

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
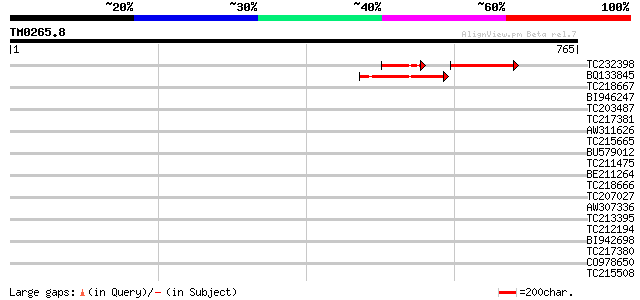
Query= TM0265.8
(765 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC232398 95 2e-28
BQ133845 102 5e-22
TC218667 similar to GB|AAP31930.1|30387529|BT006586 At3g09570 {A... 35 0.095
BI946247 33 0.36
TC203487 similar to UP|RK9_PEA (P11894) 50S ribosomal protein L9... 31 1.8
TC217381 similar to UP|O82469 (O82469) Protein phosphatase-2C , ... 31 1.8
AW311626 31 2.3
TC215665 homologue to UP|Q42792 (Q42792) Asparagine synthetase ... 31 2.3
BU579012 similar to PIR|F40201|F402 artifact-warning sequence (t... 30 3.0
TC211475 similar to UP|P79122 (P79122) Pinin, partial (5%) 30 4.0
BE211264 30 5.2
TC218666 similar to GB|AAP31930.1|30387529|BT006586 At3g09570 {A... 30 5.2
TC207027 weakly similar to UP|Q9LW95 (Q9LW95) KED, partial (17%) 30 5.2
AW307336 weakly similar to GP|22202765|dbj P0039H02.8 {Oryza sat... 29 6.8
TC213395 29 6.8
TC212194 similar to UP|Q9LK97 (Q9LK97) Exonuclease-like protein,... 29 6.8
BI942698 29 6.8
TC217380 similar to UP|O82469 (O82469) Protein phosphatase-2C , ... 29 6.8
CO978650 29 6.8
TC215508 homologue to UP|143A_SOYBN (Q96450) 14-3-3-like protein... 29 8.9
>TC232398
Length = 1054
Score = 95.1 bits (235), Expect(2) = 2e-28
Identities = 45/92 (48%), Positives = 68/92 (73%)
Frame = +1
Query: 595 RLNVGELKPTMMMLQLADRSIVTPWGVVEDVLVRVGEFEFPVDFVIIDMDEDSKIPLILG 654
R+ +++PT M LQLAD SI +GVVED+LV+V + F VDFVI+D++ED++I LILG
Sbjct: 175 RIGNQKIEPTRMTLQLADHSITRSFGVVEDILVKVHQLIFLVDFVIMDIEEDAEIRLILG 354
Query: 655 RPFLATSQAKINVGKGTISLRVADEKIGFNIF 686
PF+ T++ +++GKG + + V D+K FN+F
Sbjct: 355 WPFMVTAKCVVDMGKGNLEMSVEDQKATFNLF 450
Score = 50.1 bits (118), Expect(2) = 2e-28
Identities = 27/60 (45%), Positives = 38/60 (63%)
Frame = +2
Query: 502 LRVELPFSEVLEKMPQYAKFIKEILSKKRRLSEENEIVELTEECSVILQRKLPPKRKDPG 561
L + +PF E +++MP Y KF+K+IL KK + IV + E C ++Q KLPPK KD G
Sbjct: 2 LEITIPFGERIQQMPLYKKFLKDILIKKGKYINSETIV-VGEYCRALIQ-KLPPKFKDLG 175
>BQ133845
Length = 389
Score = 102 bits (255), Expect = 5e-22
Identities = 54/120 (45%), Positives = 78/120 (65%)
Frame = +2
Query: 473 KVPFPPFPTNIAKRRLEKQFSKFISMFKKLRVELPFSEVLEKMPQYAKFIKEILSKKRRL 532
+VP+P P+ K E+ +KF+ +FKKL + LPF E L++MP YA F+K++L+KK
Sbjct: 32 EVPYPLVPS*KDK---EQHLAKFLDIFKKLEITLPFEEALQQMPLYANFLKDMLTKKNWY 202
Query: 533 SEENEIVELTEECSVILQRKLPPKRKDPGSFTLPVNFGASKQVRALCDLGSSVNLMPLSM 592
++IV + CS ++QR LPP DPG T+P + G +AL DLG+S+NLMPLSM
Sbjct: 203 IHSDKIV-VEGNCSAVIQRILPP*HTDPGFVTMPCSIGEVAVGKALIDLGASINLMPLSM 379
>TC218667 similar to GB|AAP31930.1|30387529|BT006586 At3g09570 {Arabidopsis
thaliana;} , partial (61%)
Length = 817
Score = 35.4 bits (80), Expect = 0.095
Identities = 20/43 (46%), Positives = 28/43 (64%), Gaps = 2/43 (4%)
Frame = -3
Query: 423 TLSEPKQKIVENNNVNDEFVG--DIEPLGENKKKQKEKEEEKR 463
+L E K++ E N VN EFVG D+EP GE + +E+EE +R
Sbjct: 266 SLHEKKERH*EVNGVNGEFVGEVDVEPEGEEGEVGEEEEEVER 138
>BI946247
Length = 372
Score = 33.5 bits (75), Expect = 0.36
Identities = 14/31 (45%), Positives = 24/31 (77%)
Frame = +3
Query: 450 ENKKKQKEKEEEKRKVELEKKFTKVPFPPFP 480
+ KKK+K+K+++K+K + +KK T FPP+P
Sbjct: 114 KKKKKKKKKKKKKKKKKKKKKKTXKLFPPWP 206
Score = 28.9 bits (63), Expect = 8.9
Identities = 13/31 (41%), Positives = 22/31 (70%)
Frame = +1
Query: 450 ENKKKQKEKEEEKRKVELEKKFTKVPFPPFP 480
+ KKK+K+K+++K+K + +KK FPP P
Sbjct: 115 KKKKKKKKKKKKKKKKKKKKKKPGNCFPPGP 207
>TC203487 similar to UP|RK9_PEA (P11894) 50S ribosomal protein L9,
chloroplast precursor (CL13), partial (42%)
Length = 466
Score = 31.2 bits (69), Expect = 1.8
Identities = 13/26 (50%), Positives = 17/26 (65%)
Frame = +2
Query: 456 KEKEEEKRKVELEKKFTKVPFPPFPT 481
KE + E+ ++E EKK PFPP PT
Sbjct: 350 KEMKIEEERIEAEKKRVIFPFPPIPT 427
>TC217381 similar to UP|O82469 (O82469) Protein phosphatase-2C , partial
(35%)
Length = 742
Score = 31.2 bits (69), Expect = 1.8
Identities = 24/87 (27%), Positives = 39/87 (44%), Gaps = 4/87 (4%)
Frame = +2
Query: 314 LEGTEQLQSRRK*SEAIFKSK----IPVSEFRTSTESGSRQWQWEEELRRIDGEFH*QSI 369
L E ++S + S+A +S IP + + G R++ +E +R D H S+
Sbjct: 356 LTSQEDVKSADRISDAALESAVLQFIPCIRSGSFADIGPRRYMEDEHIRIDDLSSHLGSL 535
Query: 370 Y*FQKS*SCYKKIGDSGGTDGQANVRK 396
Y F K + Y GG + A +RK
Sbjct: 536 YNFPKPSAFYGVFDGHGGPEAAAYIRK 616
>AW311626
Length = 385
Score = 30.8 bits (68), Expect = 2.3
Identities = 14/31 (45%), Positives = 24/31 (77%), Gaps = 1/31 (3%)
Frame = +2
Query: 450 ENKKKQKEKEEEKRKVELEKKFTKVP-FPPF 479
+ KKK+K+K+++K+K + +KK K+P PPF
Sbjct: 62 KKKKKKKKKKKKKKKKKKKKKKKKIPGAPPF 154
>TC215665 homologue to UP|Q42792 (Q42792) Asparagine synthetase , complete
Length = 2492
Score = 30.8 bits (68), Expect = 2.3
Identities = 14/31 (45%), Positives = 22/31 (70%)
Frame = +1
Query: 450 ENKKKQKEKEEEKRKVELEKKFTKVPFPPFP 480
+ KKK+K+K+++K+K +KK K FPP P
Sbjct: 2176 KKKKKKKKKKKKKKKKIKKKKKKKKKFPPPP 2268
>BU579012 similar to PIR|F40201|F402 artifact-warning sequence (translated
ALU class F) - human, partial (3%)
Length = 261
Score = 30.4 bits (67), Expect = 3.0
Identities = 13/30 (43%), Positives = 22/30 (73%)
Frame = +3
Query: 450 ENKKKQKEKEEEKRKVELEKKFTKVPFPPF 479
+ KKK+K+K+++K+K + +KK K PPF
Sbjct: 21 KKKKKKKKKKKKKKKKKKKKKKKKKKTPPF 110
>TC211475 similar to UP|P79122 (P79122) Pinin, partial (5%)
Length = 436
Score = 30.0 bits (66), Expect = 4.0
Identities = 22/75 (29%), Positives = 38/75 (50%), Gaps = 2/75 (2%)
Frame = +3
Query: 411 ENCSAITLRSGATLSEPKQKIVENNNVNDEFVGDIEPLGENKKKQKE-KEEEKRKVELEK 469
++C+ + S L P + +NN N+ PL KK++K+ E+KRK+ +K
Sbjct: 42 KSCALLLWTSHPPLPLPIPRKKRSNNNNNH------PLSNTKKRKKKFPPEKKRKLPPKK 203
Query: 470 KFTKV-PFPPFPTNI 483
+ TK+ P P P +
Sbjct: 204 RKTKISPHHPSPKTL 248
>BE211264
Length = 203
Score = 29.6 bits (65), Expect = 5.2
Identities = 13/31 (41%), Positives = 22/31 (70%)
Frame = +3
Query: 450 ENKKKQKEKEEEKRKVELEKKFTKVPFPPFP 480
+ KKK+K+K+++K+K + +KKF PFP
Sbjct: 24 KKKKKKKKKKKKKKKKKKKKKFGGGAGSPFP 116
>TC218666 similar to GB|AAP31930.1|30387529|BT006586 At3g09570 {Arabidopsis
thaliana;} , partial (31%)
Length = 420
Score = 29.6 bits (65), Expect = 5.2
Identities = 30/101 (29%), Positives = 45/101 (43%), Gaps = 11/101 (10%)
Frame = -3
Query: 372 FQKS*SCYKKIGDSGGTDGQANVRKTPGMFPSDTVINPKENCSAITLRSGA----TLSEP 427
F+K SC + DS + + PS + + + SGA +L E
Sbjct: 310 FEKRPSCPYQNDDSKEQNDADKLEDVEENIPS---VREAGGLDVLLILSGAD*IQSLHEK 140
Query: 428 KQKIVENNNVNDEFVG--DIEPLGEN-----KKKQKEKEEE 461
K++ E N V+ EFVG D+EP GE K+++ E EE
Sbjct: 139 KERH*EVNAVHGEFVGVADVEPEGEEGEVG*KEEKVEGREE 17
>TC207027 weakly similar to UP|Q9LW95 (Q9LW95) KED, partial (17%)
Length = 1005
Score = 29.6 bits (65), Expect = 5.2
Identities = 28/108 (25%), Positives = 52/108 (47%), Gaps = 6/108 (5%)
Frame = +3
Query: 436 NVNDEFVGDIEPLGENKKKQK----EKEEEKRKVELEKKFTKVPFPPFPTNIAKRRLEKQ 491
+V D + +I+ GE + K K E +E+K+K + +KK K K+ K
Sbjct: 582 SVRDIDIEEIKKEGEKEDKGKDGGKEVKEKKKKEDKDKKEKK-----------KKVTGKD 728
Query: 492 FSKFISMFKKL--RVELPFSEVLEKMPQYAKFIKEILSKKRRLSEENE 537
+K +S K+ ++ +LEK + IKE+ ++ ++EEN+
Sbjct: 729 KTKDLSTLKQKLEKINGKIQPLLEKKADIERQIKEVEAEGHVVNEENK 872
>AW307336 weakly similar to GP|22202765|dbj P0039H02.8 {Oryza sativa
(japonica cultivar-group)}, partial (18%)
Length = 371
Score = 29.3 bits (64), Expect = 6.8
Identities = 21/68 (30%), Positives = 34/68 (49%), Gaps = 8/68 (11%)
Frame = +2
Query: 450 ENKKKQKEKEEEKRKVELEKKFTKVPFPPF------PTNIAKRRLEKQFSKFISMF--KK 501
+ KKK+K+K+++KRK + +KK +K+ F P + F F S KK
Sbjct: 98 KKKKKKKKKKKKKRKKKKKKKTSKLVFFTIAKKGKQPKRVKWETKSPNFKNFFSRASQKK 277
Query: 502 LRVELPFS 509
+ +PFS
Sbjct: 278 SK*IIPFS 301
>TC213395
Length = 426
Score = 29.3 bits (64), Expect = 6.8
Identities = 16/45 (35%), Positives = 29/45 (63%), Gaps = 1/45 (2%)
Frame = +2
Query: 403 SDTVINPKENCS-AITLRSGATLSEPKQKIVENNNVNDEFVGDIE 446
S ++++P N + A+T SGA + P+ + +E+ N+ND V DI+
Sbjct: 248 SPSLVSPAVNGNLAVTANSGAEVVNPEVEYIESENLND--VDDID 376
>TC212194 similar to UP|Q9LK97 (Q9LK97) Exonuclease-like protein, partial
(68%)
Length = 1015
Score = 29.3 bits (64), Expect = 6.8
Identities = 24/79 (30%), Positives = 37/79 (46%), Gaps = 3/79 (3%)
Frame = +2
Query: 655 RPFLATSQAKINVGKGTISLRVADEKIGFNIFDLKPKPVEKNDVFLV---KMMEEWSDEK 711
RPFL S ++ + +A E +G NI + PVE ++ + +EW
Sbjct: 581 RPFLNRSSSRRALR------HLAAEHLGVNIQTGEHCPVEDARAAMLLYQRNRKEWEKNI 742
Query: 712 LKQFFLKEKAGASNKKKKE 730
QF LK+K S +KKK+
Sbjct: 743 KDQFRLKQKQRKSKQKKKQ 799
>BI942698
Length = 353
Score = 29.3 bits (64), Expect = 6.8
Identities = 13/31 (41%), Positives = 22/31 (70%)
Frame = +3
Query: 450 ENKKKQKEKEEEKRKVELEKKFTKVPFPPFP 480
+ KKK+K+K+++K+K + +KK PPFP
Sbjct: 114 KKKKKKKKKKKKKKKKKKKKKPRGGARPPFP 206
>TC217380 similar to UP|O82469 (O82469) Protein phosphatase-2C , partial
(64%)
Length = 711
Score = 29.3 bits (64), Expect = 6.8
Identities = 21/83 (25%), Positives = 35/83 (41%)
Frame = +1
Query: 328 EAIFKSKIPVSEFRTSTESGSRQWQWEEELRRIDGEFH*QSIY*FQKS*SCYKKIGDSGG 387
E+ IP + + G R++ +E +R D H S+Y F + + Y GG
Sbjct: 40 ESAVLQSIPCIRSGSFADIGPRRYMEDEHIRIDDLSSHLGSLYNFPQPSAFYGVFDGHGG 219
Query: 388 TDGQANVRKTPGMFPSDTVINPK 410
+ A +RK F + V P+
Sbjct: 220 PEAAAYIRKNVTKFFFEDVNFPR 288
>CO978650
Length = 709
Score = 29.3 bits (64), Expect = 6.8
Identities = 11/27 (40%), Positives = 22/27 (80%)
Frame = +1
Query: 450 ENKKKQKEKEEEKRKVELEKKFTKVPF 476
+ KKK+K+K+++K+K + +KK + +PF
Sbjct: 25 KKKKKKKKKKKKKKKKKKKKKKSHIPF 105
>TC215508 homologue to UP|143A_SOYBN (Q96450) 14-3-3-like protein A (SGF14A),
complete
Length = 1437
Score = 28.9 bits (63), Expect = 8.9
Identities = 19/46 (41%), Positives = 26/46 (56%)
Frame = -1
Query: 452 KKKQKEKEEEKRKVELEKKFTKVPFPPFPTNIAKRRLEKQFSKFIS 497
KKK+KEKE++K LE K P ++I KR+L KFI+
Sbjct: 1197 KKKKKEKEKKK----LESLT*KKKNPTLFSSIKKRQLHSNNKKFIN 1072
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.337 0.147 0.466
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,102,854
Number of Sequences: 63676
Number of extensions: 440456
Number of successful extensions: 3455
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 3370
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3439
length of query: 765
length of database: 12,639,632
effective HSP length: 105
effective length of query: 660
effective length of database: 5,953,652
effective search space: 3929410320
effective search space used: 3929410320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0265.8


