

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0269b.2
(2064 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
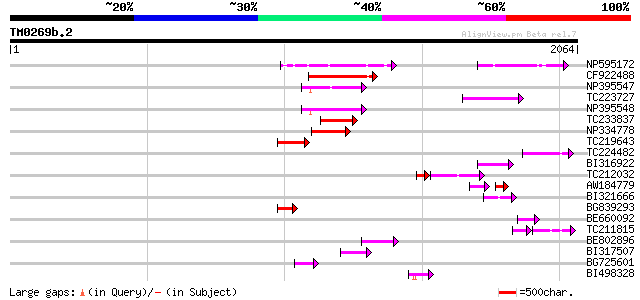
Sequences producing significant alignments: (bits) Value
NP595172 polyprotein [Glycine max] 206 1e-52
CF922488 203 7e-52
NP395547 reverse transcriptase [Glycine max] 158 2e-38
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 152 2e-36
NP395548 reverse transcriptase [Glycine max] 150 6e-36
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 145 1e-34
NP334778 reverse transcriptase [Glycine max] 137 4e-32
TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotei... 103 1e-21
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 102 1e-21
BI316922 100 7e-21
TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 72 5e-19
AW184779 59 3e-17
BI321666 72 2e-12
BG839293 67 8e-11
BE660092 weakly similar to GP|9884624|dbj retroelement pol polyp... 66 1e-10
TC211815 46 2e-10
BE802896 66 2e-10
BI317507 63 2e-09
BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] -... 62 3e-09
BI498328 59 2e-08
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 206 bits (523), Expect = 1e-52
Identities = 141/423 (33%), Positives = 215/423 (50%)
Frame = +1
Query: 986 DGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVELKLPIKEDKKP 1045
D L+ PT I DPEL LL + FA P ++ +P+K+ P
Sbjct: 1645 DTLKDLPTNI----DPEL----AILLHTYAQVFAVPASLPPQREQDHA---IPLKQGSGP 1791
Query: 1046 VKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGKMRVCIDFRDL 1105
VK P R+ +I++ I+ +L I+ + L ++ V KK+G R C D+R L
Sbjct: 1792 VKVRPYRYPHTQKDQIEKMIQEMLVQGIIQPSNSPFSLP-ILLVKKKDGSWRFCTDYRAL 1968
Query: 1106 NAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTY 1165
NA T KD + MP + ++D G +Y S LD SGY+QI + ED KTAFR G Y
Sbjct: 1969 NAITVKDSFPMPTVDELLDELHGAQYFSKLDLRSGYHQILVQPEDREKTAFRTHH--GHY 2142
Query: 1166 EWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSFERM 1225
EW+VMPFGL NA AT+Q +MN IF + F+ V+ DDI++ S S DHL HL + +
Sbjct: 2143 EWLVMPFGLTNAPATFQCLMNKIFQFALRKFVLVFFDDILIYSASWKDHLKHLESVLQTL 2322
Query: 1226 RKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLLGKI 1285
+++ L KC+FG D+LG V G+ + K +A+LD P + KQL+ LG
Sbjct: 2323 KQHQLFARLSKCSFGDTEVDYLGHKVSGLGVSMENTKVQAVLDWPTPNNVKQLRGFLGLT 2502
Query: 1286 NFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPIMASPIRGKP 1345
+ RRFI + + PL L ++D+F W E + AF +LK ++ P+++ P +P
Sbjct: 2503 GYYRRFIKSYANIA---GPLTDLLQKDSFLWNNEAEAAFVKLKKAMTEAPVLSLPDFSQP 2673
Query: 1346 MKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYFSCVKLK 1405
L A+ +G++L Q I Y S+ L + + + L + + K +
Sbjct: 2674 FILETDASGIGVGAVLGQNG-----HPIAYFSKKLAPRMQKQSAYTRELLAITEALSKFR 2838
Query: 1406 YYI 1408
+Y+
Sbjct: 2839 HYL 2847
Score = 108 bits (270), Expect = 2e-23
Identities = 94/339 (27%), Positives = 148/339 (42%), Gaps = 8/339 (2%)
Frame = +1
Query: 1702 HDGLCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSII 1761
H G H AGI + YWP + +D Y ++C CQ+ +PA L +
Sbjct: 3235 HSSPIGGH-AGITRTLARLKAQFYWPKMQEDVKAYIQKCLICQQAKSNNTLPAGLLQPL- 3408
Query: 1762 KPWPFRGW---ALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIE--FI 1816
P P + W A+D I + +S I+V ID TK+ IPL+ V+ F+
Sbjct: 3409 -PIPQQVWEDVAMDFITGLP--NSFGLSVIMVVIDRLTKYAHFIPLKADYNSKVVAEAFM 3579
Query: 1817 SSIVYRFGLPESLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLIS 1876
S IV G+P S+ +D+ VF + G L S+ Y+ Q++GQ E NK L
Sbjct: 3580 SHIVKLHGIPRSIVSDRDRVFTSTFWQHLFKLQGTTLAMSSAYHPQSDGQSEVLNKCLEM 3759
Query: 1877 LIKKHVGRKPKNWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEVYLQSCRIQR 1936
++ PK W + L + Y + S G TPFR YG+E P + Q+C I
Sbjct: 3760 YLRCFTYEHPKGWVKALPWAEFWYNTAYHMSLGMTPFRALYGRE---PPTLTRQACSIDD 3930
Query: 1937 QEEIPSEDYWNMMLDELVNLDEERLLVLDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKV 1996
E+ ++L + D + LTR + + + +KK SF +GD V
Sbjct: 3931 PAEV---------REQLTDRDALLAKLKINLTRAQQVMKRQADKKRLDVSFQIGDEVLVK 4083
Query: 1997 VLPINKKD---KRYRKWVPNWEGPFTVEKVLVNNAYSIK 2032
+ P + ++ +K + GPF V + + AY ++
Sbjct: 4084 LQPYRQHSAVLRKNQKLSMRYFGPFKVLAKIGDVAYKLE 4200
>CF922488
Length = 741
Score = 203 bits (516), Expect = 7e-52
Identities = 103/249 (41%), Positives = 162/249 (64%)
Frame = +3
Query: 1089 VIKKNGKMRVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAE 1148
V+K++GK+ +C+D+RDLN A+PKD++ +P ++VD+ S +DG+SGYNQI IA
Sbjct: 3 VLKEDGKV*MCVDYRDLN*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQIKIAP 182
Query: 1149 EDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKS 1208
ED+ KT F GT+ + M FGLKN GATYQR M +F D + ++VY+DD++VKS
Sbjct: 183 EDMEKTTFIT--LWGTFCYKAMSFGLKNVGATYQRAMVALF*DMMHKEIEVYMDDMIVKS 356
Query: 1209 PSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILD 1268
+ ++HL++LRK F R+RKY L++NP KC F V + L F+ +GIE++ NK K IL+
Sbjct: 357 RTEEEHLVNLRKLFRRLRKYRLRLNPAKCMFEVKSRKLLDFIDS*RGIEVDSNKVKVILE 536
Query: 1269 TSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELK 1328
+ P ++KQ+Q LG++N++ RFI+ L + + L K +W+ + AF+ +K
Sbjct: 537 MAKPHTEKQVQGFLGRLNYIVRFIS*LIATCEPL--FILLCKNQFVKWDHDC*VAFERIK 710
Query: 1329 VYLSSPPIM 1337
L +P ++
Sbjct: 711 QCLINPHVL 737
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 158 bits (400), Expect = 2e-38
Identities = 94/255 (36%), Positives = 131/255 (50%), Gaps = 18/255 (7%)
Frame = +1
Query: 1061 IKEEIERLLRCKFIRTARYVDWLANVVPVIKKNG------------------KMRVCIDF 1102
+++E+ +LL I W++ V V KK G + R+CID+
Sbjct: 1 VRKEVFKLLEAGLIYPISDSSWVSPVQVVPKKGGMTVVKNDRNELIPTRRVTRWRMCIDY 180
Query: 1103 RDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGAL 1162
R LN AT KD Y +P + M+ A + LDGYSGYNQI + +D KTAF CP ++
Sbjct: 181 RKLNEATRKDHYPLPFMDQMLKRLARQSFYRFLDGYSGYNQIAVDPQDQEKTAFTCPFSV 360
Query: 1163 GTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSF 1222
Y MPFGL NA T+QR M IF D +E ++V++DD S + L +L K
Sbjct: 361 FAYRR--MPFGLCNASTTFQRCMMAIFDDMVEKCIEVFMDDFSFFGASFGNCLANLEKVL 534
Query: 1223 ERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLL 1282
+R K L +N KC F V G LG + K+GIE+ K K I PP + K + S L
Sbjct: 535 QRCEKSNLVLNWEKCHFMVQEGIVLGHKISKRGIEVVKEKLDVIDKLPPPVNVKGIHSFL 714
Query: 1283 GKINFLRRFIANLSE 1297
G + F RRFI + ++
Sbjct: 715 GHVGFYRRFIKDFTK 759
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 152 bits (383), Expect = 2e-36
Identities = 80/223 (35%), Positives = 116/223 (51%), Gaps = 1/223 (0%)
Frame = +1
Query: 1649 DYLQNPVGTTDRKTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCGA 1708
+YL R + A + + G+ L+K+N D L+C+ +A I VH+G G
Sbjct: 175 EYLPEIADNDKRTLRRLAAGFFMSGSILYKRNHDMKPLRCVDAREANHMIEEVHEGSFGT 354
Query: 1709 HQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRG 1768
H G M + R G YW T+ DC + ++C CQ A + P L+ + PWPF
Sbjct: 355 HANGHAMARKILRAGYYWLTMESDCCVHVRKCHKCQAFADNVNAPPHPLNVMSSPWPFSM 534
Query: 1769 WALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFI-SSIVYRFGLPE 1827
W +D+IG I P +S H++I+VAIDYFTKWVEA +V + V+ FI I+ R+GLP
Sbjct: 535 WGIDVIGAIEPKASNGHRFILVAIDYFTKWVEAASYTDVMRGVVVRFIKKEIICRYGLPR 714
Query: 1828 SLTTDQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAA 1870
+ TD GT + + E + I+ TPY + N VE A
Sbjct: 715 KIITDNGTNLNNKMMGEICEEFKIQHHNPTPYRPKMN*AVEVA 843
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 150 bits (379), Expect = 6e-36
Identities = 88/255 (34%), Positives = 132/255 (51%), Gaps = 18/255 (7%)
Frame = +1
Query: 1061 IKEEIERLLRCKFIRTARYVDWLANVVPVIKKNG------------------KMRVCIDF 1102
+++E+ +LL I W++ V+ V KK G ++CID+
Sbjct: 1 VRKEVLKLLEVGLIYPISDSAWVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDY 180
Query: 1103 RDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAFRCPGAL 1162
R LN AT KD + +P + M++ AGH Y LD Y GYNQI + +D K AF CP
Sbjct: 181 RKLNEATRKDHFPLPFMDQMLERLAGHAYYCFLDAYFGYNQIVVDPKDQEKMAFTCP--F 354
Query: 1163 GTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLLHLRKSF 1222
G + + +PFGL NA T+Q M IF D +E ++V++DD V PS + L L
Sbjct: 355 GVFAYRRIPFGLCNAPTTFQMCMLAIFADIVEKSIEVFMDDFSVFVPSLESCLKKLEMVL 534
Query: 1223 ERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNKAKAILDTSPPTSKKQLQSLL 1282
+R + L +N KC F V G LG + +GIE+++ K I PP++ K ++S L
Sbjct: 535 QRCVETNLVLNWEKCHFMVREGIVLGHKISTRGIEVDQTKIDVIEKLPPPSNVKGIRSFL 714
Query: 1283 GKINFLRRFIANLSE 1297
G+ F RRFI + ++
Sbjct: 715 GQARFYRRFIKDFTK 759
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 145 bits (367), Expect = 1e-34
Identities = 71/134 (52%), Positives = 95/134 (69%)
Frame = +2
Query: 1133 SLLDGYSGYNQIFIAEEDVSKTAFRCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDF 1192
S +DG+SGYNQI +A EDV KT F GT+ + VM FGLKN GATYQR M +FHD
Sbjct: 5 SFMDGFSGYNQI*MAREDVEKTTFVT--LWGTFSYRVMAFGLKNTGATYQRAMVALFHDM 178
Query: 1193 IETFMQVYIDDIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVH 1252
+ ++VY+DD++ KS + +HL++L K F R++KY LK+NP KC FGV +G LGF+V
Sbjct: 179 MHKEIEVYVDDMIAKSRTETEHLVNLCKLFGRLQKYQLKLNPTKCTFGVKSGKLLGFIVS 358
Query: 1253 KKGIEINKNKAKAI 1266
+KGIEI+ K KA+
Sbjct: 359 QKGIEIDPEKVKAL 400
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 137 bits (346), Expect = 4e-32
Identities = 63/145 (43%), Positives = 98/145 (67%)
Frame = +3
Query: 1097 RVCIDFRDLNAATPKDEYHMPIAEMMVDSAAGHEYLSLLDGYSGYNQIFIAEEDVSKTAF 1156
R+C+D+RDLN A+PKD + +P ++++ + A S +DG+SGYNQI +A ED+ KT F
Sbjct: 3 RMCVDYRDLNRASPKDNFPLPHIDILMANMASFALFSFMDGFSGYNQIKMAPEDMEKTTF 182
Query: 1157 RCPGALGTYEWVVMPFGLKNAGATYQRVMNTIFHDFIETFMQVYIDDIVVKSPSRDDHLL 1216
GT+ + VM FGLKN GATY R M +F D + ++ Y+D+++ KS ++HL+
Sbjct: 183 IT--LWGTFCYKVMSFGLKNFGATYHRAMVALFQDMMHKEIEAYVDEMIAKSRMEEEHLV 356
Query: 1217 HLRKSFERMRKYGLKMNPLKCAFGV 1241
+L+ F ++RKY L++NP KC FG+
Sbjct: 357 NLQNLFGQLRKYRLRLNPRKCVFGL 431
>TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotein (Fragment),
partial (8%)
Length = 1320
Score = 103 bits (256), Expect = 1e-21
Identities = 47/114 (41%), Positives = 73/114 (63%)
Frame = +3
Query: 976 QDPLEEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVEL 1035
Q+ E VDLG G KR I I +++ ++ LL++++D FAW Y +MPGLS ++V+
Sbjct: 975 QEETELVDLGSGSGKREVKIGTGITAPIREELIILLRDYQDIFAWSYQDMPGLSSDIVQH 1154
Query: 1036 KLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLANVVPV 1089
+LP+ + PVKQ RR P+ +KIKEE+++ F+ ARY W+AN+VP+
Sbjct: 1155 RLPLNPECSPVKQKLRRMKPETSLKIKEEVKK*FDAGFLAVARYPKWVANIVPI 1316
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 102 bits (255), Expect = 1e-21
Identities = 64/184 (34%), Positives = 95/184 (50%), Gaps = 1/184 (0%)
Frame = +1
Query: 1868 EAANKTLISLIKKHVGRKPKNWHQTLGQVLWAYRNSPKESTGATPFRLAYGQEAVLPVEV 1927
EAANK + +I+K + K+WH+ L L YR S + STGATPF L YG EAVLP EV
Sbjct: 1 EAANKNIKKIIQK-MTVSYKDWHEMLPFALHGYRTSVRTSTGATPFSLVYGMEAVLPFEV 177
Query: 1928 YLQSCRIQRQEEIPSEDYWNMMLDELVNLDEERLLVLDTLTRQKDRIAKAYNKKVRAKSF 1987
+ S RI + + ++ D+L ++ +RL + + R+ A++KKV + F
Sbjct: 178 EVPSLRILAESGLKESEWAQTRYDQLNLIEGKRLTAMSHGRLYQQRMKSAFDKKVCLRKF 357
Query: 1988 MLGDYVWKVVLPINKKDKRYRKWVPNWEGPFTVEKVLVNNAYSIKELGGNR-QMTINGKY 2046
GD V K + K + KW PN+EGPF V++ A + + G +N
Sbjct: 358 HEGDLVLKKMSHAVKDHR--GKWAPNYEGPFVVKRAFSGGALVLTNMDGEELPSPMNSDV 531
Query: 2047 LKTY 2050
+K Y
Sbjct: 532 VKRY 543
>BI316922
Length = 405
Score = 100 bits (249), Expect = 7e-21
Identities = 51/133 (38%), Positives = 77/133 (57%), Gaps = 1/133 (0%)
Frame = +3
Query: 1701 VHDGLCGAHQAGIKMKWILFRQGMYWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSI 1760
+H G+CG H M + R G Y T+ K C EY K+C++C + I H+ ELH+I
Sbjct: 6 MHRGICGMHSKSQLMTTRVLRVGYY**TMRKYCTEYVKKCEEC*KFGNISHLLVEELHNI 185
Query: 1761 IKPWPFRGWALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFI-SSI 1819
+ PWPF +D++ P S RQ KY++V ID FTKW+E + ++ V +F+ +I
Sbjct: 186 VAPWPFAI*GVDILRPF-PLSKRQVKYLLVGIDQFTKWIETEHIAIISIANVRKFV*RNI 362
Query: 1820 VYRFGLPESLTTD 1832
V FG+P +L +D
Sbjct: 363 VC*FGIPNTLISD 401
>TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (3%)
Length = 803
Score = 72.4 bits (176), Expect(2) = 5e-19
Identities = 58/200 (29%), Positives = 90/200 (45%), Gaps = 6/200 (3%)
Frame = +2
Query: 1533 LEILIALGAKNVVVKGDSELVIKQLTKEYKCISENLARY*TKASNLLAKFDEARLGHVSR 1592
++ I K + V GDS LVI QL E + NL Y L FDE HV+
Sbjct: 206 VQAAIDSNVKLLKVYGDSALVIHQLRGECETRDPNLIPYQAYIKELAGFFDEISFHHVA* 385
Query: 1593 VDNQEANELAQIASGYMVDKCRLKELIEVKEKLNPSDLDILVIDNMAPNDWRKPIVDYLQ 1652
+NQ A+ LA + S + + IE + + P+ LV + W I Y++
Sbjct: 386 EENQMADALATLVSMFQLTPHGDLPYIEFRCRGRPAHC-CLVEEERDGKPWYFDIKRYVE 562
Query: 1653 N---PVGTTD---RKTKYRAMSYVIMGNELFKKNVDGTLLKCLSEDDAFIAISAVHDGLC 1706
+ P+ +D R+ + A + + G+ L+K+N D LL C++ + + VH+G
Sbjct: 563 SKEYPLEASDNDKRRKRRLAAGFFMSGSILYKRNHDMVLLHCVNGKEVENMLGEVHEGSF 742
Query: 1707 GAHQAGIKMKWILFRQGMYW 1726
G H G M + R G YW
Sbjct: 743 GTHSNGHAMARKILRAGYYW 802
Score = 42.4 bits (98), Expect(2) = 5e-19
Identities = 20/47 (42%), Positives = 29/47 (61%)
Frame = +1
Query: 1482 WKLFFDGSSHKNGTGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEA 1528
W ++FD +S+ G G+G +VSP F R+ +C+NN AEYEA
Sbjct: 52 WIVWFDRASNVLGHGVGAILVSPDNQCIPFTTRLGFDCTNNMAEYEA 192
>AW184779
Length = 432
Score = 58.5 bits (140), Expect(2) = 3e-17
Identities = 25/45 (55%), Positives = 33/45 (72%)
Frame = +3
Query: 1769 WALDLIGEINPCSSRQHKYIIVAIDYFTKWVEAIPLQNVTQDTVI 1813
W +D+IG I P +S H +I+VAIDYFTKWVEA+ +VT+ VI
Sbjct: 294 WGIDVIGAIEPKASNGHHFILVAIDYFTKWVEAVSYASVTRSVVI 428
Score = 50.4 bits (119), Expect(2) = 3e-17
Identities = 25/71 (35%), Positives = 36/71 (50%)
Frame = +1
Query: 1674 NELFKKNVDGTLLKCLSEDDAFIAISAVHDGLCGAHQAGIKMKWILFRQGMYWPTIMKDC 1733
N L+K+N D LL+C+ +A + VH+G G H M + R G YW T+ DC
Sbjct: 10 NILYKRNHDMVLLRCVDAREAEQMLVEVHEGSFGTHANIHAMAQKILRVGYYWLTMESDC 189
Query: 1734 MEYAKRCQDCQ 1744
+ +C CQ
Sbjct: 190 CIHVWKCHKCQ 222
>BI321666
Length = 430
Score = 72.4 bits (176), Expect = 2e-12
Identities = 40/122 (32%), Positives = 63/122 (50%), Gaps = 1/122 (0%)
Frame = +2
Query: 1725 YWPTIMKDCMEYAKRCQDCQRHAGIQHVPASELHSIIKPWPFRGWALDLIGEINPCSSRQ 1784
Y P+I KD +A+ C CQR + LH+I++ F W +D +G P S
Sbjct: 32 YLPSIFKDAYVHAQSCNKCQRTRSVSKRNELPLHTILEVEIFDYWGIDFVGPFPP--SFS 205
Query: 1785 HKYIIVAIDYFTKWVEAIPLQNVTQDTVIEFISSIVY-RFGLPESLTTDQGTVFVGQKVA 1843
++YI+V +DY +KWVEA+ Q VI+F+ ++ R G+P L + G+ +A
Sbjct: 206 NEYILVVVDYVSKWVEAVACQKSDAKIVIKFLKKQIFSRLGVPWVLIDNGGSHLCNA*LA 385
Query: 1844 SF 1845
F
Sbjct: 386 RF 391
>BG839293
Length = 781
Score = 67.0 bits (162), Expect = 8e-11
Identities = 31/73 (42%), Positives = 47/73 (63%)
Frame = +1
Query: 976 QDPLEEVDLGDGLEKRPTYISALIDPELKDRMVKLLKEFKDCFAWDYDEMPGLSREVVEL 1035
Q+ E VDLG G KR I I +++ ++ LLK+++D FAW Y +MPGLS ++V+
Sbjct: 511 QEETELVDLGIGSGKREVKIGTGITAPIREELIILLKDYQDIFAWSYQDMPGLSSDIVQH 690
Query: 1036 KLPIKEDKKPVKQ 1048
+LP+ + PVKQ
Sbjct: 691 QLPLNPECSPVKQ 729
>BE660092 weakly similar to GP|9884624|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (13%)
Length = 378
Score = 66.2 bits (160), Expect = 1e-10
Identities = 29/81 (35%), Positives = 48/81 (58%)
Frame = -3
Query: 1847 ESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQTLGQVLWAYRNSPKE 1906
+ +G+ STPY+ Q NGQ E +N+ + +++K V K+W L LWA+R + K
Sbjct: 361 KKYGVVHRVSTPYHPQTNGQAEISNREIKRILEKIVQPSRKDWSTRLDDALWAHRTAYKA 182
Query: 1907 STGATPFRLAYGQEAVLPVEV 1927
G +P+R+ +G+ LPVE+
Sbjct: 181 PIGMSPYRVVFGKACHLPVEI 119
>TC211815
Length = 704
Score = 45.8 bits (107), Expect(2) = 2e-10
Identities = 40/156 (25%), Positives = 71/156 (44%), Gaps = 1/156 (0%)
Frame = +2
Query: 1904 PKESTGATPFRLAYGQEAVLPVEVYLQSCRIQRQEEIPSEDYWNMMLDELVNLDEERLLV 1963
P+ +T TPF L YG +LP+EV E+ +++ + LD + + E+ +++
Sbjct: 227 PQSTTHETPF*LIYGISVMLPIEVGEVFL*RHYFAEV*NKEALQIDLDLIKQVREDTVIM 406
Query: 1964 LDTLTRQKDRIAKAYNKKVRAKSFMLGDYVWKVVLPINKKDKRYRKWVPNWEGPFTVEKV 2023
K R+ + +N K+ + G + + + + K+ N EGPF +
Sbjct: 407 T*AF---KQRMTRCFNSKLPSTV*GRGPSMEGIQRSLEVLVR--SKFTTN*EGPFKIRHN 571
Query: 2024 LVNNAYSIKELGGNRQMTI-NGKYLKTYKPTLHEIN 2058
N AY ++EL G + I N +LK Y +H N
Sbjct: 572 SKNGAYKLEELSGKVVLRIWNSMHLKVYYS*VHLCN 679
Score = 40.0 bits (92), Expect(2) = 2e-10
Identities = 20/67 (29%), Positives = 33/67 (48%)
Frame = +3
Query: 1832 DQGTVFVGQKVASFAESWGIKLLTSTPYYAQANGQVEAANKTLISLIKKHVGRKPKNWHQ 1891
D G F +K+ F IK ++ + Q N + EAANK ++ +KK + W +
Sbjct: 3 DNGLQFTNRKLNEFPSGLNIKHRVTSVKHPQTNRRAEAANKVILGDLKKLLDGANGRWVE 182
Query: 1892 TLGQVLW 1898
L ++LW
Sbjct: 183 DLVEILW 203
>BE802896
Length = 416
Score = 65.9 bits (159), Expect = 2e-10
Identities = 47/137 (34%), Positives = 71/137 (51%)
Frame = -2
Query: 1280 SLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRWEAEHQKAFDELKVYLSSPPIMAS 1339
S LG F RRFI + + S LL+ KE F + + + AFD LK L + PI+ +
Sbjct: 415 SFLGHAGFYRRFIRDFRKVALPLSNLLQ--KEVEFDFNDKCK*AFDCLKRALITTPIIQA 242
Query: 1340 PIRGKPMKLYISATDGTIGSMLAQEDEDSKERAIFYLSRVLNDAETRYTMIEKLCLCLYF 1399
P P +L A++ +G +LAQ+ D R I+Y SR L+ A+ YT EK L + F
Sbjct: 241 PDWTAPFELMCDASNYALGVVLAQK-IDKLPRVIYYSSRTLDAAQANYTTTEKELLAIVF 65
Query: 1400 SCVKLKYYIKPIDVMVF 1416
+ K Y+ ++V+
Sbjct: 64 ALEKFHSYLLGTRIIVY 14
>BI317507
Length = 359
Score = 62.8 bits (151), Expect = 2e-09
Identities = 33/114 (28%), Positives = 59/114 (50%)
Frame = -1
Query: 1203 DIVVKSPSRDDHLLHLRKSFERMRKYGLKMNPLKCAFGVIAGDFLGFVVHKKGIEINKNK 1262
+I++ SP H++HL + ++K L N KC F ++LG V+ K + ++ NK
Sbjct: 353 NILIYSPDWKSHIMHLTAVLDVLKKERLVANRKKCYFSQTTIEYLGHVISKDCVAMDSNK 174
Query: 1263 AKAILDTSPPTSKKQLQSLLGKINFLRRFIANLSEKTKSFSPLLRLKKEDAFRW 1316
K++++ P + K++ S L + R+FI + + PL L K D F+W
Sbjct: 173 VKSVIEWPVPKNVKRVCSFLRLTGYYRKFIKDYGKLAP--RPLTDLTKNDGFKW 18
>BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (1%)
Length = 285
Score = 61.6 bits (148), Expect = 3e-09
Identities = 36/89 (40%), Positives = 47/89 (52%)
Frame = -3
Query: 1036 KLPIKEDKKPVKQLPRRFHPDVLVKIKEEIERLLRCKFIRTARYVDWLANVVPVIKKNGK 1095
KL I D K V Q R+ + + +E+ +L FIR Y L +VV V K NGK
Sbjct: 274 KLAICNDVKLVTQRKRKIREERCQTV*QEVVKLAIASFIRDINYST*LFSVVMVKKPNGK 95
Query: 1096 MRVCIDFRDLNAATPKDEYHMPIAEMMVD 1124
R+C D+ DLN A PKD Y +P + M D
Sbjct: 94 WRICTDYIDLN*ACPKDAYPLPNIDHMTD 8
>BI498328
Length = 335
Score = 59.3 bits (142), Expect = 2e-08
Identities = 40/108 (37%), Positives = 53/108 (49%), Gaps = 18/108 (16%)
Frame = +1
Query: 1453 KAVKGQAIADFLADQTL----------PQEII--------TYVGIQPWKLFFDGSSHKNG 1494
KAVKG A+AD+LA L P E I T+ I W + FDG+S+ G
Sbjct: 10 KAVKGSALADYLAQ*PLQGYRPMHPEFPDEDIMALFEEKRTHEDINKWIVCFDGASNALG 189
Query: 1495 TGIGMFIVSPRGIPTKFKFRIKSNCSNNEAEYEALTSGLEILIALGAK 1542
G+G +VSP F R+ +C+NN AEYEA G++ I K
Sbjct: 190 HGVGAVLVSPDDQCIPFTARLGFDCTNNMAEYEACALGVQAAIDFDVK 333
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.334 0.144 0.454
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 88,584,721
Number of Sequences: 63676
Number of extensions: 1252750
Number of successful extensions: 9803
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 9599
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 9786
length of query: 2064
length of database: 12,639,632
effective HSP length: 112
effective length of query: 1952
effective length of database: 5,507,920
effective search space: 10751459840
effective search space used: 10751459840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0269b.2


