

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0265.5
(404 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
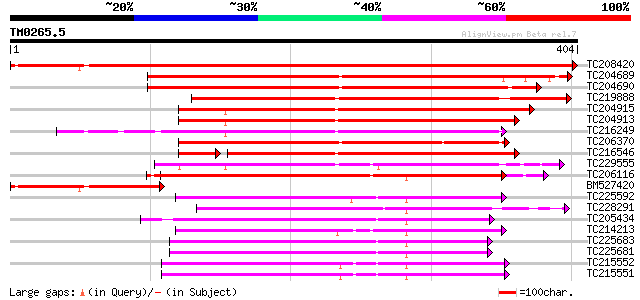
Score E
Sequences producing significant alignments: (bits) Value
TC208420 homologue to UP|Q7XXR9 (Q7XXR9) Katanin, partial (79%) 701 0.0
TC204689 homologue to UP|Q9SEA8 (Q9SEA8) Salt-induced AAA-Type A... 263 1e-70
TC204690 homologue to UP|Q9SEA8 (Q9SEA8) Salt-induced AAA-Type A... 260 7e-70
TC219888 similar to UP|Q944N4 (Q944N4) Tobacco mosaic virus heli... 225 2e-59
TC204915 weakly similar to GB|AAH46286.1|28279482|BC046286 spast... 214 4e-56
TC204913 weakly similar to UP|SAP1_YEAST (P39955) SAP1 protein, ... 212 3e-55
TC216249 weakly similar to UP|SAP1_YEAST (P39955) SAP1 protein, ... 207 7e-54
TC206370 similar to UP|Q940D1 (Q940D1) At1g64110/F22C12_22, part... 202 2e-52
TC216546 weakly similar to UP|SAP1_YEAST (P39955) SAP1 protein, ... 190 3e-51
TC229555 similar to GB|AAM62497.1|21553404|AY085265 26S proteaso... 189 1e-48
TC206116 UP|CC48_SOYBN (P54774) Cell division cycle protein 48 h... 180 1e-45
BM527420 148 4e-36
TC225592 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 ... 148 5e-36
TC228291 homologue to UP|Q9FXT8 (Q9FXT8) 26S proteasome regulato... 147 6e-36
TC205434 homologue to UP|PRS6_ARATH (Q9SEI4) 26S protease regula... 145 3e-35
TC214213 homologue to UP|PRSA_BRACM (O23894) 26S protease regula... 142 2e-34
TC225683 homologue to UP|Q9SZD4 (Q9SZD4) 26S proteasome subunit ... 140 8e-34
TC225681 homologue to UP|Q9SZD4 (Q9SZD4) 26S proteasome subunit ... 140 8e-34
TC215552 UP|PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit ... 135 3e-32
TC215551 homologue to UP|PRS7_SPIOL (Q41365) 26S protease regula... 135 3e-32
>TC208420 homologue to UP|Q7XXR9 (Q7XXR9) Katanin, partial (79%)
Length = 1523
Score = 701 bits (1810), Expect = 0.0
Identities = 357/407 (87%), Positives = 379/407 (92%), Gaps = 3/407 (0%)
Frame = +1
Query: 1 MADEEPMPTRWSFQDFKLYYDSKFGRKKVVQNGENA-DKAVGNGSSMSV--VSNGNVHSK 57
MAD+ PMPTRWSFQDFKLYYD+KFGRKKV +NG+ A DKAV NG+S++V VSNGN K
Sbjct: 97 MADD-PMPTRWSFQDFKLYYDAKFGRKKVAENGDTATDKAVSNGNSVTVTIVSNGN---K 264
Query: 58 RSSDMAIYEQLRSQGQNGIHTNDVSPNNMDERPQKSLLPPFESAEMRTLAESLSRDIIRG 117
R+S+MA+YEQ R +G N IHTN P +DERPQKSLLPPFESAEMR LAESLSRDIIRG
Sbjct: 265 RASEMAVYEQFRGEGLNQIHTNGFVPTIVDERPQKSLLPPFESAEMRALAESLSRDIIRG 444
Query: 118 SPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAV 177
SPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAV
Sbjct: 445 SPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAV 624
Query: 178 ATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARS 237
ATECKTTFFNISASSVVSKWRGDSEKLVKVLF+LARHHAPSTIFLDEIDAIISQRGEARS
Sbjct: 625 ATECKTTFFNISASSVVSKWRGDSEKLVKVLFELARHHAPSTIFLDEIDAIISQRGEARS 804
Query: 238 EHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEAR 297
EHEASRRLKTELLIQMDGLT+TDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEP AR
Sbjct: 805 EHEASRRLKTELLIQMDGLTKTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPVAR 984
Query: 298 VAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQREDLV 357
AMFEELLP QPDEE IPYD+LV++TEGYSGSDIRLLCKE AMQPLRRLMSQLEQ +D+V
Sbjct: 985 RAMFEELLPQQPDEEPIPYDILVDKTEGYSGSDIRLLCKETAMQPLRRLMSQLEQSQDVV 1164
Query: 358 PEEELPKVGPIRPEDIQAALKNTRPSAHLHAHKYDKFNADYGSQILQ 404
PEEELPKVGPI+ EDI+ AL+NTRPSAHLHAHKYDKFNADYGSQILQ
Sbjct: 1165PEEELPKVGPIKSEDIETALRNTRPSAHLHAHKYDKFNADYGSQILQ 1305
>TC204689 homologue to UP|Q9SEA8 (Q9SEA8) Salt-induced AAA-Type ATPase,
partial (98%)
Length = 1726
Score = 263 bits (672), Expect = 1e-70
Identities = 146/331 (44%), Positives = 205/331 (61%), Gaps = 28/331 (8%)
Frame = +3
Query: 99 ESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWK 158
E E L L+ IIR P+VKW + GLE+AK+ L+EAV++P+K+P++FTG PW+
Sbjct: 498 EDPEQAKLRAGLNSAIIREKPNVKWNDVAGLESAKQALQEAVILPVKFPQFFTGKRRPWR 677
Query: 159 GILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPS 218
LL+GPPGTGK+ LAKAVATE ++TFF++S+S +VSKW G+SEKLV LF++AR APS
Sbjct: 678 AFLLYGPPGTGKSYLAKAVATEAESTFFSVSSSDLVSKWMGESEKLVSNLFEMARESAPS 857
Query: 219 TIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAA 278
IF+DEID++ QRGE +E EASRR+KTELL+QM G+ D+ V VLAATN P+ LD A
Sbjct: 858 IIFIDEIDSLCGQRGEG-NESEASRRIKTELLVQMQGVGHNDQKVLVLAATNTPYALDQA 1034
Query: 279 MLRRLEKRILVPLPEPEARVAMFEELLPPQPDE-ESIPYDLLVNQTEGYSGSDIRLLCKE 337
+ RR +KRI +PLP+ +AR MF+ L P ++ L ++TEG+SGSDI + K+
Sbjct: 1035IRRRFDKRIYIPLPDLKARQHMFKVHLGDTPHNLTESDFEYLASRTEGFSGSDISVCVKD 1214
Query: 338 VAMQPLRRLMSQL----------------EQREDLVPEEELPKVG--------PIRPEDI 373
V +P+R+ + +Q +EL G PIR D
Sbjct: 1215VLFEPVRKTQDAMFFFKNPEGMWIPCGPKQQGAVQTSMQELAAKGLASKILPPPIRRTDF 1394
Query: 374 QAALKNTRPS---AHLHAHKYDKFNADYGSQ 401
+ L RP+ A L H ++F ++G +
Sbjct: 1395EKVLARQRPTVSKADLDVH--ERFTKEFGEE 1481
>TC204690 homologue to UP|Q9SEA8 (Q9SEA8) Salt-induced AAA-Type ATPase,
partial (94%)
Length = 1630
Score = 260 bits (665), Expect = 7e-70
Identities = 135/282 (47%), Positives = 190/282 (66%), Gaps = 1/282 (0%)
Frame = +2
Query: 99 ESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGLLSPWK 158
E E L L+ I+R P+VKW + GLE+AK+ L+EAV++P+K+P++FTG PW+
Sbjct: 449 EDPEQAKLRAGLNSAIVREKPNVKWNDVAGLESAKQALQEAVILPVKFPQFFTGKRRPWR 628
Query: 159 GILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPS 218
LL+GPPGTGK+ LAKAVATE +TFF++S+S +VSKW G+SEKLV LFQ+AR APS
Sbjct: 629 AFLLYGPPGTGKSYLAKAVATEADSTFFSVSSSDLVSKWMGESEKLVSNLFQMARESAPS 808
Query: 219 TIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAA 278
IF+DEID++ QRGE +E EASRR+KTELL+QM G+ D+ V VLAATN P+ LD A
Sbjct: 809 IIFVDEIDSLCGQRGEG-NESEASRRIKTELLVQMQGVGHNDQKVLVLAATNTPYALDQA 985
Query: 279 MLRRLEKRILVPLPEPEARVAMFEELLPPQPDE-ESIPYDLLVNQTEGYSGSDIRLLCKE 337
+ RR +KRI +PLP+ +AR MF+ L P ++ L +TEG+SGSDI + K+
Sbjct: 986 IRRRFDKRIYIPLPDLKARQHMFKVHLGDTPHNLAESDFEHLARKTEGFSGSDISVCVKD 1165
Query: 338 VAMQPLRRLMSQLEQREDLVPEEELPKVGPIRPEDIQAALKN 379
V +P+R+ + + PE+ GP + +Q +++
Sbjct: 1166VLFEPVRKTQDAMFFFRN--PEDMWIPCGPKQQSAVQTTMQD 1285
>TC219888 similar to UP|Q944N4 (Q944N4) Tobacco mosaic virus helicase
domain-binding protein (Fragment), partial (50%)
Length = 1119
Score = 225 bits (574), Expect = 2e-59
Identities = 128/274 (46%), Positives = 179/274 (64%), Gaps = 3/274 (1%)
Frame = +3
Query: 130 ENAKRLLKEAVVMPIKYPKYFTGLLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNIS 189
E AK+ L E V++P K FTGL P +G+LLFGPPG GKTMLAKAVA+E + TFFN++
Sbjct: 3 EKAKQALMEMVILPTKRRDLFTGLRRPARGLLLFGPPGNGKTMLAKAVASESQATFFNVT 182
Query: 190 ASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTEL 249
A+S+ SKW G++EKLV+ LF +A PS IF+DEID+I+S R +E++ASRRLK+E
Sbjct: 183 AASLTSKWVGEAEKLVRTLFMVAISRQPSVIFIDEIDSIMSTR--LANENDASRRLKSEF 356
Query: 250 LIQMDGLT-RTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQ 308
LIQ DG+T D++V V+ ATN P ELD A+LRRL KRI +PLP+ R + + L Q
Sbjct: 357 LIQFDGVTSNPDDIVIVIGATNKPQELDDAVLRRLVKRIYIPLPDENVRKLLLKHKLKGQ 536
Query: 309 P-DEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQREDLVPEEELPKVGP 367
S + LV +TEGYSGSD++ LC+E AM P+R L + + + +V
Sbjct: 537 AFSLPSRDLERLVKETEGYSGSDLQALCEEAAMMPIRELGAD-------ILTVKANQVRG 695
Query: 368 IRPEDIQAALKNTRPSAHLHA-HKYDKFNADYGS 400
+R ED + A+ RPS + + +++N ++GS
Sbjct: 696 LRYEDFKKAMATIRPSLNKSKWEELERWNEEFGS 797
>TC204915 weakly similar to GB|AAH46286.1|28279482|BC046286 spastic
paraplegia 4 homolog {Mus musculus;} , partial (21%)
Length = 1470
Score = 214 bits (546), Expect = 4e-56
Identities = 117/257 (45%), Positives = 163/257 (62%), Gaps = 3/257 (1%)
Frame = +1
Query: 121 VKWESIKGLENAKRLLKEAVVMPIKYPKYFTG--LLSPWKGILLFGPPGTGKTMLAKAVA 178
V ++ I LEN K LKE V++P++ P+ F L P KGILLFGPPGTGKTMLAKAVA
Sbjct: 271 VTFDDIGALENVKDTLKELVMLPLQRPELFCKGQLAKPCKGILLFGPPGTGKTMLAKAVA 450
Query: 179 TECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARSE 238
TE F NIS SS+ SKW G+ EK VK +F LA APS IF+DE+D+++ +R E SE
Sbjct: 451 TEAGANFINISMSSITSKWFGEGEKYVKAVFSLASKIAPSVIFVDEVDSMLGRR-ENPSE 627
Query: 239 HEASRRLKTELLIQMDGL-TRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEAR 297
HEA R++K E ++ DGL T+ E V VLAATN P++LD A++RRL +R++V LP+ R
Sbjct: 628 HEAMRKMKNEFMVNWDGLRTKDKERVLVLAATNRPFDLDEAVIRRLPRRLMVNLPDAPNR 807
Query: 298 VAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQREDLV 357
+ +L + + ++ + N T+GYSGSD++ LC A P+R ++ + ++ L
Sbjct: 808 EKILRVILVKEDLAPDVDFEAIANMTDGYSGSDLKNLCVTAAHCPIREILEKEKKERSLA 987
Query: 358 PEEELPKVGPIRPEDIQ 374
E P G DI+
Sbjct: 988 LSESKPLPGLCGSGDIR 1038
>TC204913 weakly similar to UP|SAP1_YEAST (P39955) SAP1 protein, partial
(14%)
Length = 1080
Score = 212 bits (539), Expect = 3e-55
Identities = 112/246 (45%), Positives = 159/246 (64%), Gaps = 3/246 (1%)
Frame = +3
Query: 121 VKWESIKGLENAKRLLKEAVVMPIKYPKYFTG--LLSPWKGILLFGPPGTGKTMLAKAVA 178
V ++ I LEN K LKE V++P++ P+ F L P KGILLFGPPGTGKTMLAKAVA
Sbjct: 300 VTFDDIGALENVKETLKELVMLPLQRPELFGKGQLAKPCKGILLFGPPGTGKTMLAKAVA 479
Query: 179 TECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARSE 238
TE F NIS SS+ SKW G+ EK VK +F LA APS IF+DE+D+++ +R E E
Sbjct: 480 TEAGANFINISMSSITSKWFGEGEKYVKAVFSLASKIAPSVIFVDEVDSMLGRR-ENPGE 656
Query: 239 HEASRRLKTELLIQMDGL-TRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEAR 297
HEA R++K E ++ DGL T+ E + VLAATN P++LD A++RRL +R++V LP+ R
Sbjct: 657 HEAMRKMKNEFMVNWDGLRTKDKERILVLAATNRPFDLDEAVIRRLPRRLMVNLPDAPNR 836
Query: 298 VAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQREDLV 357
+ +L + + ++ + N T+GYSGSD++ LC A P+R+++ + ++ L
Sbjct: 837 GKIVRVILAKEDLAPDVDFEAIANMTDGYSGSDLKNLCVTAAQCPIRQILEKEKKERSLA 1016
Query: 358 PEEELP 363
E P
Sbjct: 1017LAENQP 1034
>TC216249 weakly similar to UP|SAP1_YEAST (P39955) SAP1 protein, partial (14%)
Length = 2222
Score = 207 bits (527), Expect = 7e-54
Identities = 128/325 (39%), Positives = 191/325 (58%), Gaps = 4/325 (1%)
Frame = +2
Query: 34 ENADKAVGNGSSMSVVSNGNVHSKRSSDMAIYEQLRSQGQNGIHTNDVSPNNMDERPQKS 93
ENA+K +G S ++ N +K S + + + G + S N + +KS
Sbjct: 719 ENAEKIIGWALSHHLMQNSE--AKPDSKLVLSCESILYGIGILQ----SIQNESKSLKKS 880
Query: 94 LLPPFESAEMRTLAESLSRDIIRGSP-DVKWESIKGLENAKRLLKEAVVMPIKYPKYFTG 152
L E + L D+I S DV ++ I LE K LKE V++P++ P+ F
Sbjct: 881 LKDVVTENEFE---KRLLADVIPPSDIDVTFDDIGALEKVKDTLKELVMLPLQRPELFCK 1051
Query: 153 --LLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQ 210
L P KGILLFGPPGTGKTMLAKA+ATE F NIS SS+ SKW G+ EK VK +F
Sbjct: 1052 GQLTKPCKGILLFGPPGTGKTMLAKAIATEAGANFINISMSSITSKWFGEGEKYVKAVFS 1231
Query: 211 LARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGL-TRTDELVFVLAAT 269
LA +PS IF+DE+D+++ +R E EHEA R++K E ++ DGL T+ E V VLAAT
Sbjct: 1232 LASKISPSVIFVDEVDSMLGRR-ENPGEHEAMRKMKNEFMVNWDGLRTKETERVLVLAAT 1408
Query: 270 NLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGS 329
N P++LD A++RR+ +R++V LP+ R + + +L + + D + + T+GYSGS
Sbjct: 1409 NRPFDLDEAVIRRMPRRLMVNLPDAPNRAKILKVILAKEELSPDVDLDAVASMTDGYSGS 1588
Query: 330 DIRLLCKEVAMQPLRRLMSQLEQRE 354
D++ LC A +P++ ++ + E++E
Sbjct: 1589 DLKNLCVTAAHRPIKEILEK-EKKE 1660
>TC206370 similar to UP|Q940D1 (Q940D1) At1g64110/F22C12_22, partial (38%)
Length = 1502
Score = 202 bits (514), Expect = 2e-52
Identities = 111/238 (46%), Positives = 157/238 (65%), Gaps = 2/238 (0%)
Frame = +2
Query: 121 VKWESIKGLENAKRLLKEAVVMPIKYPKYFTG-LLSPWKGILLFGPPGTGKTMLAKAVAT 179
V + I L+ K L+E V++P++ P F G LL P +GILLFGPPGTGKTMLAKA+A
Sbjct: 53 VTFADIGALDEIKESLQELVMLPLRRPDLFKGGLLKPCRGILLFGPPGTGKTMLAKAIAN 232
Query: 180 ECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARSEH 239
E +F N+S S++ SKW G+ EK V+ LF LA AP+ IF+DE+D+++ QR EH
Sbjct: 233 EAGASFINVSMSTITSKWFGEDEKNVRALFTLAAKVAPTIIFVDEVDSMLGQRTRV-GEH 409
Query: 240 EASRRLKTELLIQMDG-LTRTDELVFVLAATNLPWELDAAMLRRLEKRILVPLPEPEARV 298
EA R++K E + DG LT +E + VLAATN P++LD A++RR E+RILV LP E R
Sbjct: 410 EAMRKIKNEFMTHWDGLLTGPNEQILVLAATNRPFDLDEAIIRRFERRILVGLPSVENRE 589
Query: 299 AMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQREDL 356
+ + LL + E++ + L TEGY+GSD++ LC A +P+R L+ Q E+ +D+
Sbjct: 590 MILKTLLAKE-KHENLDFKELATMTEGYTGSDLKNLCITAAYRPVRELIQQ-ERMKDM 757
>TC216546 weakly similar to UP|SAP1_YEAST (P39955) SAP1 protein, partial
(14%)
Length = 1321
Score = 190 bits (482), Expect(2) = 3e-51
Identities = 98/209 (46%), Positives = 138/209 (65%), Gaps = 1/209 (0%)
Frame = +3
Query: 156 PWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHH 215
P KGILLFGPPGTGKTMLAKAVATE F NIS SS+ SKW G+ EK VK +F LA
Sbjct: 138 PCKGILLFGPPGTGKTMLAKAVATEAGANFINISMSSITSKWFGEGEKYVKAVFSLASKI 317
Query: 216 APSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGL-TRTDELVFVLAATNLPWE 274
APS IF+DE+D+++ +R E EHEA R++K E ++ DGL T+ E V VLAATN P++
Sbjct: 318 APSVIFVDEVDSMLGRR-ENPGEHEAMRKMKNEFMVNWDGLRTKDTERVLVLAATNRPFD 494
Query: 275 LDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLL 334
LD A++RRL +R++V LP+ R + + +L + I D + + T+GYSGSD++ L
Sbjct: 495 LDEAVIRRLPRRLMVNLPDAPNRAKILKVILEKEDLSSDIDMDAIASMTDGYSGSDLKNL 674
Query: 335 CKEVAMQPLRRLMSQLEQREDLVPEEELP 363
C A +P++ ++ + ++ + E P
Sbjct: 675 CVTAAHRPIKEILEKEKKEQAAAVSEGRP 761
Score = 30.0 bits (66), Expect(2) = 3e-51
Identities = 13/30 (43%), Positives = 20/30 (66%)
Frame = +2
Query: 121 VKWESIKGLENAKRLLKEAVVMPIKYPKYF 150
V ++ I LEN K LKE V++P++ P+ F
Sbjct: 26 VTFDDIGALENVKDTLKELVMLPLQRPELF 115
>TC229555 similar to GB|AAM62497.1|21553404|AY085265 26S proteasome
regulatory particle chain RPT6-like protein {Arabidopsis
thaliana;} , partial (85%)
Length = 1155
Score = 189 bits (481), Expect = 1e-48
Identities = 118/311 (37%), Positives = 179/311 (56%), Gaps = 19/311 (6%)
Frame = +2
Query: 104 RTLAESLSRDIIRGSP---------------DVKWESIKGLENAKRLLKEAVVMPIKYPK 148
+ +A+ L R +I+ +P DV++ SI GLE K+ L E V++P+K P
Sbjct: 137 KEIAKRLGRPLIQTNPYEDVIACDIINPDHIDVEFNSIGGLETIKQALFELVILPLKRPD 316
Query: 149 YFTG--LLSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVK 206
F+ LL P KG+LL+GPPGTGKTMLAKA+A E + F N+ S+++SKW GD++KLV
Sbjct: 317 LFSHGKLLGPQKGVLLYGPPGTGKTMLAKAIAKESRAVFINVRISNLMSKWFGDAQKLVA 496
Query: 207 VLFQLARHHAPSTIFLDEIDAIISQRGEARSEHEASRRLKTELLIQMDGLTRTDE--LVF 264
+F LA P+ IF+DE+D+ + +R ++HEA +KTE + DG T TD+ V
Sbjct: 497 AVFSLAYKLQPAIIFIDEVDSFLGKR--RGTDHEAMLNMKTEFMALWDGFT-TDQNAQVM 667
Query: 265 VLAATNLPWELDAAMLRRLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTE 324
VLAATN P ELD A+LRRL + + +P+ R + + +L + E++I + + E
Sbjct: 668 VLAATNRPSELDEAILRRLPQAFEIGIPDQRERAEILKVVLKGERVEDNIDFGHIAGLCE 847
Query: 325 GYSGSDIRLLCKEVAMQPLRRLMSQLEQREDLVPEEELPKVGPIRPEDIQAALKNTRPSA 384
GY+GSD+ LCK+ A P+R L+ + E + P+ P+ D + AL T
Sbjct: 848 GYTGSDLFDLCKKAAYFPIRELLDE----EKKGKQSHAPR--PLSQLDFEKALA-TSKKT 1006
Query: 385 HLHAHKYDKFN 395
+ A +Y F+
Sbjct: 1007KVAASEYGGFS 1039
>TC206116 UP|CC48_SOYBN (P54774) Cell division cycle protein 48 homolog
(Valosin containing protein homolog) (VCP), complete
Length = 2583
Score = 180 bits (456), Expect = 1e-45
Identities = 98/261 (37%), Positives = 160/261 (60%), Gaps = 4/261 (1%)
Frame = +1
Query: 98 FESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSP 156
F++A + T S R+ + P+V WE I GLEN KR L+E V P+++P+ F +SP
Sbjct: 1387 FQTA-LGTSNPSALRETVVEVPNVSWEDIGGLENVKRELQETVQYPVEHPEKFEKFGMSP 1563
Query: 157 WKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHA 216
KG+L +GPPG GKT+LAKA+A EC+ F ++ +++ W G+SE V+ +F AR A
Sbjct: 1564 SKGVLFYGPPGCGKTLLAKAIANECQANFISVKGPELLTMWFGESEANVREIFDKARQSA 1743
Query: 217 PSTIFLDEIDAIISQRGEARSE-HEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWEL 275
P +F DE+D+I +QRG + + A+ R+ +LL +MDG++ + VF++ ATN P +
Sbjct: 1744 PCVLFFDELDSIATQRGSSVGDAGGAADRVLNQLLTEMDGMS-AKKTVFIIGATNRPDII 1920
Query: 276 DAAMLR--RLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRL 333
D A+LR RL++ I +PLP+ ++R +F+ L P +++ L T+G+SG+DI
Sbjct: 1921 DPALLRPGRLDQLIYIPLPDEDSRHQIFKACLRKSPIAKNVDLRALARHTQGFSGADITE 2100
Query: 334 LCKEVAMQPLRRLMSQLEQRE 354
+C+ +R + + +RE
Sbjct: 2101 ICQRACKYAIRENIEKDIERE 2163
Score = 155 bits (391), Expect = 4e-38
Identities = 93/280 (33%), Positives = 156/280 (55%), Gaps = 3/280 (1%)
Frame = +1
Query: 108 ESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPP 166
E L R+ +V ++ + G+ ++E V +P+++P+ F + + P KGILL+GPP
Sbjct: 595 EPLKREDEERLDEVGYDDVGGVRKQMAQIRELVELPLRHPQLFKSIGVKPPKGILLYGPP 774
Query: 167 GTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEID 226
G+GKT++A+AVA E FF I+ ++SK G+SE ++ F+ A +APS IF+DEID
Sbjct: 775 GSGKTLIARAVANETGAFFFCINGPEIMSKLAGESESNLRKAFEEAEKNAPSIIFIDEID 954
Query: 227 AIISQRGEARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RLE 284
+I +R + E E RR+ ++LL MDGL ++ V V+ ATN P +D A+ R R +
Sbjct: 955 SIAPKREKTHGEVE--RRIVSQLLTLMDGL-KSRAHVIVIGATNRPNSIDPALRRFGRFD 1125
Query: 285 KRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLR 344
+ I + +P+ R+ + + + + + T GY G+D+ LC E A+Q +R
Sbjct: 1126REIDIGVPDEVGRLEVLRIHTKNMKLSDDVDLERIAKDTHGYVGADLAALCTEAALQCIR 1305
Query: 345 RLMSQLEQREDLVPEEELPKVGPIRPEDIQAALKNTRPSA 384
M ++ ++ + E L + + E Q AL + PSA
Sbjct: 1306EKMDVIDLEDETIDAEVLNSMA-VTNEHFQTALGTSNPSA 1422
>BM527420
Length = 420
Score = 148 bits (374), Expect = 4e-36
Identities = 81/115 (70%), Positives = 91/115 (78%), Gaps = 5/115 (4%)
Frame = +3
Query: 1 MADEEPMPTRWSFQDFKLYYDSKFGRKKVVQNGENA-DKAVGNGSSMSV----VSNGNVH 55
MAD+ PMPTRWSFQDFKL YD+KFGRKKV +NG++A KAV NG+ SV VSNGN
Sbjct: 87 MADD-PMPTRWSFQDFKLCYDAKFGRKKVAENGDDAAGKAVSNGNGNSVTVAIVSNGN-- 257
Query: 56 SKRSSDMAIYEQLRSQGQNGIHTNDVSPNNMDERPQKSLLPPFESAEMRTLAESL 110
KR+S+MA+YEQ RS+GQN IHTN P DERPQKSLLPPFESAEMR LAESL
Sbjct: 258 -KRASEMAVYEQFRSEGQNQIHTNGFVPTLTDERPQKSLLPPFESAEMRALAESL 419
>TC225592 homologue to GB|AAP78936.1|32306505|BT009668 At5g19990 {Arabidopsis
thaliana;} , partial (94%)
Length = 1612
Score = 148 bits (373), Expect = 5e-36
Identities = 91/241 (37%), Positives = 142/241 (58%), Gaps = 5/241 (2%)
Frame = +3
Query: 119 PDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTMLAKAV 177
PD ++ I GL+ + +KE + +PIK+P+ F L ++ KG+LL+GPPGTGKT+LA+AV
Sbjct: 564 PDSTYDMIGGLDQQIKEIKEVIELPIKHPELFESLGIAQPKGVLLYGPPGTGKTLLARAV 743
Query: 178 ATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRGEARS 237
A TF +S S +V K+ G+ ++V+ LF +AR HAPS IF+DEID+I S R E+ S
Sbjct: 744 AHHTDCTFIRVSGSELVQKYIGEGSRMVRELFVMAREHAPSIIFMDEIDSIGSARMESGS 923
Query: 238 EHEAS--RRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RLEKRILVPLPE 293
+ S +R ELL Q+DG +++ + VL ATN LD A+LR R++++I P P
Sbjct: 924 GNGDSEVQRTMLELLNQLDGFEASNK-IKVLMATNRIDILDQALLRPGRIDRKIEFPNPN 1100
Query: 294 PEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQR 353
E+R+ + + I + + G SG++++ +C E M LR + Q
Sbjct: 1101EESRLDILKIHSRRMNLMRGIDLKKIAEKMNGASGAELKAVCTEAGMFALRERRVHVTQE 1280
Query: 354 E 354
+
Sbjct: 1281D 1283
>TC228291 homologue to UP|Q9FXT8 (Q9FXT8) 26S proteasome regulatory particle
triple-A ATPase subunit4, partial (62%)
Length = 1079
Score = 147 bits (372), Expect = 6e-36
Identities = 90/270 (33%), Positives = 153/270 (56%), Gaps = 4/270 (1%)
Frame = +3
Query: 134 RLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASS 192
R L+E++ +P+ P+ F + + P KG+LL+GPPGTGKT+LA+A+A+ + F + +S+
Sbjct: 6 RELRESIELPLMNPELFIRVGIKPPKGVLLYGPPGTGKTLLARAIASNIEANFLKVVSSA 185
Query: 193 VVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQR-GEARSEHEASRRLKTELLI 251
++ K+ G+S +L++ +F AR H P IF+DEIDAI +R E S +R ELL
Sbjct: 186 IIDKYIGESARLIREMFGYARDHQPCIIFMDEIDAIGGRRFSEGTSADREIQRTLMELLN 365
Query: 252 QMDGLTRTDELVFVLAATNLPWELDAAMLR--RLEKRILVPLPEPEARVAMFEELLPPQP 309
Q+DG + ++ ++ ATN P LD A+LR RL+++I +PLP ++R+ + +
Sbjct: 366 QLDGFDQLGKVKMIM-ATNRPDVLDPALLRPGRLDRKIEIPLPNEQSRMEILKIHAAGIA 542
Query: 310 DEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQREDLVPEEELPKVGPIR 369
I Y+ +V EG++G+D+R +C E M +R +R+ ++ E+ + V +
Sbjct: 543 KHGEIDYEAVVKLAEGFNGADLRNVCTEAGMAAIR------AERDYVIHEDFMKAVRKLN 704
Query: 370 PEDIQAALKNTRPSAHLHAHKYDKFNADYG 399
K SAH ++AD+G
Sbjct: 705 D------AKKLESSAH--------YSADFG 752
>TC205434 homologue to UP|PRS6_ARATH (Q9SEI4) 26S protease regulatory subunit
6B homolog (26S proteasome AAA-ATPase subunit RPT3)
(Regulatory particle triple-A ATPase subunit 3), partial
(96%)
Length = 1573
Score = 145 bits (366), Expect = 3e-35
Identities = 87/256 (33%), Positives = 149/256 (57%), Gaps = 4/256 (1%)
Frame = +1
Query: 94 LLPPFESAEMRTLAESLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL 153
+LPP + + L++S PDV + I G + K+ ++EAV +P+ + + + +
Sbjct: 508 VLPPEADSSISLLSQS-------EKPDVTYNDIGGCDIQKQEIREAVELPLTHHELYKQI 666
Query: 154 -LSPWKGILLFGPPGTGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLA 212
+ P +G+LL+GPPGTGKTMLAKAVA F + S V K+ G+ ++V+ +F+LA
Sbjct: 667 GIDPPRGVLLYGPPGTGKTMLAKAVANHTTAAFIRVVGSEFVQKYLGEGPRMVRDVFRLA 846
Query: 213 RHHAPSTIFLDEIDAIISQRGEARSEHEAS-RRLKTELLIQMDGLTRTDELVFVLAATNL 271
+ +AP+ IF+DE+DAI + R +A++ + +R+ ELL QMDG +T V V+ ATN
Sbjct: 847 KENAPAIIFIDEVDAIATARFDAQTGADREVQRILMELLNQMDGFDQTVN-VKVIMATNR 1023
Query: 272 PWELDAAMLR--RLEKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGS 329
LD A+LR RL+++I PLP+ + +F+ + + + V++ + S +
Sbjct: 1024ADTLDPALLRPGRLDRKIEFPLPDRRQKRLVFQVCTAKMNLSDEVDLEDYVSRPDKISAA 1203
Query: 330 DIRLLCKEVAMQPLRR 345
+I +C+E M +R+
Sbjct: 1204EIAAICQEAGMHAVRK 1251
>TC214213 homologue to UP|PRSA_BRACM (O23894) 26S protease regulatory subunit
6A homolog (TAT-binding protein homolog 1) (TBP-1),
partial (64%)
Length = 1086
Score = 142 bits (359), Expect = 2e-34
Identities = 86/241 (35%), Positives = 139/241 (56%), Gaps = 5/241 (2%)
Frame = +1
Query: 119 PDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTMLAKAV 177
P + I GLE + L EA+V+P+ + + F L + P KG+LL+GPPGTGKT++A+A
Sbjct: 34 PTEDYNDIGGLEKQIQELVEAIVLPMTHKERFQKLGVRPPKGVLLYGPPGTGKTLMARAC 213
Query: 178 ATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQR--GEA 235
A + TF ++ +V + GD KLV+ FQLA+ +P IF+DEIDAI ++R E
Sbjct: 214 AAQTNATFLKLAGPQLVQMFIGDGAKLVRDAFQLAKEKSPCIIFIDEIDAIGTKRFDSEV 393
Query: 236 RSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RLEKRILVPLPE 293
+ E R + ELL Q+DG + +D+ + V+AATN LD A++R RL+++I P P
Sbjct: 394 SGDREVQRTM-LELLNQLDGFS-SDDRIKVIAATNRADILDPALMRSGRLDRKIEFPHPS 567
Query: 294 PEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLRRLMSQLEQR 353
EAR + + + ++ L T+ ++G+ ++ +C E M LRR +++
Sbjct: 568 EEARARILQIHSRKMNVHPDVNFEELARSTDDFNGAQLKAVCVEAGMLALRRDATEVNHE 747
Query: 354 E 354
+
Sbjct: 748 D 750
>TC225683 homologue to UP|Q9SZD4 (Q9SZD4) 26S proteasome subunit 4-like
protein (26S proteasome subunit AtRPT2a), partial (97%)
Length = 1513
Score = 140 bits (354), Expect = 8e-34
Identities = 84/234 (35%), Positives = 137/234 (57%), Gaps = 4/234 (1%)
Frame = +2
Query: 115 IRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTML 173
+ +P + I GL+ + +KEAV +P+ +P+ + + + P KG++L+G PGTGKT+L
Sbjct: 545 VEKAPLESYADIGGLDAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGTGKTLL 724
Query: 174 AKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRG 233
AKAVA TF + S ++ K+ GD KLV+ LF++A +PS +F+DEIDA+ ++R
Sbjct: 725 AKAVANSTSATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPSIVFIDEIDAVGTKRY 904
Query: 234 EARSEHEAS-RRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RLEKRILVP 290
+A S E +R ELL Q+DG + V V+ ATN LD A+LR R++++I P
Sbjct: 905 DAHSGGEREIQRTMLELLNQLDGFDSRGD-VKVILATNRIESLDPALLRPGRIDRKIEFP 1081
Query: 291 LPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLR 344
LP+ + R +F+ + + + V + +SG+DI+ +C E + LR
Sbjct: 1082LPDIKTRRRIFQIHTSRMTLADDVNLEEFVMTKDEFSGADIKAICTEAGLLALR 1243
>TC225681 homologue to UP|Q9SZD4 (Q9SZD4) 26S proteasome subunit 4-like
protein (26S proteasome subunit AtRPT2a), partial (93%)
Length = 1433
Score = 140 bits (354), Expect = 8e-34
Identities = 84/234 (35%), Positives = 137/234 (57%), Gaps = 4/234 (1%)
Frame = +1
Query: 115 IRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPGTGKTML 173
+ +P + I GL+ + +KEAV +P+ +P+ + + + P KG++L+G PGTGKT+L
Sbjct: 442 VEKAPLESYADIGGLDAQIQEIKEAVELPLTHPELYEDIGIKPPKGVILYGEPGTGKTLL 621
Query: 174 AKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDAIISQRG 233
AKAVA TF + S ++ K+ GD KLV+ LF++A +PS +F+DEIDA+ ++R
Sbjct: 622 AKAVANSTSATFLRVVGSELIQKYLGDGPKLVRELFRVADDLSPSIVFIDEIDAVGTKRY 801
Query: 234 EARSEHEAS-RRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RLEKRILVP 290
+A S E +R ELL Q+DG + V V+ ATN LD A+LR R++++I P
Sbjct: 802 DAHSGGEREIQRTMLELLNQLDGFDSRGD-VKVILATNRIESLDPALLRPGRIDRKIEFP 978
Query: 291 LPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPLR 344
LP+ + R +F+ + + + V + +SG+DI+ +C E + LR
Sbjct: 979 LPDIKTRRRIFQIHTSRMTLADDVNLEEFVMTKDEFSGADIKAICTEAGLLALR 1140
>TC215552 UP|PRS7_ARATH (Q9SSB5) 26S protease regulatory subunit 7 (26S
proteasome subunit 7) (26S proteasome AAA-ATPase subunit
RPT1a) (Regulatory particle triple-A ATPase subunit 1a),
partial (73%)
Length = 1082
Score = 135 bits (341), Expect = 3e-32
Identities = 83/253 (32%), Positives = 137/253 (53%), Gaps = 5/253 (1%)
Frame = +1
Query: 109 SLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPG 167
S++ + PDV + + G + ++E V +P+ +P+ F L + P KG+L +GPPG
Sbjct: 106 SVTMMTVEEKPDVTYNDVGGCKEQIEKMREVVELPMLHPEKFVKLGIDPPKGVLCYGPPG 285
Query: 168 TGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDA 227
TGKT+LA+AVA F + S +V K+ G+ ++V+ LFQ+AR +F DE+DA
Sbjct: 286 TGKTLLARAVANRTDACFIRVIGSELVQKYVGEGARMVRELFQMARSKKACIVFFDEVDA 465
Query: 228 IISQRGE--ARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RL 283
I R + ++E R + E++ Q+DG + VL ATN P LD A+LR RL
Sbjct: 466 IGGARFDDGVGGDNEVQRTM-LEIVNQLDGFDARGN-IKVLMATNRPDTLDPALLRPGRL 639
Query: 284 EKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPL 343
++++ LP+ E+R +F+ E I ++LL +G+DIR +C E M +
Sbjct: 640 DRKVEFGLPDLESRTQIFKIHTRTMNCERDIRFELLARLCPNSTGADIRSVCTEAGMYAI 819
Query: 344 RRLMSQLEQREDL 356
R + +++ L
Sbjct: 820 RARRKTVTEKDFL 858
>TC215551 homologue to UP|PRS7_SPIOL (Q41365) 26S protease regulatory subunit
7 (26S proteasome subunit 7) (26S proteasome AAA-ATPase
subunit RPT1) (Regulatory particle triple-A ATPase
subunit 1), complete
Length = 1561
Score = 135 bits (341), Expect = 3e-32
Identities = 83/253 (32%), Positives = 137/253 (53%), Gaps = 5/253 (1%)
Frame = +3
Query: 109 SLSRDIIRGSPDVKWESIKGLENAKRLLKEAVVMPIKYPKYFTGL-LSPWKGILLFGPPG 167
S++ + PDV + + G + ++E V +P+ +P+ F L + P KG+L +GPPG
Sbjct: 513 SVTMMTVEEKPDVTYNDVGGCKEQIEKMREVVELPMLHPEKFVKLGIDPPKGVLCYGPPG 692
Query: 168 TGKTMLAKAVATECKTTFFNISASSVVSKWRGDSEKLVKVLFQLARHHAPSTIFLDEIDA 227
TGKT+LA+AVA F + S +V K+ G+ ++V+ LFQ+AR +F DE+DA
Sbjct: 693 TGKTLLARAVANRTDACFIRVIGSELVQKYVGEGARMVRELFQMARSKKACIVFFDEVDA 872
Query: 228 IISQRGE--ARSEHEASRRLKTELLIQMDGLTRTDELVFVLAATNLPWELDAAMLR--RL 283
I R + ++E R + E++ Q+DG + VL ATN P LD A+LR RL
Sbjct: 873 IGGARFDDGVGGDNEVQRTM-LEIVNQLDGFDARGN-IKVLMATNRPDTLDPALLRPGRL 1046
Query: 284 EKRILVPLPEPEARVAMFEELLPPQPDEESIPYDLLVNQTEGYSGSDIRLLCKEVAMQPL 343
++++ LP+ E+R +F+ E I ++LL +G+DIR +C E M +
Sbjct: 1047DRKVEFGLPDLESRTQIFKIHTRTMNCERDIRFELLARLCPNSTGADIRSVCTEAGMYAI 1226
Query: 344 RRLMSQLEQREDL 356
R + +++ L
Sbjct: 1227RARRKTVTEKDFL 1265
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.133 0.381
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,918,444
Number of Sequences: 63676
Number of extensions: 202797
Number of successful extensions: 1202
Number of sequences better than 10.0: 125
Number of HSP's better than 10.0 without gapping: 1115
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1122
length of query: 404
length of database: 12,639,632
effective HSP length: 99
effective length of query: 305
effective length of database: 6,335,708
effective search space: 1932390940
effective search space used: 1932390940
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0265.5


