERP ++NRCLK+FIASF
Sbjct: 755 RIHLLWGENDKIFNLELAQNMKEQLGDGTTFEAIKKAGHMVNMERPRLFNRCLKQFIASF 934
Query: 61 FA 62
A
Sbjct: 935 LA 940
>TC225033 similar to UP|O65758 (O65758) Histone H1, partial (81%)
Length = 1228
Score = 104 bits (260), Expect = 8e-24
Identities = 47/62 (75%), Positives = 56/62 (89%)
Frame = -1
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFIASF 60
RIHLLWGEND+IF ELA+NMKEQLGDG TF+ IKKAGH+V++ERP ++NRCLK+FIASF
Sbjct: 373 RIHLLWGENDKIFNLELAQNMKEQLGDGTTFEAIKKAGHMVNMERPRLFNRCLKQFIASF 194
Query: 61 FA 62
A
Sbjct: 193 LA 188
>BE057229 weakly similar to GP|13605611|gb| At2g18360/T30D6.13 {Arabidopsis
thaliana}, partial (19%)
Length = 345
Score = 104 bits (260), Expect = 8e-24
Identities = 47/62 (75%), Positives = 56/62 (89%)
Frame = +2
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFIASF 60
RIHLLWGEND+IF ELA+NMKEQLGDG TF+ IKKAGH+V++ERP ++NRCLK+FIASF
Sbjct: 56 RIHLLWGENDKIFNLELAQNMKEQLGDGTTFEAIKKAGHMVNMERPRLFNRCLKQFIASF 235
Query: 61 FA 62
A
Sbjct: 236 LA 241
>TC211574
Length = 1008
Score = 47.8 bits (112), Expect = 1e-06
Identities = 21/54 (38%), Positives = 33/54 (60%)
Frame = +2
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERPCVYNRCLKRFI 57
++WGE D +F ELA ++ LG+ A IK AGH +++E+P + LK F+
Sbjct: 503 IIWGEKDLVFPMELAHRLQRHLGENAQLVVIKNAGHALNVEKPKEMYKNLKSFL 664
>TC208820 similar to UP|Q6NL07 (Q6NL07) At1g13820, partial (83%)
Length = 1422
Score = 39.7 bits (91), Expect = 3e-04
Identities = 19/43 (44%), Positives = 26/43 (60%)
Frame = +2
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERP 46
++WGEND+I + A + +L D A + I GHL HLERP
Sbjct: 1022 IIWGENDRIISNKFAVRLHCELPD-AIIRQIPNCGHLPHLERP 1147
>BU546722
Length = 611
Score = 38.9 bits (89), Expect = 5e-04
Identities = 19/43 (44%), Positives = 26/43 (60%)
Frame = -1
Query: 4 LLWGENDQIFKRELAKNMKEQLGDGATFQGIKKAGHLVHLERP 46
++WGEND+I + A + +L D A + I GHL HLERP
Sbjct: 329 IIWGENDRIISNKFAVRLHCELRD-AIIRQIPYCGHLPHLERP 204
>TC209900 similar to UP|Q8LSY8 (Q8LSY8) Phosphoribosyltransferase (Fragment),
partial (83%)
Length = 575
Score = 26.2 bits (56), Expect = 3.6
Identities = 12/29 (41%), Positives = 18/29 (61%)
Frame = -1
Query: 27 DGATFQGIKKAGHLVHLERPCVYNRCLKR 55
D T+Q + K+G + H +PC + CLKR
Sbjct: 434 DWGTWQPVAKSG-MPHNVKPCQTSHCLKR 351
>TC205251 similar to UP|Q9ZP50 (Q9ZP50) FtsH-like protein Pftf precursor,
partial (8%)
Length = 419
Score = 26.2 bits (56), Expect = 3.6
Identities = 13/33 (39%), Positives = 19/33 (57%)
Frame = -1
Query: 11 QIFKRELAKNMKEQLGDGATFQGIKKAGHLVHL 43
++FK+ L +KEQLG FQ + H +HL
Sbjct: 167 RLFKQWLVLELKEQLGFQLDFQPTQTI*HGIHL 69
>TC232027 homologue to UP|Q7PDP0 (Q7PDP0) Erythrocyte membrane protein
PFEMP3, partial (3%)
Length = 335
Score = 25.8 bits (55), Expect = 4.7
Identities = 9/14 (64%), Positives = 13/14 (92%)
Frame = -3
Query: 46 PCVYNRCLKRFIAS 59
P VYNRCL++F++S
Sbjct: 255 PHVYNRCLQKFLSS 214
>TC216648 weakly similar to UP|Q9FFQ8 (Q9FFQ8) Emb|CAB62360.1, partial (41%)
Length = 2064
Score = 25.0 bits (53), Expect = 7.9
Identities = 11/25 (44%), Positives = 16/25 (64%)
Frame = -1
Query: 38 GHLVHLERPCVYNRCLKRFIASFFA 62
G L+ P + N+ L RF++SFFA
Sbjct: 1512 GLLITFTFPLILNQSL*RFVSSFFA 1438
>BM143422
Length = 424
Score = 25.0 bits (53), Expect = 7.9
Identities = 12/26 (46%), Positives = 17/26 (65%)
Frame = -1
Query: 1 RIHLLWGENDQIFKRELAKNMKEQLG 26
RIHL+ D+ +REL +N K Q+G
Sbjct: 409 RIHLI*EGVDERVQRELGQNGKSQIG 332
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.328 0.142 0.447
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,113,914
Number of Sequences: 63676
Number of extensions: 28266
Number of successful extensions: 185
Number of sequences better than 10.0: 25
Number of HSP's better than 10.0 without gapping: 183
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 183
length of query: 63
length of database: 12,639,632
effective HSP length: 39
effective length of query: 24
effective length of database: 10,156,268
effective search space: 243750432
effective search space used: 243750432
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0283.14.1


