

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
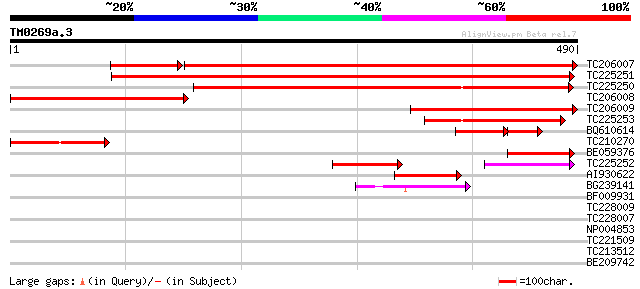
Query= TM0269a.3
(490 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC206007 similar to GB|AAO64828.1|29028898|BT005893 At3g56480 {A... 614 0.0
TC225251 weakly similar to GB|AAO64828.1|29028898|BT005893 At3g5... 513 e-145
TC225250 weakly similar to GB|AAO64828.1|29028898|BT005893 At3g5... 412 e-115
TC206008 similar to GB|AAO64828.1|29028898|BT005893 At3g56480 {A... 265 5e-71
TC206009 similar to GB|AAO64828.1|29028898|BT005893 At3g56480 {A... 254 5e-68
TC225253 homologue to UP|Q8T0J6 (Q8T0J6) GH26763p, partial (10%) 102 4e-22
BQ610614 94 1e-19
TC210270 84 2e-16
BE059376 73 3e-13
TC225252 63 3e-10
AI930622 58 1e-08
BG239141 50 2e-06
BF009931 41 0.001
TC228009 similar to UP|Q9LT13 (Q9LT13) Gb|AAF01580.1, partial (33%) 33 0.29
TC228007 similar to UP|Q9LT13 (Q9LT13) Gb|AAF01580.1, partial (51%) 32 0.84
NP004853 nodulin 31 1.4
TC221509 weakly similar to GB|AAP68225.1|31711738|BT008786 At3g4... 31 1.4
TC213512 30 1.9
BE209742 29 4.2
>TC206007 similar to GB|AAO64828.1|29028898|BT005893 At3g56480 {Arabidopsis
thaliana;} , partial (61%)
Length = 1530
Score = 614 bits (1583), Expect(2) = 0.0
Identities = 312/339 (92%), Positives = 325/339 (95%)
Frame = +2
Query: 152 ELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQ 211
+LIQEKFEVKKL+NFLKQASEDAKKLVNQEKSFACAEIESAR+VVLRIGEALEEQE+ SQ
Sbjct: 194 QLIQEKFEVKKLVNFLKQASEDAKKLVNQEKSFACAEIESARAVVLRIGEALEEQEKASQ 373
Query: 212 ASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQKELTMS 271
ASKPQDVDGL+EEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQKELTMS
Sbjct: 374 ASKPQDVDGLIEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQKELTMS 553
Query: 272 KRDEENKSHPYMLHGSEALGSYLKVQPRSGEVPQVSKCSFQWYRLSSEGSWREVISGADK 331
KRDEENKS YML GSEALGSYL+VQP S EVPQVSKCSFQWYRLSSEGSWREVISGA+K
Sbjct: 554 KRDEENKSRLYMLDGSEALGSYLRVQPCSAEVPQVSKCSFQWYRLSSEGSWREVISGANK 733
Query: 332 SIYAPDPFDVGRILQVDIVSNGKKLTLTTNPIQTASGLGSHVEALLRKSNTDFHVVISQM 391
+IYAPDP DVGRILQVDIVSNGKKLTLTTNPIQT SGLGSHVE LLRKSNTDF+VVISQM
Sbjct: 734 TIYAPDPSDVGRILQVDIVSNGKKLTLTTNPIQTVSGLGSHVETLLRKSNTDFNVVISQM 913
Query: 392 NGKDHSSRSTHSFNVGRMRIKLCRGWITKAREIYSPSMQLCGVRGDASNAAKALFWQARK 451
NGKDHSS STHSFNVGRMRIKLCRGWITKAREIYSPSMQLCGVR D +NAAK LFWQARK
Sbjct: 914 NGKDHSSHSTHSFNVGRMRIKLCRGWITKAREIYSPSMQLCGVRSDVANAAKVLFWQARK 1093
Query: 452 GLSFVLTFESEKERNVAIMIARKYALDCNVVLAGPDDLV 490
GLSFVLTFESE++RN AIM+ARKYALDCNVVLAGPDDLV
Sbjct: 1094GLSFVLTFESERDRNSAIMVARKYALDCNVVLAGPDDLV 1210
Score = 109 bits (272), Expect(2) = 0.0
Identities = 58/62 (93%), Positives = 60/62 (96%)
Frame = +1
Query: 88 AAAKLSNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLT 147
AAAKLSNEAK+R+VASLEGHVLLKKLRDALE LR RFAGRNKEDVEKAISMVEALAVKLT
Sbjct: 1 AAAKLSNEAKLRDVASLEGHVLLKKLRDALESLRGRFAGRNKEDVEKAISMVEALAVKLT 180
Query: 148 QN 149
QN
Sbjct: 181 QN 186
>TC225251 weakly similar to GB|AAO64828.1|29028898|BT005893 At3g56480
{Arabidopsis thaliana;} , partial (48%)
Length = 1272
Score = 513 bits (1320), Expect = e-145
Identities = 262/401 (65%), Positives = 331/401 (82%), Gaps = 1/401 (0%)
Frame = +1
Query: 89 AAKLSNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQ 148
AAKLS EA++RE ASLE HVLLKKLRDALE L+ R AGRNK+DVE+AI++VEALAV+LTQ
Sbjct: 10 AAKLSEEARLREAASLEKHVLLKKLRDALESLKGRVAGRNKDDVEEAIAIVEALAVQLTQ 189
Query: 149 NEGELIQEKFEVKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQ 208
EGELIQEK EVKKL NFLKQASEDAKKLV++E++FA AEIE AR+ V R+ EAL+E E+
Sbjct: 190 REGELIQEKAEVKKLANFLKQASEDAKKLVDEERAFARAEIEDARAAVQRVEEALQEHER 369
Query: 209 DSQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQKEL 268
SQAS QD++ L++EVQEARRIK+LHQPSKVM ME+ELRALR Q+ EK+ ++LQKEL
Sbjct: 370 MSQASGKQDMEQLMKEVQEARRIKMLHQPSKVMDMEHELRALRAQLAEKTRQYLRLQKEL 549
Query: 269 TMSKRDEENKSHPYMLHGSEALGSYLKVQPRSGEVPQVSKCSFQWYRLSSEGSWREVISG 328
T +K+ EN H Y L G+E LGSYL++QP S P+VS+C+ QWY +SS+G+ +E+ISG
Sbjct: 550 TRTKKGGENVPHLYELEGNETLGSYLQIQPCSDNAPEVSRCAIQWYCVSSDGAKKELISG 729
Query: 329 ADKSIYAPDPFDVGRILQVDIVSNGKKLTLTT-NPIQTASGLGSHVEALLRKSNTDFHVV 387
A KS+YAP+PFDVGRILQVDI+S + +TL+T PI A+GLG++VEAL+RK +T+F+VV
Sbjct: 730 ATKSVYAPEPFDVGRILQVDIISENEHITLSTAGPIDPAAGLGTYVEALVRKHDTEFNVV 909
Query: 388 ISQMNGKDHSSRSTHSFNVGRMRIKLCRGWITKAREIYSPSMQLCGVRGDASNAAKALFW 447
++QMNG H + S H +VG+MRIKLC+G T A+E YS SMQLCGVRG + AA+ALFW
Sbjct: 910 VTQMNGSHHPTESIHVLHVGKMRIKLCKGKTTIAKEYYSSSMQLCGVRGGGNAAAQALFW 1089
Query: 448 QARKGLSFVLTFESEKERNVAIMIARKYALDCNVVLAGPDD 488
Q ++G SFVL FESE+ERN AIM+AR++A DCN++LAGPDD
Sbjct: 1090QPKQGHSFVLAFESERERNAAIMLARRFAFDCNIMLAGPDD 1212
>TC225250 weakly similar to GB|AAO64828.1|29028898|BT005893 At3g56480
{Arabidopsis thaliana;} , partial (54%)
Length = 1302
Score = 412 bits (1059), Expect = e-115
Identities = 207/329 (62%), Positives = 269/329 (80%), Gaps = 1/329 (0%)
Frame = +2
Query: 160 VKKLLNFLKQASEDAKKLVNQEKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVD 219
VKKL NFLKQASEDAKKLVN+E++FA EIE+AR+ V R+ EAL+E E+ SQAS QD++
Sbjct: 2 VKKLTNFLKQASEDAKKLVNEERAFARFEIENARAAVHRVEEALQEHERMSQASGKQDLE 181
Query: 220 GLVEEVQEARRIKLLHQPSKVMAMEYELRALRDQIREKSIFSIKLQKELTMSKRDEENKS 279
L++EVQEARRIK+LHQPSKVM ME+ELRALR Q+ EKS ++LQKEL +K+ EEN
Sbjct: 182 QLMKEVQEARRIKMLHQPSKVMDMEHELRALRAQLAEKSRLYLRLQKELARTKKGEENAP 361
Query: 280 HPYMLHGSEALGSYLKVQPRSGEVPQVSKCSFQWYRLSSEGSWREVISGADKSIYAPDPF 339
H Y L G+E LGSYL++QP S P++S+CS QWYR+SSEG+ +E+ISGA KS+YAP+PF
Sbjct: 362 HLYELEGTETLGSYLQIQPCSDNGPELSECSIQWYRVSSEGAKKELISGATKSVYAPEPF 541
Query: 340 DVGRILQVDIVSNGKKLTL-TTNPIQTASGLGSHVEALLRKSNTDFHVVISQMNGKDHSS 398
DVGR+LQVDI+S G+++TL TT PI A+GLG++VEAL+RK +T+F+VV++Q DH +
Sbjct: 542 DVGRVLQVDIISEGQQITLSTTGPIDPAAGLGTYVEALVRKHDTEFNVVVTQ-TVSDHPN 718
Query: 399 RSTHSFNVGRMRIKLCRGWITKAREIYSPSMQLCGVRGDASNAAKALFWQARKGLSFVLT 458
S H +VG+MR+KLC+G T A+E YS SMQLCGVRG + AA+A+FWQ ++GLSFVL
Sbjct: 719 ESIHVLHVGKMRMKLCKGNTTIAKEYYSSSMQLCGVRGGGNAAAQAVFWQPKQGLSFVLA 898
Query: 459 FESEKERNVAIMIARKYALDCNVVLAGPD 487
FESE+ERN AIM+AR++A DCN++LAGPD
Sbjct: 899 FESERERNAAIMLARRFAFDCNIILAGPD 985
>TC206008 similar to GB|AAO64828.1|29028898|BT005893 At3g56480 {Arabidopsis
thaliana;} , partial (18%)
Length = 795
Score = 265 bits (676), Expect = 5e-71
Identities = 139/154 (90%), Positives = 145/154 (93%)
Frame = +2
Query: 1 MTKISPEIEIKMPMEAVPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDVIE 60
MTKISPE+EI+MPMEA PPVSADVSFIS+ FPKYKL AD+Q+LEE VEDNQGPSLKDVIE
Sbjct: 116 MTKISPEVEIRMPMEAFPPVSADVSFISNSFPKYKLDADNQVLEEPVEDNQGPSLKDVIE 295
Query: 61 QEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYL 120
QEA NLSDQHKRISVRDLA KFDKNLAAAAKLSNEAK+REVASLEGHVLLKKLRDALE L
Sbjct: 296 QEAFNLSDQHKRISVRDLASKFDKNLAAAAKLSNEAKLREVASLEGHVLLKKLRDALESL 475
Query: 121 RVRFAGRNKEDVEKAISMVEALAVKLTQNEGELI 154
R RFAG NKEDVEKAISMVEALAVKLTQNEGELI
Sbjct: 476 RGRFAGINKEDVEKAISMVEALAVKLTQNEGELI 577
Score = 32.3 bits (72), Expect = 0.49
Identities = 13/14 (92%), Positives = 14/14 (99%)
Frame = +1
Query: 477 LDCNVVLAGPDDLV 490
+DCNVVLAGPDDLV
Sbjct: 571 IDCNVVLAGPDDLV 612
>TC206009 similar to GB|AAO64828.1|29028898|BT005893 At3g56480 {Arabidopsis
thaliana;} , partial (18%)
Length = 722
Score = 254 bits (650), Expect = 5e-68
Identities = 125/144 (86%), Positives = 131/144 (90%)
Frame = +2
Query: 347 VDIVSNGKKLTLTTNPIQTASGLGSHVEALLRKSNTDFHVVISQMNGKDHSSRSTHSFNV 406
VDIVSNGK T +PIQT SGL HVE LLRKSNTDF+VVISQMNGKDHSS STHSFNV
Sbjct: 8 VDIVSNGKXXXXTXDPIQTVSGLXXHVETLLRKSNTDFNVVISQMNGKDHSSHSTHSFNV 187
Query: 407 GRMRIKLCRGWITKAREIYSPSMQLCGVRGDASNAAKALFWQARKGLSFVLTFESEKERN 466
GRMRIKLCRGWITKAREIYSPSMQLCGVR D +NAAKALFWQARKGLSFVLTFESE++RN
Sbjct: 188 GRMRIKLCRGWITKAREIYSPSMQLCGVRSDIANAAKALFWQARKGLSFVLTFESERDRN 367
Query: 467 VAIMIARKYALDCNVVLAGPDDLV 490
AIM+ARKYALDCNVVLAGPDDLV
Sbjct: 368 AAIMVARKYALDCNVVLAGPDDLV 439
>TC225253 homologue to UP|Q8T0J6 (Q8T0J6) GH26763p, partial (10%)
Length = 551
Score = 102 bits (254), Expect = 4e-22
Identities = 54/122 (44%), Positives = 77/122 (62%)
Frame = +3
Query: 359 TTNPIQTASGLGSHVEALLRKSNTDFHVVISQMNGKDHSSRSTHSFNVGRMRIKLCRGWI 418
TT PI A+GLG++VEAL+RK +T+F+VV++Q DH + S H + G+ +KLC+G
Sbjct: 189 TTGPIDPAAGLGTYVEALVRKHDTEFNVVVTQ-TVSDHPNESIHVLHAGKTTMKLCKGNT 365
Query: 419 TKAREIYSPSMQLCGVRGDASNAAKALFWQARKGLSFVLTFESEKERNVAIMIARKYALD 478
T A+E Y SMQL G RG A+ ++ F + G F +E+ RN IM+AR+ A D
Sbjct: 366 TIAKEYYISSMQLYGARGGANACTQSSFRLPKHGHCFASPSVAER*RNAVIMLARRLAFD 545
Query: 479 CN 480
CN
Sbjct: 546 CN 551
>BQ610614
Length = 462
Score = 94.0 bits (232), Expect = 1e-19
Identities = 44/46 (95%), Positives = 45/46 (97%)
Frame = +3
Query: 386 VVISQMNGKDHSSRSTHSFNVGRMRIKLCRGWITKAREIYSPSMQL 431
VVISQMNGKDHSS STHSFNVGRMRIKLCRGWITKAREIYSPSMQ+
Sbjct: 21 VVISQMNGKDHSSHSTHSFNVGRMRIKLCRGWITKAREIYSPSMQV 158
Score = 57.8 bits (138), Expect = 1e-08
Identities = 26/30 (86%), Positives = 27/30 (89%)
Frame = +1
Query: 431 LCGVRGDASNAAKALFWQARKGLSFVLTFE 460
LCGVR D +NAAK LFWQARKGLSFVLTFE
Sbjct: 373 LCGVRSDVANAAKVLFWQARKGLSFVLTFE 462
>TC210270
Length = 610
Score = 83.6 bits (205), Expect = 2e-16
Identities = 43/86 (50%), Positives = 61/86 (70%)
Frame = +2
Query: 1 MTKISPEIEIKMPMEAVPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDVIE 60
MT+++ + M EAVP VS+DV F S RFP YK+GA++QI+ ET +D + S+K+V+
Sbjct: 308 MTRVTRDFGDTMQREAVPAVSSDVVFASARFPNYKIGANNQIM-ETKDDPKVLSMKEVVA 484
Query: 61 QEATNLSDQHKRISVRDLACKFDKNL 86
+E L +QH R+SVRDLA KF+K L
Sbjct: 485 RETAQLLEQHNRLSVRDLASKFEKGL 562
>BE059376
Length = 184
Score = 72.8 bits (177), Expect = 3e-13
Identities = 34/58 (58%), Positives = 44/58 (75%)
Frame = +2
Query: 431 LCGVRGDASNAAKALFWQARKGLSFVLTFESEKERNVAIMIARKYALDCNVVLAGPDD 488
L GVRG + A +ALF Q +G SFVL FES++ERN M+AR++A DCN++LAGPDD
Sbjct: 5 LWGVRGGGNAAPQALFGQPMQGHSFVLAFESKEERNAPFMLARRFAFDCNIMLAGPDD 178
>TC225252
Length = 532
Score = 63.2 bits (152), Expect = 3e-10
Identities = 29/60 (48%), Positives = 39/60 (64%)
Frame = +1
Query: 280 HPYMLHGSEALGSYLKVQPRSGEVPQVSKCSFQWYRLSSEGSWREVISGADKSIYAPDPF 339
H Y L G+E LGSYL++QP S P+VSKCS QWYR+SS+G+ + ++K P F
Sbjct: 7 HLYELEGNETLGSYLQIQPCSDNAPEVSKCSIQWYRVSSDGAKKRTYLRSNKISLCP*TF 186
Score = 55.8 bits (133), Expect = 4e-08
Identities = 29/78 (37%), Positives = 44/78 (56%)
Frame = +2
Query: 411 IKLCRGWITKAREIYSPSMQLCGVRGDASNAAKALFWQARKGLSFVLTFESEKERNVAIM 470
+ +C+ + + ++Q G+ K L A K + FESE+ERN AIM
Sbjct: 41 VLICKFNLALIMPLKFQNVQFSGIVYHLMVPKKELISGATKSVYAPEPFESERERNAAIM 220
Query: 471 IARKYALDCNVVLAGPDD 488
+AR++A DCN++LAGPDD
Sbjct: 221 LARRFAFDCNIMLAGPDD 274
Score = 32.7 bits (73), Expect = 0.38
Identities = 13/21 (61%), Positives = 18/21 (84%)
Frame = +2
Query: 323 REVISGADKSIYAPDPFDVGR 343
+E+ISGA KS+YAP+PF+ R
Sbjct: 137 KELISGATKSVYAPEPFESER 199
>AI930622
Length = 190
Score = 57.8 bits (138), Expect = 1e-08
Identities = 27/59 (45%), Positives = 44/59 (73%), Gaps = 1/59 (1%)
Frame = +2
Query: 333 IYAPDPFDVGRILQVDIVSNGKKLT-LTTNPIQTASGLGSHVEALLRKSNTDFHVVISQ 390
+YAP+PFDVGR+L VDI+ G ++T TT I ++ G++V AL+RK +T+++ V++Q
Sbjct: 2 VYAPEPFDVGRVLHVDIIIYGHQITWSTTGTIDPSAESGTYV*ALVRKHDTEYNAVVTQ 178
>BG239141
Length = 301
Score = 50.4 bits (119), Expect = 2e-06
Identities = 34/106 (32%), Positives = 57/106 (53%), Gaps = 7/106 (6%)
Frame = +2
Query: 300 SGEVPQVSKCSFQWYRLSSEGSWREVISGADKSIYAPDPFD------VGRILQVDIVSNG 353
S P+VS CS W R ++ + P+ F +GRILQ+ I+S
Sbjct: 2 SDNAPEVSICSIHWIVC------RMMVPKMILFLEQPNQFMPLNLSMLGRILQLYIMSEN 163
Query: 354 KKLTLTT-NPIQTASGLGSHVEALLRKSNTDFHVVISQMNGKDHSS 398
+ +TL+T P+ ++GL + VEAL+ +T+ +VV++ MNG H++
Sbjct: 164 EHITLSTAGPLDPSAGLITFVEALMH*PDTELNVVVTLMNGSHHTT 301
>BF009931
Length = 235
Score = 41.2 bits (95), Expect = 0.001
Identities = 22/73 (30%), Positives = 41/73 (56%)
Frame = +2
Query: 2 TKISPEIEIKMPMEAVPPVSADVSFISDRFPKYKLGADSQILEETVEDNQGPSLKDVIEQ 61
TKI+ + + + +P + D+ S FP YK+ ++QI+ ET + +G ++ +I +
Sbjct: 2 TKITHNFKNTIQKDTIPIICTDIILTSLTFPNYKIEINNQII-ETKNNPKGVCIERMITR 178
Query: 62 EATNLSDQHKRIS 74
E T L +Q + IS
Sbjct: 179ETTQLVNQQRHIS 217
>TC228009 similar to UP|Q9LT13 (Q9LT13) Gb|AAF01580.1, partial (33%)
Length = 588
Score = 33.1 bits (74), Expect = 0.29
Identities = 14/43 (32%), Positives = 25/43 (57%)
Frame = +1
Query: 307 SKCSFQWYRLSSEGSWREVISGADKSIYAPDPFDVGRILQVDI 349
+ C+F+W R +GS+ I GA + Y + DVG +L +++
Sbjct: 31 TSCNFEWIRXXEDGSF-NYIDGAKQPTYLVNADDVGTLLAIEV 156
>TC228007 similar to UP|Q9LT13 (Q9LT13) Gb|AAF01580.1, partial (51%)
Length = 1707
Score = 31.6 bits (70), Expect = 0.84
Identities = 14/43 (32%), Positives = 25/43 (57%)
Frame = +2
Query: 307 SKCSFQWYRLSSEGSWREVISGADKSIYAPDPFDVGRILQVDI 349
+ C+F+W R +GS+ I GA + Y + DVG +L +++
Sbjct: 851 TSCNFEWIRHLEDGSF-NYIDGAKQPTYLVNADDVGTLLAIEV 976
>NP004853 nodulin
Length = 1431
Score = 30.8 bits (68), Expect = 1.4
Identities = 41/189 (21%), Positives = 86/189 (44%), Gaps = 20/189 (10%)
Frame = +1
Query: 127 RNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVK---KLLNFLKQA------SEDAKKL 177
R + ++ A + + TQ +G+ +E+ +V+ K+ ++A SE AKK
Sbjct: 580 RQGQTLQNAAKIDAETKIISTQRQGDGKKEEIKVRTEVKVFENEREAVVAEANSELAKKK 759
Query: 178 VNQEKSFACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDG---------LVEEVQEA 228
++ AE+E+A++V LR E E E+ + ++ + + +VQEA
Sbjct: 760 AVWAQTAQVAEVEAAKAVALREAELQREVERMNALTRTEKLKAEFLSKASVEYETKVQEA 939
Query: 229 R-RIKLLHQPSKVMAMEYELRA-LRDQIREKSIFSIKLQKELTMSKRDEENKSHPYMLHG 286
+ + ++ + E E A + + E + FS + + E + + +E + ++
Sbjct: 940 NWELYKKQKAAEAILFEKEKEAEAQKALAEAAFFSRQQEAEAELFAKKKEAEG---LVAL 1110
Query: 287 SEALGSYLK 295
+A G+YLK
Sbjct: 1111GQAQGAYLK 1137
>TC221509 weakly similar to GB|AAP68225.1|31711738|BT008786 At3g49240
{Arabidopsis thaliana;} , partial (23%)
Length = 806
Score = 30.8 bits (68), Expect = 1.4
Identities = 31/124 (25%), Positives = 48/124 (38%)
Frame = +2
Query: 44 EETVEDNQGPSLKDVIEQEATNLSDQHKRISVRDLACKFDKNLAAAAKLSNEAKIREVAS 103
EE E +G K+ E+E T L ++ +R+ A K++ AA A+ +
Sbjct: 317 EEFQEFVKGELRKEGREEELTKLIEEKERLKAEAKA----KDIEAAEAAKRSARAAVASL 484
Query: 104 LEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAVKLTQNEGELIQEKFEVKKL 163
L + K D+ +E S E LA K T+ EG+ E+
Sbjct: 485 LPSKLFGNKETDSESKTEGEIQALGEESANATASEAE-LAEKTTETEGDRASEQLTG*SC 661
Query: 164 LNFL 167
NFL
Sbjct: 662 CNFL 673
>TC213512
Length = 560
Score = 30.4 bits (67), Expect = 1.9
Identities = 23/73 (31%), Positives = 37/73 (50%)
Frame = +2
Query: 185 ACAEIESARSVVLRIGEALEEQEQDSQASKPQDVDGLVEEVQEARRIKLLHQPSKVMAME 244
A A++ ARSV+ IGE +E D+ +K DV+ + +E I L +P+ E
Sbjct: 305 AVAQVTQARSVLTLIGERPTHEEVDNARAKLADVEAQLS--RELEEIVLQARPA-----E 463
Query: 245 YELRALRDQIREK 257
E++ R Q E+
Sbjct: 464 IEIQGWRAQQAER 502
>BE209742
Length = 432
Score = 29.3 bits (64), Expect = 4.2
Identities = 19/74 (25%), Positives = 34/74 (45%)
Frame = +1
Query: 85 NLAAAAKLSNEAKIREVASLEGHVLLKKLRDALEYLRVRFAGRNKEDVEKAISMVEALAV 144
N+ + K R V + GH LLKK DAL G+ ++ ++K +S E++
Sbjct: 91 NVVPTVTMLGVVKARLVGATRGHALLKKKSDAL-------TGQLRQILKKIVSTKESMGD 249
Query: 145 KLTQNEGELIQEKF 158
+ + L + K+
Sbjct: 250 IMKTSSFALTEAKY 291
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.131 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,796,517
Number of Sequences: 63676
Number of extensions: 197438
Number of successful extensions: 960
Number of sequences better than 10.0: 42
Number of HSP's better than 10.0 without gapping: 950
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 953
length of query: 490
length of database: 12,639,632
effective HSP length: 101
effective length of query: 389
effective length of database: 6,208,356
effective search space: 2415050484
effective search space used: 2415050484
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0269a.3


