

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
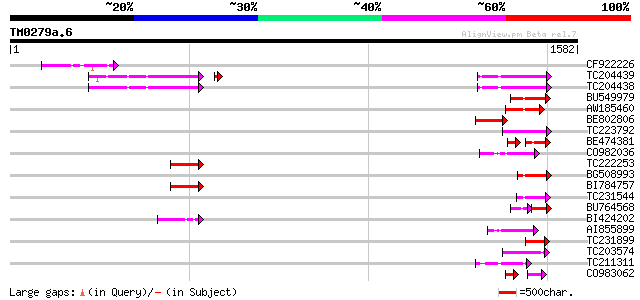
Query= TM0279a.6
(1582 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CF922226 156 6e-38
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 146 4e-35
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 144 4e-34
BU549979 102 1e-21
AW185460 101 3e-21
BE802806 95 3e-19
TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polypro... 94 5e-19
BE474381 similar to GP|21434|emb|CA ORF4 {Solanum tuberosum}, pa... 64 3e-16
CO982036 84 6e-16
TC222253 similar to UP|Q9ZQE4 (Q9ZQE4) Copia-like retroelement p... 81 3e-15
BG508993 79 2e-14
BI784757 73 9e-13
TC231544 weakly similar to UP|Q9FLR2 (Q9FLR2) Polyprotein-like, ... 68 3e-11
BU764568 50 1e-10
BI424202 65 2e-10
AI855899 similar to GP|2244960|emb| retrotransposon like protein... 65 2e-10
TC231899 similar to UP|Q850H7 (Q850H7) Gag-pol polyprotein (Frag... 64 7e-10
TC203574 weakly similar to UP|Q850I3 (Q850I3) Gag-pol polyprotei... 63 1e-09
TC211311 weakly similar to UP|O24587 (O24587) Pol protein, parti... 62 2e-09
CO983062 40 2e-08
>CF922226
Length = 667
Score = 156 bits (395), Expect = 6e-38
Identities = 90/228 (39%), Positives = 127/228 (55%), Gaps = 13/228 (5%)
Frame = -3
Query: 90 MSKTLMNKLFAKQRLYSLKMQEGGDLQAHVYAFNNILADLTRLGVTVDDEDKTIILLCSL 149
M+K+L+N+L+ KQ LYS KM E + + FN ++ DL + VT+DDED+ ++LLC L
Sbjct: 665 MTKSLVNRLYXKQSLYSFKMHEDRSVGEQLDLFNKLILDLENIDVTIDDEDQALLLLCYL 486
Query: 150 PGSYDHLVTTLTYGKDSITLDSISSTLLQHAQRRRSVEEGGGSSGEGLFVKGGQDRGRGK 209
P SY H TL +G+DS++LD + T L + E+ +SGEGL RGK
Sbjct: 485 PKSYSHFKETLLFGRDSVSLDEV-QTALNSKELNERKEKKSSASGEGL-------TARGK 330
Query: 210 GKAVDSG-KKKRSKSKDRKTAE-------CYSCKQIGHWKRDCPNRSGKSGNS-----SS 256
DS KK+ K +++K E CY CK+ GH ++ CP R G++ S
Sbjct: 329 TFKKDSEFDKKKQKPENQKNGEGNIFKIRCYHCKKEGHTRKVCPERQKNGGSNNRKKDSG 150
Query: 257 AANVVQSDGSCSEEDLLCVSYVKCTDAWVLDSGCSCHMTPHREWFNSF 304
A +VQ DG S E L+ VS W++DSGCS HMTP++ WF F
Sbjct: 149 NAAIVQDDGYESAEALM-VSEKNPETKWIMDSGCSWHMTPNKSWFEQF 9
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 146 bits (369), Expect(2) = 4e-35
Identities = 98/332 (29%), Positives = 161/332 (47%), Gaps = 12/332 (3%)
Frame = +1
Query: 220 RSKSKDRKTAECYSCKQIGHWKRDC------PNRSGKSGNSSSAANVVQSDGSCSEEDLL 273
+ K RK C+ C + GH K C P+ +S NS V + S L+
Sbjct: 1477 QQKKSKRKKWRCHYCGKYGHIKPFCYHLHGHPHHGTQSSNSRKKMMWVPKHKAVS---LV 1647
Query: 274 CVSYVKCT--DAWVLDSGCSCHMTPHREWFNSFKSCDFGYVYLGDDKPCIIKGM*QVKIA 331
+ ++ + + W LDSGCS HMT +E+ + + C YV GD I GM +
Sbjct: 1648 VHTSLRASAKEDWYLDSGCSRHMTGVKEFLLNIEPCSTSYVTFGDGSKGKIIGMGK---- 1815
Query: 332 LDDGGVRTLSQVRYVPEVTKNLISLGTLHENGYSFKSEENRDILRVSKGAMTVMRAKRTA 391
L G+ +L++V V +T NLIS+ L + G++ ++ ++ K + +M+ R+
Sbjct: 1816 LVHDGLPSLNKVLLVKGLTANLISISQLCDEGFNVNFTKSECLVTNEKSEV-LMKGSRSK 1992
Query: 392 GNIYKLLGSTVMGDVASVETDDDATKLWHMRLGHLSERGMMELYKRNLLKGVRSCTIG-- 449
N Y + + +D ++WH R GHL RGM ++ + ++G+ + I
Sbjct: 1993 DNCYLWTPQETSYSSTCLSSKEDEVRIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEG 2172
Query: 450 -LCKYCVLGKQCRVRFKTGQHKTKG-ILDYVHSDVWGPTKEPSVGGFRYFVTFTDDFSRK 507
+C C +GKQ ++ + QH+T +L+ +H D+ GP + S+GG RY DDFSR
Sbjct: 2173 RICGECQIGKQVKMSHQKLQHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRF 2352
Query: 508 VWVYFLKYKSEVFAKFKLWKAEVENQTGRKIK 539
WV F++ KSE F FK ++ + IK
Sbjct: 2353 TWVNFIREKSETFEVFKELSLRLQREKDCVIK 2448
Score = 129 bits (323), Expect = 1e-29
Identities = 69/207 (33%), Positives = 117/207 (56%)
Frame = +1
Query: 1304 LGMEIHREKGAKKLWLSQNSYVEGVLSRFDMSKANPVSTPLANHFKLSLEQCPKTASEIE 1363
LG+++ + + + ++LSQ+ Y + ++ +F M A+ TP H KLS ++ + +
Sbjct: 3892 LGLQVKQMEDS--IFLSQSRYAKNIVKKFGMENASHKRTPAPTHLKLSKDEAGTSVDQS- 4062
Query: 1364 GMSKISHASAVGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKGTADR 1423
+ S +G L+Y + +RPD+ A ++ + P H VK IL+Y+ GT+D
Sbjct: 4063 -----LYRSMIGSLLY-LTASRPDITYAVGVCARYQANPKISHLTQVKRILKYVNGTSDY 4224
Query: 1424 GIMFSREQGVVPLVVGYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIVAMSTTE 1483
GIM+ P++VGY D+D+AG DDR+ST+G F L I W S Q+ V++ST E
Sbjct: 4225 GIMYCHCSN--PMLVGYCDADWAGSADDRKSTSGGCFYLGNNLISWFSKKQNCVSLSTAE 4398
Query: 1484 AEYMAVAEAVKEALWLTGLVKKLGVEQ 1510
AEY+A + + +W+ ++K+ VEQ
Sbjct: 4399 AEYIAAGSSCSQLVWMKQMLKEYNVEQ 4479
Score = 30.4 bits (67), Expect = 6.4
Identities = 42/142 (29%), Positives = 57/142 (39%), Gaps = 2/142 (1%)
Frame = +2
Query: 1091 RRWSP*KRMRHGILFSCHMERELLVASGCTRKSQQ*QKKKVK--SSRLA*LQRGIHNRRG 1148
+ WS K M+ G F E L SG +R KKV +R L + +
Sbjct: 3257 KNWSNSKGMKSGS*FLGLRELM*LAPSGSSRTKPM---KKVS*PETRPDWLLKATLRLKV 3427
Query: 1149 LIMMRFSVWWLDTLLSGQY*LW*PAGICTESRWM*KQFSSMVI*RSKSTWRSQMDSVKLE 1208
+MR LD S Y + + + +RWM*+ M * KS W SQ D
Sbjct: 3428 *TLMRLLPQLLDLSPSDYYLV*LVSSNSSCTRWM*RAHF*MDT*MKKSMWSSQRDLQTRL 3607
Query: 1209 MADLFAN*RGLCMA*SSLQGSG 1230
+ ++ R L M *S LQ G
Sbjct: 3608 IQIMYTGSRRLSMD*SKLQELG 3673
Score = 21.9 bits (45), Expect(2) = 4e-35
Identities = 11/21 (52%), Positives = 14/21 (66%)
Frame = +2
Query: 572 HHSRMVLQRG*TGL*QRRQGA 592
HH+RM RG TGL +R G+
Sbjct: 2546 HHNRMG*LRGKTGLCKRLLGS 2608
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 144 bits (362), Expect = 4e-34
Identities = 96/329 (29%), Positives = 160/329 (48%), Gaps = 9/329 (2%)
Frame = +1
Query: 220 RSKSKDRKTAECYSCKQIGHWKRDCPNRSGKSGN---SSSAANVVQSDGSCSEEDLLCVS 276
+ K RK C+ C + GH K C + G + SSS+ + L+ +
Sbjct: 1480 QQKKSKRKKWRCHYCGKYGHIKPFCYHLHGHPHHGTQSSSSGRKMMWVPKHKIVSLVVHT 1659
Query: 277 YVKCT--DAWVLDSGCSCHMTPHREWFNSFKSCDFGYVYLGDDKPCIIKGM*QVKIALDD 334
++ + + W LDSGCS HMT +E+ + + C YV GD I GM + L
Sbjct: 1660 SLRASAKEDWYLDSGCSRHMTGVKEFLVNIEPCSTSYVTFGDGSKGKITGMGK----LVH 1827
Query: 335 GGVRTLSQVRYVPEVTKNLISLGTLHENGYSFKSEENRDILRVSKGAMTVMRAKRTAGNI 394
G+ +L++V V +T NLIS+ L + G++ ++ ++ K + +M+ R+ N
Sbjct: 1828 DGLPSLNKVLLVKGLTANLISISQLCDEGFNVNFTKSECLVTNEKSEV-LMKGSRSKDNC 2004
Query: 395 YKLLGSTVMGDVASVETDDDATKLWHMRLGHLSERGMMELYKRNLLKGVRSCTIG---LC 451
Y + + +D K+WH R GHL RGM ++ + ++G+ + I +C
Sbjct: 2005 YLWTPQETSYSSTCLFSKEDEVKIWHQRFGHLHLRGMKKIIDKGAVRGIPNLKIEEGRIC 2184
Query: 452 KYCVLGKQCRVRFKTGQHKTKG-ILDYVHSDVWGPTKEPSVGGFRYFVTFTDDFSRKVWV 510
C +GKQ ++ + QH+T +L+ +H D+ GP + S+GG RY DDFSR WV
Sbjct: 2185 GECQIGKQVKMSHQKLQHQTTSRVLELLHMDLMGPMQVESLGGKRYAYVVVDDFSRFTWV 2364
Query: 511 YFLKYKSEVFAKFKLWKAEVENQTGRKIK 539
F++ KS+ F FK ++ + IK
Sbjct: 2365 NFIREKSDTFEVFKELSLRLQREKDCVIK 2451
Score = 125 bits (314), Expect = 1e-28
Identities = 68/207 (32%), Positives = 116/207 (55%)
Frame = +1
Query: 1304 LGMEIHREKGAKKLWLSQNSYVEGVLSRFDMSKANPVSTPLANHFKLSLEQCPKTASEIE 1363
LG+++ + + + ++LSQ+ Y + ++ +F M A+ TP H KLS ++ + +
Sbjct: 3895 LGLQVKQMEDS--IFLSQSKYAKNIVKKFGMENASHKRTPAPTHLKLSKDEAGTSVDQS- 4065
Query: 1364 GMSKISHASAVGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKGTADR 1423
+ S +G L+Y + +RPD+ A ++ + P H VK IL+Y+ GT+D
Sbjct: 4066 -----LYRSMIGSLLY-LTASRPDITYAVGVCARYQANPKISHLNQVKRILKYVNGTSDY 4227
Query: 1424 GIMFSREQGVVPLVVGYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIVAMSTTE 1483
GIM+ ++VGY D+D+AG DDR+ST+G F L I W S Q+ V++ST E
Sbjct: 4228 GIMYCHCSD--SMLVGYCDADWAGSADDRKSTSGGCFYLGTNLISWFSKKQNCVSLSTAE 4401
Query: 1484 AEYMAVAEAVKEALWLTGLVKKLGVEQ 1510
AEY+A + + +W+ ++K+ VEQ
Sbjct: 4402 AEYIAAGSSCSQLVWMKQMLKEYNVEQ 4482
Score = 35.0 bits (79), Expect = 0.26
Identities = 51/223 (22%), Positives = 80/223 (35%), Gaps = 42/223 (18%)
Frame = +1
Query: 91 SKTLMNKL-FAKQRLYSLKMQEGGDLQAHVYAFNNILADLTRLGVTVDDEDKTIILLCSL 149
SK M++L + +LKM+E + I T LG + DE +L SL
Sbjct: 358 SKVKMSRLQLLATKFENLKMKEEECIHDFHMNILEIANACTALGERMTDEKLVRKILRSL 537
Query: 150 PGSYDHLVTTLTYGKD--SITLDSISSTL------LQHAQRRRSVEEGGGSSGEGLFVKG 201
P +D VT + +D ++ +D + +L L ++S S+ EG +
Sbjct: 538 PKRFDMKVTAIEEAQDICNMRVDELIGSLQTFELGLSDRTEKKSKNLAFVSNDEGEEDEY 717
Query: 202 GQDRGRGKGKAV---------------------------------DSGKKKRSKSKDRKT 228
D G AV + KK K K
Sbjct: 718 DLDTDEGLTNAVVLLGKQFNKVLNRMDRRQKPHVRNIPFDIRKGSEYQKKSDEKPSHSKG 897
Query: 229 AECYSCKQIGHWKRDCPNRSGKSGNSSSAANVVQSDGSCSEED 271
+C+ C+ GH K +CP K S V +SD + SE++
Sbjct: 898 IQCHGCEGYGHIKAECPTHLKKQRKGLS---VCRSDDTESEQE 1017
>BU549979
Length = 615
Score = 102 bits (254), Expect = 1e-21
Identities = 47/112 (41%), Positives = 71/112 (62%)
Frame = -1
Query: 1397 KFMSKPGKQHWEAVKWILRYLKGTADRGIMFSREQGVVPLVVGYVDSDYAGDLDDRRSTT 1456
++ S PG HW+ K ++RYL+GT D +M+ + + V+GY DSD+AG +D RRST+
Sbjct: 606 RYQSNPGIDHWKTAKKVMRYLQGTKDYMLMYKQTNCLE--VIGYSDSDFAGCVDSRRSTS 433
Query: 1457 GYVFTLAGGPICWKSSVQSIVAMSTTEAEYMAVAEAVKEALWLTGLVKKLGV 1508
GY+F LA G + W+SS Q+++A ST E E++ EA +WL + L V
Sbjct: 432 GYIFMLADGVVSWRSSKQTLIATSTMEVEFVPCFEATSHGVWLKSFMSSLRV 277
>AW185460
Length = 411
Score = 101 bits (251), Expect = 3e-21
Identities = 53/108 (49%), Positives = 69/108 (63%)
Frame = +2
Query: 1384 TRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKGTADRGIMFSREQGVVPLVVGYVDS 1443
TRPD+ A S + +FM P + H+ A K ILRYL+GT GI ++ E ++GY DS
Sbjct: 92 TRPDIMYATSLLSRFMQSPSQIHFGAGKRILRYLQGTKAFGIWYTTETNSE--LLGYTDS 265
Query: 1444 DYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIVAMSTTEAEYMAVAE 1491
D+AG DD +ST+GY F+L G W S Q+ VA ST EAEY+AVAE
Sbjct: 266 DWAGSTDDMKSTSGYAFSLGSGMFSWASKKQATVAQSTAEAEYVAVAE 409
>BE802806
Length = 285
Score = 94.7 bits (234), Expect = 3e-19
Identities = 43/89 (48%), Positives = 68/89 (76%)
Frame = -2
Query: 1299 AAKKILGMEIHREKGAKKLWLSQNSYVEGVLSRFDMSKANPVSTPLANHFKLSLEQCPKT 1358
+A++ILG++IHR++ +L+LSQ++Y++ V+ RF M ++ PVSTPL +H KLS+ Q P+T
Sbjct: 269 SARRILGIDIHRDRAKGELFLSQSNYLKKVVERFRMHQSKPVSTPLGHHTKLSVTQAPET 90
Query: 1359 ASEIEGMSKISHASAVGCLMYVMVCTRPD 1387
A E M++ +A+ VG +MY MVC+RPD
Sbjct: 89 AEERSKMNQTPYANGVGSIMYGMVCSRPD 3
>TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polyprotein, partial
(7%)
Length = 804
Score = 94.0 bits (232), Expect = 5e-19
Identities = 47/138 (34%), Positives = 82/138 (59%), Gaps = 1/138 (0%)
Frame = +1
Query: 1374 VGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKGTADRGIMFS-REQG 1432
+G L Y + +RP++ A S + +FM +P H +A K +LR +KGT G++F + +
Sbjct: 34 IGSLRY-LCNSRPNICFAVSLISRFMKRPRLSHMQAAKRVLRLIKGTIGSGVLFPFKAKS 210
Query: 1433 VVPLVVGYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIVAMSTTEAEYMAVAEA 1492
P ++GY DSD+ D + +ST GY+F P+ S Q ++A+ST EAEY+A +
Sbjct: 211 GKPDLLGYTDSDWKRDPEQEKSTGGYLFMYNDAPVA*SSKKQDVIALSTCEAEYVAASLG 390
Query: 1493 VKEALWLTGLVKKLGVEQ 1510
+A+W+ L+++L + +
Sbjct: 391 ACQAVWMMNLLEELKLRE 444
>BE474381 similar to GP|21434|emb|CA ORF4 {Solanum tuberosum}, partial (20%)
Length = 406
Score = 64.3 bits (155), Expect(2) = 3e-16
Identities = 33/70 (47%), Positives = 47/70 (67%)
Frame = +3
Query: 1439 GYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIVAMSTTEAEYMAVAEAVKEALW 1498
GY D+D+AG + DRRST+GY T GG + S QS+VA S+ EAE+ A+A + E LW
Sbjct: 153 GYTDADWAGSVTDRRSTSGYC-TFVGGNLVS*SKKQSVVARSSAEAEFRALAHGICETLW 329
Query: 1499 LTGLVKKLGV 1508
+ L+++L V
Sbjct: 330 VKKLLQELKV 359
Score = 40.8 bits (94), Expect(2) = 3e-16
Identities = 20/38 (52%), Positives = 25/38 (65%)
Frame = +1
Query: 1388 LAQAASQVCKFMSKPGKQHWEAVKWILRYLKGTADRGI 1425
+A A S V +FM PG +H EA ILRYLKG+ RG+
Sbjct: 7 IAFAVSMVSQFMHAPGHEHLEAAFRILRYLKGSPGRGL 120
>CO982036
Length = 674
Score = 83.6 bits (205), Expect = 6e-16
Identities = 56/166 (33%), Positives = 83/166 (49%), Gaps = 1/166 (0%)
Frame = -2
Query: 1312 KGAKKLWLSQNSYVEGVLSRFDMSKANPVSTPLANHFKLSLEQCPKTASEIEGMSKISHA 1371
K L S + + + R +A P+S+P+ KLS K+ S++ +
Sbjct: 514 KSMPDLLFSLRTSIFEIFCRKPR*QAQPISSPMTTTCKLS-----KSDSDLFS-GPTFYR 353
Query: 1372 SAVGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKGTADRGIMFSREQ 1431
S VG L Y V RP+++ A ++VC+FMS P HW VK ILRYLKG+ G+
Sbjct: 352 SVVGALQYTTVI-RPEISFAVNKVCQFMSNPLDSHWTEVKRILRYLKGSLSYGL*LKPAI 176
Query: 1432 GVVPLVV-GYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSI 1476
PL + G+ D+D+A +DD+RST+G L I W Q +
Sbjct: 175 SSQPLPIRGFCDADWASAVDDKRSTSGAAVFLGPNLISWWXXKQQV 38
>TC222253 similar to UP|Q9ZQE4 (Q9ZQE4) Copia-like retroelement pol
polyprotein, partial (4%)
Length = 919
Score = 81.3 bits (199), Expect = 3e-15
Identities = 43/92 (46%), Positives = 58/92 (62%), Gaps = 1/92 (1%)
Frame = +1
Query: 449 GLCKYCVLGKQCRVRFKTGQH-KTKGILDYVHSDVWGPTKEPSVGGFRYFVTFTDDFSRK 507
G+C C +GK+ R F TG+ + K +L VH D+ + P+ G YF+TF DDFS+K
Sbjct: 325 GVCDTCEIGKKHRESFPTGKSWRMKKLLKIVHLDLC-TVEIPTHGDNNYFITFIDDFSKK 501
Query: 508 VWVYFLKYKSEVFAKFKLWKAEVENQTGRKIK 539
+WVYFLK KSE FK++KA E Q G K+K
Sbjct: 502 MWVYFLKQKSEACNAFKMFKAFAEKQNGCKVK 597
>BG508993
Length = 374
Score = 78.6 bits (192), Expect = 2e-14
Identities = 40/93 (43%), Positives = 56/93 (60%)
Frame = +1
Query: 1418 KGTADRGIMFSREQGVVPLVVGYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIV 1477
KGT D G+ +S +VG+ DSD+AGD+DDR+STTG+VF + W S Q IV
Sbjct: 4 KGTIDFGLFYSPSNNYK--LVGFCDSDFAGDVDDRKSTTGFVFFMGDCVFTWSSKKQGIV 177
Query: 1478 AMSTTEAEYMAVAEAVKEALWLTGLVKKLGVEQ 1510
+ T EAEY+A A+WL L+++L + Q
Sbjct: 178 TLFTCEAEYVAATSCTCHAIWLRRLLEELQLLQ 276
>BI784757
Length = 430
Score = 73.2 bits (178), Expect = 9e-13
Identities = 37/91 (40%), Positives = 56/91 (60%), Gaps = 1/91 (1%)
Frame = +1
Query: 450 LCKYCVLGKQCRVRFKTGQH-KTKGILDYVHSDVWGPTKEPSVGGFRYFVTFTDDFSRKV 508
+C C+ KQ R FK + K L+ ++SDV GP + S+GG RYF++F D+ +RKV
Sbjct: 64 VCDGCLQCKQSRSTFKQNVPIRAKEKLEVIYSDVCGPMQTESLGGNRYFISFIDELTRKV 243
Query: 509 WVYFLKYKSEVFAKFKLWKAEVENQTGRKIK 539
WVY ++ KS+ F F+ +K + Q+G IK
Sbjct: 244 WVYLIRRKSDFFEVFEKFKNMAKKQSGSLIK 336
>TC231544 weakly similar to UP|Q9FLR2 (Q9FLR2) Polyprotein-like, partial (16%)
Length = 662
Score = 68.2 bits (165), Expect = 3e-11
Identities = 38/98 (38%), Positives = 58/98 (58%), Gaps = 2/98 (2%)
Frame = +3
Query: 1413 ILRYLKGTADRGIMFSREQGVVPLVVGYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSS 1472
+L+YLKG +G+ FSRE + ++G+ D+D+A +D +S T Y F L I WK+
Sbjct: 30 VLKYLKGCPRKGLSFSRESPIQ--ILGFSDADWATCIDSSKSITWYCFFLGSSLISWKAK 203
Query: 1473 VQSIV--AMSTTEAEYMAVAEAVKEALWLTGLVKKLGV 1508
Q+ V + S++EA+Y A+ E WLT L+K L V
Sbjct: 204 KQNTVSRSSSSSEAKYRALTSTTCELQWLTYLLKDLHV 317
>BU764568
Length = 420
Score = 49.7 bits (117), Expect(2) = 1e-10
Identities = 25/56 (44%), Positives = 34/56 (60%)
Frame = +3
Query: 1455 TTGYVFTLAGGPICWKSSVQSIVAMSTTEAEYMAVAEAVKEALWLTGLVKKLGVEQ 1510
T+GY + G I WKS QS+VA S+ EAEY A+A E +WL L+ +L E+
Sbjct: 165 TSGYCVLIGGNLISWKSKKQSVVAKSSAEAEYRAMALVTCELIWLKQLL*ELKFEE 332
Score = 36.6 bits (83), Expect(2) = 1e-10
Identities = 19/62 (30%), Positives = 31/62 (49%)
Frame = +1
Query: 1397 KFMSKPGKQHWEAVKWILRYLKGTADRGIMFSREQGVVPLVVGYVDSDYAGDLDDRRSTT 1456
+F++ P + HW AV IL+ K +G+++ E ++GY D+D G DR
Sbjct: 4 QFLNSPCQDHWNAVS*ILK*TKSAPGKGLIY--EDKGHSQIIGYSDAD*VGSPSDRHQDI 177
Query: 1457 GY 1458
Y
Sbjct: 178 VY 183
>BI424202
Length = 421
Score = 65.5 bits (158), Expect = 2e-10
Identities = 33/128 (25%), Positives = 68/128 (52%)
Frame = -3
Query: 412 DDDATKLWHMRLGHLSERGMMELYKRNLLKGVRSCTIGLCKYCVLGKQCRVRFKTGQHKT 471
+++++ LWH RLGH+S + L +L + C+ GKQ + K G ++
Sbjct: 395 NEESSMLWHRRLGHISIERIKRLVNEGVLSTLDFADFETYVDCIKGKQTN-KSKKGAKRS 219
Query: 472 KGILDYVHSDVWGPTKEPSVGGFRYFVTFTDDFSRKVWVYFLKYKSEVFAKFKLWKAEVE 531
+L+ +H+D+ P + + +YF+TF DD+SR +++YF+ + ++ K + ++
Sbjct: 218 SNLLEIIHTDICCPDMDAN--SLKYFITFIDDYSRYMYLYFI-LRMKL*MSLKFLRLKLR 48
Query: 532 NQTGRKIK 539
N K++
Sbjct: 47 NNVENKLR 24
>AI855899 similar to GP|2244960|emb| retrotransposon like protein {Arabidopsis
thaliana}, partial (18%)
Length = 418
Score = 65.5 bits (158), Expect = 2e-10
Identities = 47/143 (32%), Positives = 71/143 (48%), Gaps = 1/143 (0%)
Frame = +1
Query: 1333 DMSKANPVSTPLANHFKLSLEQCPKTASEIEGMSKISHASAVGCLMYVMVCTRPDLAQAA 1392
+M N +STP+ + +KLS K SE+ + + VG L YV + TRP++A
Sbjct: 10 NMLDCNGISTPMVSSYKLS-----KFGSELLPNAH-QYRDIVGALQYVTL-TRPNIAYNV 168
Query: 1393 SQVCKFMSKPGKQHWEAVKWILRYLKGTADRGIMFSREQGVVPLVV-GYVDSDYAGDLDD 1451
++V +FMS P + + VK ILRYL GT +G++ + + Y D D+ D +
Sbjct: 169 NKVSEFMSSPLQSY*LTVKRILRYLSGTVTQGLLLQPAHMDAKISLRAYNDLDWGSDPAE 348
Query: 1452 RRSTTGYVFTLAGGPICWKSSVQ 1474
RST+G I W S Q
Sbjct: 349 MRSTSGSCIFSGSNLIAWSSKKQ 417
>TC231899 similar to UP|Q850H7 (Q850H7) Gag-pol polyprotein (Fragment), partial
(30%)
Length = 687
Score = 63.5 bits (153), Expect = 7e-10
Identities = 29/68 (42%), Positives = 44/68 (64%)
Frame = +2
Query: 1439 GYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIVAMSTTEAEYMAVAEAVKEALW 1498
GY D+D+AG DRRST+GY + G + WKS Q++VA S+ EAEY ++A E +W
Sbjct: 23 GYCDADWAGCPMDRRSTSGYCVFIGGNLVSWKSKKQTVVARSSAEAEYRSMAMVTCELMW 202
Query: 1499 LTGLVKKL 1506
+ +++L
Sbjct: 203 IKQFLQEL 226
>TC203574 weakly similar to UP|Q850I3 (Q850I3) Gag-pol polyprotein (Fragment),
partial (25%)
Length = 1037
Score = 62.8 bits (151), Expect = 1e-09
Identities = 46/133 (34%), Positives = 70/133 (52%)
Frame = +1
Query: 1374 VGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKGTADRGIMFSREQGV 1433
VG L Y+ + T D+ + V +F + P HW V ILR +KG +G++ ++G
Sbjct: 640 VGKLNYLTL-TITDIYFHVNVVSQFFNAPHDSHWNVVVQILR*IKGLPGKGLI-DMDKGQ 813
Query: 1434 VPLVVGYVDSDYAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIVAMSTTEAEYMAVAEAV 1493
+V + ++++ D DRRST GY + G I WKS Q A S+TE Y AVA
Sbjct: 814 ANTIV-HRNANWERDASDRRSTIGYCVFIGGDLILWKSKKQI*AARSSTEV-YWAVANTT 987
Query: 1494 KEALWLTGLVKKL 1506
E L L ++++L
Sbjct: 988 CELL*LKLMLQEL 1026
>TC211311 weakly similar to UP|O24587 (O24587) Pol protein, partial (15%)
Length = 1213
Score = 62.0 bits (149), Expect = 2e-09
Identities = 46/156 (29%), Positives = 74/156 (46%)
Frame = +3
Query: 1301 KKILGMEIHREKGAKKLWLSQNSYVEGVLSRFDMSKANPVSTPLANHFKLSLEQCPKTAS 1360
+K+ G+ IH+EK Y + L RF M +A P++TP+ + ++ S
Sbjct: 633 QKVYGIFIHQEK-----------YTKSHLKRFRMDEAKPMATPMHRSTIIDKDEKGNHTS 779
Query: 1361 EIEGMSKISHASAVGCLMYVMVCTRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLKGT 1420
E ++ + L Y + +RPD+ +F S P H AVK ILRYL GT
Sbjct: 780 *KE------YSGMIDSLSY-LTSSRPDIVFVVCLCARFQSYPKISHVTAVKRILRYLVGT 938
Query: 1421 ADRGIMFSREQGVVPLVVGYVDSDYAGDLDDRRSTT 1456
+ + F + ++GY D +AGD +R+ST+
Sbjct: 939 TNHCLWFKKRSEFD--LLGYCDVYFAGDKVERKSTS 1040
>CO983062
Length = 390
Score = 40.4 bits (93), Expect(2) = 2e-08
Identities = 20/54 (37%), Positives = 28/54 (51%)
Frame = +3
Query: 1445 YAGDLDDRRSTTGYVFTLAGGPICWKSSVQSIVAMSTTEAEYMAVAEAVKEALW 1498
+A ++DDRRST+ I W S Q + A S+TEAEY + + E W
Sbjct: 228 WASNIDDRRSTSRAAIFPGRNLISWWSKKQKVTARSSTEAEYQILVQTSIELTW 389
Score = 38.1 bits (87), Expect(2) = 2e-08
Identities = 16/35 (45%), Positives = 23/35 (65%)
Frame = +2
Query: 1384 TRPDLAQAASQVCKFMSKPGKQHWEAVKWILRYLK 1418
TRP++ ++V ++M+ P HW VK ILRYLK
Sbjct: 44 TRPEINYVVNKVXQYMTNPLDSHWAVVKRILRYLK 148
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.341 0.149 0.500
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 76,301,870
Number of Sequences: 63676
Number of extensions: 1189326
Number of successful extensions: 12840
Number of sequences better than 10.0: 122
Number of HSP's better than 10.0 without gapping: 10129
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 11850
length of query: 1582
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1472
effective length of database: 5,635,272
effective search space: 8295120384
effective search space used: 8295120384
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (22.0 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0279a.6


