

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0278.5.1
(30 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
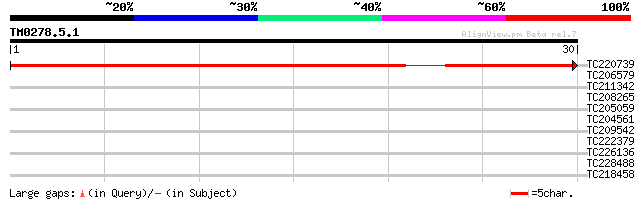
Score E
Sequences producing significant alignments: (bits) Value
TC220739 weakly similar to UP|PRP4_HUMAN (O43172) U4/U6 small nu... 45 1e-05
TC206579 weakly similar to UP|TAF5_YEAST (P38129) Transcription ... 27 1.9
TC211342 similar to UP|Q9LVF2 (Q9LVF2) Arabidopsis thaliana geno... 27 3.3
TC208265 similar to UP|Q6S7B0 (Q6S7B0) TAF5, partial (36%) 26 4.3
TC205059 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-... 26 5.6
TC204561 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-... 26 5.6
TC209542 25 7.3
TC222379 homologue to UP|Q9HGV7 (Q9HGV7) Gbeta like protein, par... 25 9.6
TC226136 similar to UP|O48847 (O48847) Expressed protein (At2g32... 25 9.6
TC228488 similar to UP|Q8LF96 (Q8LF96) PRL1 protein, partial (70%) 25 9.6
TC218458 similar to PIR|T01391|T01391 WD-repeat protein T4I9.10 ... 25 9.6
>TC220739 weakly similar to UP|PRP4_HUMAN (O43172) U4/U6 small nuclear
ribonucleoprotein Prp4 (U4/U6 snRNP 60 kDa protein) (WD
splicing factor Prp4) (hPrp4), partial (34%)
Length = 1206
Score = 44.7 bits (104), Expect = 1e-05
Identities = 23/30 (76%), Positives = 24/30 (79%)
Frame = +1
Query: 1 GGYIVTVSHDRTIKLWSSNMTNDNEHTMDV 30
GG IVTVSHDRTIKLWSSN T +E MDV
Sbjct: 862 GGSIVTVSHDRTIKLWSSNPT--DEQAMDV 945
>TC206579 weakly similar to UP|TAF5_YEAST (P38129) Transcription initiation
factor TFIID subunit 5 (TBP-associated factor 5)
(TBP-associated factor 90 kDa) (TAFII-90), partial (4%)
Length = 835
Score = 27.3 bits (59), Expect = 1.9
Identities = 10/23 (43%), Positives = 14/23 (60%)
Frame = +1
Query: 1 GGYIVTVSHDRTIKLWSSNMTND 23
G Y++T S D T +LWS + D
Sbjct: 460 GAYLITASSDTTARLWSMSTGED 528
>TC211342 similar to UP|Q9LVF2 (Q9LVF2) Arabidopsis thaliana genomic DNA,
chromosome 3, P1 clone: MIL23, partial (29%)
Length = 862
Score = 26.6 bits (57), Expect = 3.3
Identities = 10/16 (62%), Positives = 13/16 (80%)
Frame = +1
Query: 1 GGYIVTVSHDRTIKLW 16
G +IVT SHDR+I+ W
Sbjct: 634 GDFIVTGSHDRSIRRW 681
>TC208265 similar to UP|Q6S7B0 (Q6S7B0) TAF5, partial (36%)
Length = 1048
Score = 26.2 bits (56), Expect = 4.3
Identities = 9/17 (52%), Positives = 12/17 (69%)
Frame = +2
Query: 1 GGYIVTVSHDRTIKLWS 17
G Y + SHDRT ++WS
Sbjct: 149 GHYFASSSHDRTARIWS 199
>TC205059 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-binding
protein beta subunit-like protein, complete
Length = 1492
Score = 25.8 bits (55), Expect = 5.6
Identities = 10/15 (66%), Positives = 13/15 (86%)
Frame = +2
Query: 4 IVTVSHDRTIKLWSS 18
IV+ S DRTIKLW++
Sbjct: 578 IVSASRDRTIKLWNT 622
Score = 25.0 bits (53), Expect = 9.6
Identities = 11/19 (57%), Positives = 15/19 (78%)
Frame = +2
Query: 4 IVTVSHDRTIKLWSSNMTN 22
IV+ S DRT+K+W N+TN
Sbjct: 716 IVSASWDRTVKVW--NLTN 766
>TC204561 homologue to UP|GBLP_SOYBN (Q39836) Guanine nucleotide-binding
protein beta subunit-like protein, complete
Length = 1419
Score = 25.8 bits (55), Expect = 5.6
Identities = 10/15 (66%), Positives = 13/15 (86%)
Frame = +1
Query: 4 IVTVSHDRTIKLWSS 18
IV+ S DRTIKLW++
Sbjct: 502 IVSASRDRTIKLWNT 546
Score = 25.0 bits (53), Expect = 9.6
Identities = 11/19 (57%), Positives = 15/19 (78%)
Frame = +1
Query: 4 IVTVSHDRTIKLWSSNMTN 22
IV+ S DRT+K+W N+TN
Sbjct: 640 IVSASWDRTVKVW--NLTN 690
>TC209542
Length = 1445
Score = 25.4 bits (54), Expect = 7.3
Identities = 8/17 (47%), Positives = 13/17 (76%)
Frame = +2
Query: 1 GGYIVTVSHDRTIKLWS 17
G Y+++ DRTI+LW+
Sbjct: 80 GNYVLSCGKDRTIRLWN 130
>TC222379 homologue to UP|Q9HGV7 (Q9HGV7) Gbeta like protein, partial (84%)
Length = 950
Score = 25.0 bits (53), Expect = 9.6
Identities = 13/25 (52%), Positives = 17/25 (68%)
Frame = +1
Query: 4 IVTVSHDRTIKLWSSNMTNDNEHTM 28
IV+ S DRTIKLW N D ++T+
Sbjct: 205 IVSGSRDRTIKLW--NTLGDCKYTI 273
>TC226136 similar to UP|O48847 (O48847) Expressed protein (At2g32700/F24L7.16),
partial (42%)
Length = 1463
Score = 25.0 bits (53), Expect = 9.6
Identities = 7/18 (38%), Positives = 13/18 (71%)
Frame = +3
Query: 2 GYIVTVSHDRTIKLWSSN 19
G + + SHD+++K+W N
Sbjct: 1017 GMVASASHDKSVKIWK*N 1070
>TC228488 similar to UP|Q8LF96 (Q8LF96) PRL1 protein, partial (70%)
Length = 1303
Score = 25.0 bits (53), Expect = 9.6
Identities = 9/13 (69%), Positives = 11/13 (84%)
Frame = +3
Query: 4 IVTVSHDRTIKLW 16
+VT SHD TIK+W
Sbjct: 933 VVTGSHDTTIKMW 971
>TC218458 similar to PIR|T01391|T01391 WD-repeat protein T4I9.10 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(30%)
Length = 788
Score = 25.0 bits (53), Expect = 9.6
Identities = 12/28 (42%), Positives = 18/28 (63%), Gaps = 5/28 (17%)
Frame = +1
Query: 1 GGYIVTVSHDRTIKLW-----SSNMTND 23
G Y ++ D+TIKLW SSN+T++
Sbjct: 85 GRYFISNGKDQTIKLWDIRKMSSNVTSN 168
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.128 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,469,090
Number of Sequences: 63676
Number of extensions: 6728
Number of successful extensions: 97
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 79
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 97
length of query: 30
length of database: 12,639,632
effective HSP length: 6
effective length of query: 24
effective length of database: 12,257,576
effective search space: 294181824
effective search space used: 294181824
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0278.5.1


