

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0265.11
(1543 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
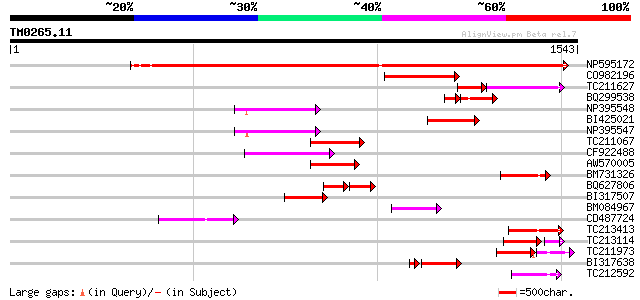
Score E
Sequences producing significant alignments: (bits) Value
NP595172 polyprotein [Glycine max] 1086 0.0
CO982196 210 4e-54
TC211627 122 2e-51
BQ299538 127 3e-43
NP395548 reverse transcriptase [Glycine max] 172 8e-43
BI425021 167 3e-41
NP395547 reverse transcriptase [Glycine max] 163 5e-40
TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%) 155 1e-37
CF922488 138 2e-32
AW570005 137 5e-32
BM731326 weakly similar to GP|21740635|em OSJNBb0043H09.2 {Oryza... 130 3e-30
BQ627806 75 1e-29
BI317507 128 2e-29
BM084967 122 1e-27
CD487724 121 3e-27
TC213413 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, parti... 120 6e-27
TC213114 weakly similar to UP|Q8W150 (Q8W150) Polyprotein, parti... 101 3e-25
TC211973 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, parti... 114 4e-25
BI317638 weakly similar to GP|9294238|dbj| contains similarity t... 100 2e-24
TC212592 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, parti... 104 3e-22
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 1086 bits (2808), Expect = 0.0
Identities = 563/1198 (46%), Positives = 769/1198 (63%), Gaps = 6/1198 (0%)
Frame = +1
Query: 330 KISPTLHKCPERAMRVLILGDGETMDEDGEIVMLEEEAEEIEEEETVECKLMGVLGCMGE 389
K SP HKCP R + +L L + + D+ E VM+ EEA ++ + M G
Sbjct: 997 KFSPA-HKCPNRQVMLLQLEETDE-DQTDEQVMVTEEANMDDDTHHLSLNAM-----RGS 1155
Query: 390 HQTQTMKLAGQVANVELLILIDSGASHNFISPKITSALGLVITPITTRSIKLGDGHKVVT 449
+ T++ GQV + + IL+D G+S NFI P++ L L + P + +G+G +
Sbjct: 1156 NGVGTIRFTGQVGGIAVKILVDGGSSDNFIQPRVAQVLKLPVEPAPNLRVLVGNGQILSA 1335
Query: 450 RGICRGIKAKLGSIEITIDALVLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQV 509
GI + + + E+ + +L++ G D++LG +WL+TLG + D+ LT++F ++
Sbjct: 1336 EGIVQQLPLHIQGQEVKVPVYLLQISGADVILGSTWLATLGPHVADYAALTLKFFQNDKF 1515
Query: 510 VQLQGMRGKDSYQSNLHSFLSDTHGREQMEWWGPQL---QVVEATKKG--HEIPQELRAV 564
+ LQG ++ Q+ LH F + + E + QL +V E T K I EL +
Sbjct: 1516 ITLQGEGNSEATQAQLHHFRRLQNTKSIEECFAIQLIQKEVPEDTLKDLPTNIDPELAIL 1695
Query: 565 LADFHEVFTKKIQLPPVRSRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLEAGI 624
L + +VF LPP R + H I L GP+ VRPYRYPH QK++IE+ + E+L GI
Sbjct: 1696 LHTYAQVFAVPASLPPQREQDHAIPLKQGSGPVKVRPYRYPHTQKDQIEKMIQEMLVQGI 1875
Query: 625 IRPSMSSFSSPVILVKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSK 684
I+PS S FS P++LVKKKD SWR C DYRALN T+ D +P+P VDELLDEL+GA FSK
Sbjct: 1876 IQPSNSPFSLPILLVKKKDGSWRFCTDYRALNAITVKDSFPMPTVDELLDELHGAQYFSK 2055
Query: 685 IDLKSGYHQIRVHEEDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKF 744
+DL+SGYHQI V ED KT FRTH+GHYE+LVMPFGL NAPATFQ +MN IF+ LRKF
Sbjct: 2056 LDLRSGYHQILVQPEDREKTAFRTHHGHYEWLVMPFGLTNAPATFQCLMNKIFQFALRKF 2235
Query: 745 VLVFFDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGV 804
VLVFFDDILIYS S +H HL+ VL L + A SKC FG ++DYLGH +SG GV
Sbjct: 2236 VLVFFDDILIYSASWKDHLKHLESVLQTLKQHQLFARLSKCSFGDTEVDYLGHKVSGLGV 2415
Query: 805 SVDPEKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWGEE 864
S++ KVQ +++WP P NVK +RGFLGLTGYYR+FIK+Y +A PLT+L +KD+F W E
Sbjct: 2416 SMENTKVQAVLDWPTPNNVKQLRGFLGLTGYYRRFIKSYANIAGPLTDLLQKDSFLWNNE 2595
Query: 865 AKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGN 924
A+ AF KLK MT +PVL+LPDF + F +E DA+G G+GAVL Q PIAYFSK L+
Sbjct: 2596 AEAAFVKLKKAMTEAPVLSLPDFSQPFILETDASGIGVGAVLGQNGHPIAYFSKKLAPRM 2775
Query: 925 LTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLG 984
+S Y +EL+A+ + +RHYLLG F + TD +SLK + Q + +P+QQ WL K LG
Sbjct: 2776 QKQSAYTRELLAITEALSKFRHYLLGNKFIIRTDQRSLKSLMDQSLQTPEQQAWLHKFLG 2955
Query: 985 YQFEVKYKPGQENKAADALSRRGDESDFFSLVSFPLWDDRKKLLEEIASDPYIKNLLQDV 1044
Y F+++YKPG++N+AADALSR F S P ++L + SDP++K L++
Sbjct: 2956 YDFKIEYKPGKDNQAADALSRM-----FMLAWSEPHSIFLEELRARLISDPHLKQLMETY 3120
Query: 1045 QQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYRRLAES 1104
+Q + ++V +G+L ++ R+V+ + +LQEYH SP GGH+G RT RL
Sbjct: 3121 KQGADAS-HYTVREGLLYWKDRVVIPAEEEIVNKILQEYHSSPIGGHAGITRTLARLKAQ 3297
Query: 1105 LYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSKG 1164
YW MQ V+ +++ C CQ+ K + P GLLQP P+P VWED+++DFI GLP S G
Sbjct: 3298 FYWPKMQEDVKAYIQKCLICQQAKSNNTLPAGLLQPLPIPQQVWEDVAMDFITGLPNSFG 3477
Query: 1165 YEAILVVVDRLSKYSHFILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFW 1224
I+VV+DRL+KY+HFI LK Y +K +AE F+ IV+LHGIP ++ SDRD +F S FW
Sbjct: 3478 LSVIMVVIDRLTKYAHFIPLKADYNSKVVAEAFMSHIVKLHGIPRSIVSDRDRVFTSTFW 3657
Query: 1225 SELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNS 1284
LFKLQGT L MSS+YHP++DGQ+EV+N+CLE YLRCF +HPK W +PWAEFWYN+
Sbjct: 3658 QHLFKLQGTTLAMSSAYHPQSDGQSEVLNKCLEMYLRCFTYEHPKGWVKALPWAEFWYNT 3837
Query: 1285 TFHCSIGQTPFEVVYGRKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQM 1344
+H S+G TPF +YGR+ P + R + V +L++RD L +L+ +L RAQ+ M
Sbjct: 3838 AYHMSLGMTPFRALYGREPPTLTRQACSIDDPAEVREQLTDRDALLAKLKINLTRAQQVM 4017
Query: 1345 ASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKL 1404
A+KKR D+SF IG+ V +KL+P+RQHS V R NQKL+ R++GPF++ KIG VAYKL
Sbjct: 4018 KRQADKKRLDVSFQIGDEVLVKLQPYRQHSAVLRKNQKLSMRYFGPFKVLAKIGDVAYKL 4197
Query: 1405 KLPVESRIHPVFHVSLLKRAIGNYQVQGQLPTDLGIEEATDI-YPEAILGTRIIRQGDSE 1463
+LP +RIHPVFHVS LK G Q LP L + E + P IL +RII +G ++
Sbjct: 4198 ELPSAARIHPVFHVSQLKPFNGTAQ-DPYLPLPLTVTEMGPVMQPVKILASRIIIRGHNQ 4374
Query: 1464 VHQSLIKWKNRSIDDVTWEDNEVLAGQFPEFCLEDKAVSKEGGIERDSNTKWGMGLGK 1521
+ Q L++W+N D+ TWED E + +P F LEDK V K G N GM G+
Sbjct: 4375 IEQILVQWENGLQDEATWEDIEDIKASYPTFNLEDKVVFKGEG-----NVTNGMSRGE 4533
>CO982196
Length = 812
Score = 210 bits (534), Expect = 4e-54
Identities = 101/206 (49%), Positives = 141/206 (68%), Gaps = 3/206 (1%)
Frame = +1
Query: 1021 WDDRKKLL---EEIASDPYIKNLLQDVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIP 1077
W R++L EEI + + + Q + KPG+++ G L ++ RLVLS S IP
Sbjct: 193 WCKRRELADWEEEIQAYLELYEIYQGILTKTTKKPGYAIRGGKLYFKDRLVLSKNSTKIP 372
Query: 1078 WLLQEYHGSPTGGHSGFLRTYRRLAESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGL 1137
LL+E SP GGHSGF RT++R+A ++W GM++T RD+V AC+ C+R K S +SP GL
Sbjct: 373 LLLKELQDSPLGGHSGFFRTFKRVANVVFWQGMKKTTRDYVAACEICRRNKTSTLSPAGL 552
Query: 1138 LQPFPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVF 1197
L P+P VW D+S+DFI GLPK++G + ILVVVDRL+KY+HF L HPYTAK +AE+F
Sbjct: 553 L*LLPIPTKVWTDISMDFIGGLPKAQGKDNILVVVDRLTKYAHFFALSHPYTAKEVAELF 732
Query: 1198 VREIVRLHGIPNTVKSDRDPIFVSHF 1223
++E+VRLHG P ++ SD +F+S F
Sbjct: 733 IKELVRLHGFPASIVSDXXRLFMSLF 810
>TC211627
Length = 1034
Score = 122 bits (307), Expect(2) = 2e-51
Identities = 74/212 (34%), Positives = 111/212 (51%)
Frame = +3
Query: 1297 VVYGRKAPPIIRFLTNETKVVAVALELSERDEALKQLRTHLQRAQEQMASYANKKRRDLS 1356
V+YG+ P + + + V AV L L LQ+ Q+ M A+ RRDL+
Sbjct: 240 VMYGKPPPALPLYSAGTSTVEAVDAILHSLATIHHTLTCRLQKYQDSMKRIADSHRRDLT 419
Query: 1357 FDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVESRIHPVF 1416
F+IG+WV+++L P+RQ S+ KL+ RFYGP+QI+ ++G VAY+L+LP S+IHP+F
Sbjct: 420 FNIGDWVYVRL*PYRQTSIQSTYT-KLSKRFYGPYQIQARVGQVAYRLQLPPTSKIHPIF 596
Query: 1417 HVSLLKRAIGNYQVQGQLPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSI 1476
HVSLLK G + + P L ++ + Q L++W N +
Sbjct: 597 HVSLLKVHHGPIPPELLALPPFSTTNHPLVQPLQFLDWKMDESTTPPIPQVLVQWTNLAP 776
Query: 1477 DDVTWEDNEVLAGQFPEFCLEDKAVSKEGGIE 1508
+D TWE L + LEDK + GGI+
Sbjct: 777 EDTTWESWTQLKDIYD---LEDKVCFQTGGID 863
Score = 100 bits (248), Expect(2) = 2e-51
Identities = 45/79 (56%), Positives = 57/79 (71%)
Frame = +2
Query: 1218 IFVSHFWSELFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPW 1277
IF+S W ELF + GTKL+ S++YHP+TDGQTEVINR LE YLR F DHP+ W ++
Sbjct: 2 IFISGLWHELFHISGTKLRFSTAYHPQTDGQTEVINRILEQYLRAFVHDHPQHWFKFLSL 181
Query: 1278 AEFWYNSTFHCSIGQTPFE 1296
AE YN++ H IG +PFE
Sbjct: 182 AE*CYNTSVHSGIGFSPFE 238
>BQ299538
Length = 426
Score = 127 bits (319), Expect(2) = 3e-43
Identities = 60/101 (59%), Positives = 72/101 (70%)
Frame = +3
Query: 1227 LFKLQGTKLKMSSSYHPETDGQTEVINRCLESYLRCFASDHPKTWSFWVPWAEFWYNSTF 1286
+ KL GT LKMS+SYHP DGQT +N CLE++LRCF +D PK W+ WAE+WYN+ F
Sbjct: 129 IVKLPGTYLKMSTSYHP*IDGQT--VNHCLETFLRCFVADQPKM*VQWLSWAEYWYNTNF 302
Query: 1287 HCSIGQTPFEVVYGRKAPPIIRFLTNETKVVAVALELSERD 1327
H S G TPFEVVYGRK P + RFL E +V AV EL +RD
Sbjct: 303 HASTGTTPFEVVYGRKPPVLNRFLPGEVRVEAVRRELQDRD 425
Score = 68.2 bits (165), Expect(2) = 3e-43
Identities = 28/44 (63%), Positives = 36/44 (81%)
Frame = +1
Query: 1184 LKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSEL 1227
LKHPY+A+ +AE+F +E+V LHG+P +V SD DPIFVS FW EL
Sbjct: 1 LKHPYSARVLAEIFTKEVVHLHGVPASVLSDEDPIFVSSFWKEL 132
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 172 bits (437), Expect = 8e-43
Identities = 97/254 (38%), Positives = 141/254 (55%), Gaps = 19/254 (7%)
Frame = +1
Query: 612 IERQVTELLEAGIIRP-SMSSFSSPVILVKKKD------------------KSWRMCVDY 652
+ ++V +LLE G+I P S S++ SPV++V KK+ SW++C+DY
Sbjct: 1 VRKEVLKLLEVGLIYPISDSAWVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDY 180
Query: 653 RALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEEDIPKTTFRTHNGH 712
R LN+AT D +P+P +D++L+ L G + +D GY+QI V +D K F G
Sbjct: 181 RKLNEATRKDHFPLPFMDQMLERLAGHAYYCFLDAYFGYNQIVVDPKDQEKMAFTCPFGV 360
Query: 713 YEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSLLEHRDHLKLVLTV 772
+ Y +PFGL NAP TFQ M IF + K + VF DD ++ SL L++VL
Sbjct: 361 FAYRRIPFGLCNAPTTFQMCMLAIFADIVEKSIEVFMDDFSVFVPSLESCLKKLEMVLQR 540
Query: 773 LLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEWPEPKNVKGVRGFLGL 832
+ + V N KC F + LGH IS G+ VD K+ I + P P NVKG+R FLG
Sbjct: 541 CVETNLVLNWEKCHFMVREGIVLGHKISTRGIEVDQTKIDVIEKLPPPSNVKGIRSFLGQ 720
Query: 833 TGYYRKFIKNYGKM 846
+YR+FIK++ K+
Sbjct: 721 ARFYRRFIKDFTKV 762
>BI425021
Length = 426
Score = 167 bits (424), Expect = 3e-41
Identities = 74/142 (52%), Positives = 99/142 (69%)
Frame = -1
Query: 1138 LQPFPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKYSHFILLKHPYTAKSIAEVF 1197
L P PVP WEDLS+DFI+GLP G+ I VVV+R SK H L +TA +A +F
Sbjct: 426 LCPLPVPQRPWEDLSMDFIVGLPPYHGHTTIFVVVNRFSKGIHLGTLPTSHTAHMVASLF 247
Query: 1198 VREIVRLHGIPNTVKSDRDPIFVSHFWSELFKLQGTKLKMSSSYHPETDGQTEVINRCLE 1257
+ +++LHG P ++ SDRDP+F+SHFW +LF+L GT L+MSS+YHP+TDGQTEV+NR +E
Sbjct: 246 LNIVIKLHGFPRSIVSDRDPLFISHFWQDLFRLSGTVLRMSSAYHPQTDGQTEVLNRVIE 67
Query: 1258 SYLRCFASDHPKTWSFWVPWAE 1279
YLR F P+ ++PW E
Sbjct: 66 QYLRAFVHGRPRNLGRFIPWVE 1
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 163 bits (413), Expect = 5e-40
Identities = 101/254 (39%), Positives = 138/254 (53%), Gaps = 19/254 (7%)
Frame = +1
Query: 612 IERQVTELLEAGIIRP-SMSSFSSPVILVKKK--------DKS----------WRMCVDY 652
+ ++V +LLEAG+I P S SS+ SPV +V KK D++ WRMC+DY
Sbjct: 1 VRKEVFKLLEAGLIYPISDSSWVSPVQVVPKKGGMTVVKNDRNELIPTRRVTRWRMCIDY 180
Query: 653 RALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHEEDIPKTTFRTHNGH 712
R LN+AT D YP+P +D++L L + +D SGY+QI V +D KT F
Sbjct: 181 RKLNEATRKDHYPLPFMDQMLKRLARQSFYRFLDGYSGYNQIAVDPQDQEKTAFTCPFSV 360
Query: 713 YEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKSLLEHRDHLKLVLTV 772
+ Y MPFGL NA TFQ M IF + K + VF DD + S +L+ VL
Sbjct: 361 FAYRRMPFGLCNASTTFQRCMMAIFDDMVEKCIEVFMDDFSFFGASFGNCLANLEKVLQR 540
Query: 773 LLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEWPEPKNVKGVRGFLGL 832
++ V N KC F + LGH IS G+ V EK+ I + P P NVKG+ FLG
Sbjct: 541 CEKSNLVLNWEKCHFMVQEGIVLGHKISKRGIEVVKEKLDVIDKLPPPVNVKGIHSFLGH 720
Query: 833 TGYYRKFIKNYGKM 846
G+YR+FIK++ K+
Sbjct: 721 VGFYRRFIKDFTKV 762
>TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%)
Length = 589
Score = 155 bits (393), Expect = 1e-37
Identities = 75/148 (50%), Positives = 103/148 (68%), Gaps = 1/148 (0%)
Frame = +1
Query: 818 PEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDN-FTWGEEAKQAFHKLKTVM 876
P K+V +R F GL +YR+F+ N+ +A PL EL KK+ FTWGE+ +QAF LK +
Sbjct: 97 PTLKSVGDIRSFHGLASFYRRFVPNFSTVASPLNELVKKNMAFTWGEKQEQAFALLKEKL 276
Query: 877 TSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMA 936
T +PVLALPDF K+FE+ECDA+G G+ AVL+Q PIAYFS+ L + L Y+KEL A
Sbjct: 277 TKAPVLALPDFSKTFELECDASGVGVRAVLLQGGHPIAYFSEKLHSATLNYPTYDKELYA 456
Query: 937 LVLCIQHWRHYLLGKAFTVYTDHKSLKH 964
L+ Q W H+L+ K F +++DH+SLK+
Sbjct: 457 LIRAPQTWEHFLVCKEFVIHSDHQSLKY 540
Score = 35.4 bits (80), Expect = 0.19
Identities = 15/37 (40%), Positives = 22/37 (58%)
Frame = +2
Query: 792 IDYLGHIISGAGVSVDPEKVQCIVEWPEPKNVKGVRG 828
I + G ++ GV +DPEK++ I EWP P V + G
Sbjct: 17 IFFSGFVVGRNGVQMDPEKIKAIQEWPPP*KVWEILG 127
>CF922488
Length = 741
Score = 138 bits (348), Expect = 2e-32
Identities = 89/245 (36%), Positives = 129/245 (52%), Gaps = 1/245 (0%)
Frame = +3
Query: 639 VKKKDKSWRMCVDYRALNKATIPDKYPIPIVDELLDELNGAVIFSKIDLKSGYHQIRVHE 698
V K+D MCVDYR LN A+ DK+P+P ++ L+D FS +D SGY+QI++
Sbjct: 3 VLKEDGKV*MCVDYRDLN*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQIKIAP 182
Query: 699 EDIPKTTFRTHNGHYEYLVMPFGLMNAPATFQAIMNDIFRPYLRKFVLVFFDDILIYSKS 758
ED+ KTTF T G + Y M FGL N AT+Q M +F + K + V+ DD+++ S++
Sbjct: 183 EDMEKTTFITLWGTFCYKAMSFGLKNVGATYQRAMVALF*DMMHKEIEVYMDDMIVKSRT 362
Query: 759 LLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDPEKVQCIVEWP 818
EH +L+ + L N +KC F L I S G+ VD KV+ I+E
Sbjct: 363 EEEHLVNLRKLFRRLRKYRLRLNPAKCMFEVKSRKLLDFIDS*RGIEVDSNKVKVILEMA 542
Query: 819 EPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFT-WGEEAKQAFHKLKTVMT 877
+P K V+GFLG Y +FI +PL L K+ F W + AF ++K +
Sbjct: 543 KPHTEKQVQGFLGRLNYIVRFIS*LIATCEPLFILLCKNQFVKWDHDC*VAFERIKQCLI 722
Query: 878 SSPVL 882
+ VL
Sbjct: 723 NPHVL 737
>AW570005
Length = 413
Score = 137 bits (344), Expect = 5e-32
Identities = 68/133 (51%), Positives = 86/133 (64%)
Frame = -2
Query: 818 PEPKNVKGVRGFLGLTGYYRKFIKNYGKMAKPLTELTKKDNFTWGEEAKQAFHKLKTVMT 877
P P+ + +RGFL LTG+YR+FIK Y MA PL+ L KD+F W EA AF LK V+T
Sbjct: 412 PPPRTARSLRGFLRLTGFYRRFIKGYAAMAAPLSHLLTKDSFVWSPEADVAFQALKNVVT 233
Query: 878 SSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQPIAYFSKALSNGNLTKSVYEKELMAL 937
++ VLALPDF K F VE DA+G +GAVL Q+ PIA+FSK + S Y EL A+
Sbjct: 232 NTLVLALPDFTKPFTVETDASGSDMGAVLSQEGHPIAFFSKEFCPKLVRSSTYVHELAAI 53
Query: 938 VLCIQHWRHYLLG 950
++ WR YLLG
Sbjct: 52 TNVVKKWRQYLLG 14
>BM731326 weakly similar to GP|21740635|em OSJNBb0043H09.2 {Oryza sativa
(japonica cultivar-group)}, partial (3%)
Length = 424
Score = 130 bits (328), Expect = 3e-30
Identities = 66/135 (48%), Positives = 93/135 (68%), Gaps = 1/135 (0%)
Frame = +1
Query: 1337 LQRAQEQMASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDK 1396
L+RAQ M +AN RR ++G+WV+LK+RPHRQ S+ R++ KL ARFYGP+ + +
Sbjct: 25 LERAQSLMVKHANNHRRPHDINVGDWVYLKIRPHRQGSMPPRLHPKLTARFYGPYLVMRQ 204
Query: 1397 IGTVAYKLKLPVESRIHPVFHVSLLKRAIGNYQVQGQLPTDLGIEEATDIY-PEAILGTR 1455
+G VA++L+LP E+RIHPVFHVS LKRA+GN+Q Q +LP DL E ++Y P IL R
Sbjct: 205 VGAVAFQLQLPSEARIHPVFHVSQLKRALGNHQAQEELPPDL--EHQAELYFPVQILKIR 378
Query: 1456 IIRQGDSEVHQSLIK 1470
+++ Q LI+
Sbjct: 379 EVQKQHEVERQVLIR 423
>BQ627806
Length = 435
Score = 75.1 bits (183), Expect(2) = 1e-29
Identities = 29/71 (40%), Positives = 49/71 (68%)
Frame = +1
Query: 925 LTKSVYEKELMALVLCIQHWRHYLLGKAFTVYTDHKSLKHFLQQKISSPDQQCWLAKLLG 984
L S Y +EL A+ + ++ WR YLLG F + TDH+SLK + Q + +P+QQ +LA+L+G
Sbjct: 220 LRASTYVRELAAITVAVKKWRQYLLGHHFVILTDHRSLKELMSQAVQTPEQQIYLARLMG 399
Query: 985 YQFEVKYKPGQ 995
+ + ++Y+ G+
Sbjct: 400 FDYTIQYRAGK 432
Score = 74.7 bits (182), Expect(2) = 1e-29
Identities = 35/68 (51%), Positives = 46/68 (67%)
Frame = +3
Query: 853 LTKKDNFTWGEEAKQAFHKLKTVMTSSPVLALPDFDKSFEVECDAAGRGIGAVLMQQRQP 912
L KD F W EEA +AF +LK + +PVL LPDF+ SF VE DA+G G+GA+L Q P
Sbjct: 3 LLVKDQFHWNEEADRAFSQLKLALCQAPVLGLPDFNSSFVVETDASGIGMGAILSQNHHP 182
Query: 913 IAYFSKAL 920
+A+F A+
Sbjct: 183 LAFFIYAI 206
>BI317507
Length = 359
Score = 128 bits (322), Expect = 2e-29
Identities = 65/116 (56%), Positives = 79/116 (68%), Gaps = 1/116 (0%)
Frame = -1
Query: 749 FDDILIYSKSLLEHRDHLKLVLTVLLSNSFVANQSKCKFGCAQIDYLGHIISGAGVSVDP 808
F +ILIYS H HL VL VL VAN+ KC F I+YLGH+IS V++D
Sbjct: 359 FYNILIYSPDWKSHIMHLTAVLDVLKKERLVANRKKCYFSQTTIEYLGHVISKDCVAMDS 180
Query: 809 EKVQCIVEWPEPKNVKGVRGFLGLTGYYRKFIKNYGKMA-KPLTELTKKDNFTWGE 863
KV+ ++EWP PKNVK V FL LTGYYRKFIK+YGK+A +PLT+LTK D F WG+
Sbjct: 179 NKVKSVIEWPVPKNVKRVCSFLRLTGYYRKFIKDYGKLAPRPLTDLTKNDGFKWGD 12
>BM084967
Length = 426
Score = 122 bits (306), Expect = 1e-27
Identities = 64/135 (47%), Positives = 81/135 (59%)
Frame = -2
Query: 1040 LLQDVQQDPQSKPGFSVLQGVLLYQGRLVLSPTSPSIPWLLQEYHGSPTGGHSGFLRTYR 1099
L Q + +P + P SV+Q ++L G + L IP LL EYH SPT H G +T
Sbjct: 410 LQQAILAEPTTYPDHSVIQDLILKNGCIWLPSGFSFIPTLLLEYHSSPTDAHIGVTKTMA 231
Query: 1100 RLAESLYWVGMQRTVRDFVRACDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGL 1159
RL+E+ W+G+++ V FV AC CQ KY A GLL P PVP WEDLS +FIIGL
Sbjct: 230 RLSENFTWIGIRKDVEQFVAACLDCQYTKYEAQKMAGLLCPLPVPCRPWEDLSFNFIIGL 51
Query: 1160 PKSKGYEAILVVVDR 1174
+ +GY AILVVV R
Sbjct: 50 SEFRGYTAILVVVGR 6
>CD487724
Length = 676
Score = 121 bits (303), Expect = 3e-27
Identities = 73/219 (33%), Positives = 112/219 (50%), Gaps = 3/219 (1%)
Frame = +3
Query: 406 LLILIDSGASHNFISPKITSALGLVITPITTRSIKLGDGHKVVTRGICRGIKAKLGSIEI 465
L+ L+D G++HNF+ + S LGL + +G+GH + IC I + +IE
Sbjct: 24 LVYLVDGGSTHNFVQQPLVSQLGLPCRSTPPLRVMVGNGHHLKCTTICEAIPISIQNIEF 203
Query: 466 TIDALVLELGGLDLVLGVSWLSTLGKVIMDWKLLTMQFVYGEQVVQLQGMRGKDSYQSNL 525
+ VL + G ++VLGV WL TLG +++D+ L+MQF Y ++VQL+G N
Sbjct: 204 LVHLYVLPIVGANIVLGVQWLKTLGPILVDYNSLSMQFFYQHRLVQLKGESEAQLGLLNH 383
Query: 526 HSF--LSDTHGREQMEWWGPQLQVVEAT-KKGHEIPQELRAVLADFHEVFTKKIQLPPVR 582
H L TH E + ++ + + +PQ ++ +L F +F LPP R
Sbjct: 384 HQLRRLHQTH--EPVTYFHIAILTENTSPTSSPPLPQPIQHLLDQFSALFQ*PQGLPPAR 557
Query: 583 SRVHQITLNPEHGPINVRPYRYPHHQKEEIERQVTELLE 621
H I L P P+N+R Y YPH+ EIE QV +L+
Sbjct: 558 ETDHHIHLLP*SEPVNMRLY*YPHY*NNEIEHQVNLMLQ 674
>TC213413 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, partial (5%)
Length = 761
Score = 120 bits (300), Expect = 6e-27
Identities = 63/151 (41%), Positives = 96/151 (62%), Gaps = 2/151 (1%)
Frame = +1
Query: 1358 DIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYKLKLPVESRIHPVFH 1417
+IG+WV +KLRPHRQ S + KL R+YGPF++++++G V Y+LKL SRIHPVFH
Sbjct: 1 NIGDWVLVKLRPHRQGSASETTYSKLTKRYYGPFEVQERLGKVVYRLKLTAHSRIHPVFH 180
Query: 1418 VSLLKRAIGNYQV--QGQLPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRS 1475
VSLLK +G+ + G LP + EEAT+ P ++ ++++ + L++W + S
Sbjct: 181 VSLLKAFVGDPETTHAGPLPV-MRTEEATNT-PLTVIDSKLVPADNGPRRMVLVQWPSAS 354
Query: 1476 IDDVTWEDNEVLAGQFPEFCLEDKAVSKEGG 1506
D +WED +VL ++ LEDK +S+E G
Sbjct: 355 RQDASWEDWQVLRERYN---LEDKVLSEERG 438
>TC213114 weakly similar to UP|Q8W150 (Q8W150) Polyprotein, partial (7%)
Length = 810
Score = 101 bits (251), Expect(2) = 3e-25
Identities = 46/104 (44%), Positives = 73/104 (69%)
Frame = +3
Query: 1344 MASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAARFYGPFQIEDKIGTVAYK 1403
M +YA++ RR ++ +G+WV+LKL+P+R S+ K+ N+KL+ RFYGP+QI+ +IG VA++
Sbjct: 3 MKAYADRSRRAVTLSVGDWVYLKLQPYRLKSLAKKRNEKLSPRFYGPYQIKKQIGLVAFE 182
Query: 1404 LKLPVESRIHPVFHVSLLKRAIGNYQVQGQLPTDLGIEEATDIY 1447
L LP +IHPVFH SLLK+A+ LP L + +++ +
Sbjct: 183 LDLPPARKIHPVFHASLLKKAVAATANPQPLPLMLSEDLSSEFF 314
Score = 34.3 bits (77), Expect(2) = 3e-25
Identities = 19/56 (33%), Positives = 31/56 (54%)
Frame = +2
Query: 1455 RIIRQGDSEVHQSLIKWKNRSIDDVTWEDNEVLAGQFPEFCLEDKAVSKEGGIERD 1510
+ + + + + LI+ ++ + TWE EV+ QFP F LEDK V+ GG+ D
Sbjct: 326 KAVHNNSNGIAEVLIQLEDLPDFEATWESVEVIKEQFPSFHLEDK-VTLLGGVLLD 490
>TC211973 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, partial (4%)
Length = 730
Score = 114 bits (284), Expect = 4e-25
Identities = 57/116 (49%), Positives = 82/116 (70%), Gaps = 11/116 (9%)
Frame = +1
Query: 1326 RDEALKQLRTHLQRAQEQMASYANKKRRDLSFDIGEWVFLKLRPHRQHSVVKRINQKLAA 1385
RD L LR +L ++ + M + NK+RRD+ + +G+ VFLK++P+R+ S+ KRIN+KL+
Sbjct: 43 RDGLLATLRENLLKS*DIM*ANTNKRRRDIEYVVGD*VFLKMQPYRRRSLAKRINEKLSP 222
Query: 1386 RFYGPFQIEDKIGTVAYKLKLPVESRIHPVFHVSLLKRA-----------IGNYQV 1430
RFY PFQ+ +K+GT+AYKL LP +IHPVFHVSLLK+A IGNY++
Sbjct: 223 RFYAPFQVFNKVGTIAYKLDLPSHIKIHPVFHVSLLKKASPNLYHLCYLRIGNYKL 390
Score = 48.5 bits (114), Expect = 2e-05
Identities = 34/103 (33%), Positives = 51/103 (49%)
Frame = +3
Query: 1434 LPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDVTWEDNEVLAGQFPE 1493
LP L + Y +++L +R ++ G+ +V LI+WKN + +WE L F
Sbjct: 351 LPPMLSEDWKLQTYSDSVLDSRELQPGNVKV---LIQWKNLPPSENSWESVAKLQEIFSI 521
Query: 1494 FCLEDKAVSKEGGIERDSNTKWGMGLGKPKIWKVYVRKKGKGA 1536
+ LEDK GGI++ + KP I KVY RK +GA
Sbjct: 522 YHLEDKVSLLGGGIDKHKH--------KPPIPKVYTRKHPRGA 626
>BI317638 weakly similar to GP|9294238|dbj| contains similarity to reverse
transcriptase~gene_id:K11J14.5 {Arabidopsis thaliana},
partial (5%)
Length = 420
Score = 100 bits (249), Expect(2) = 2e-24
Identities = 52/108 (48%), Positives = 66/108 (60%)
Frame = -2
Query: 1121 CDTCQRQKYSAMSPGGLLQPFPVPNAVWEDLSLDFIIGLPKSKGYEAILVVVDRLSKYSH 1180
C CQ KY LL P VP+ WEDLSLDFI GL + AILVVVD SK H
Sbjct: 326 CLDCQHTKYETKRIVDLLCPLLVPHRPWEDLSLDFITGLLPYHVHTAILVVVDHFSKGIH 147
Query: 1181 FILLKHPYTAKSIAEVFVREIVRLHGIPNTVKSDRDPIFVSHFWSELF 1228
+L +TA ++A +F+ + +LHG+P ++ SD D +FVSHFW ELF
Sbjct: 146 LGMLPSSHTAHTVACLFIDSVAKLHGLPRSLVSDCDLLFVSHFWQELF 3
Score = 32.3 bits (72), Expect(2) = 2e-24
Identities = 12/27 (44%), Positives = 19/27 (69%)
Frame = -1
Query: 1088 TGGHSGFLRTYRRLAESLYWVGMQRTV 1114
TGGH+G +T L++++YW GM+ V
Sbjct: 414 TGGHTGIAKTLA*LSKNIYWFGMRTDV 334
>TC212592 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, partial (4%)
Length = 664
Score = 104 bits (259), Expect = 3e-22
Identities = 64/140 (45%), Positives = 84/140 (59%), Gaps = 3/140 (2%)
Frame = +2
Query: 1365 LKLRPHRQHSVV--KRINQKLAARFYGPFQIEDKIGTVAYKLKLPVESRIHPVFHVSLLK 1422
LKLRPHRQ S + I KL+ RF+GPFQ+ +++G VAY+L+LPV++++HPVFH SLLK
Sbjct: 17 LKLRPHRQSSAKGPEPITGKLSKRFFGPFQVVERVGKVAYRLQLPVDAKLHPVFHCSLLK 196
Query: 1423 RAIGN-YQVQGQLPTDLGIEEATDIYPEAILGTRIIRQGDSEVHQSLIKWKNRSIDDVTW 1481
GN LP L + + I P IL TR + D EV L++W+ S DD TW
Sbjct: 197 PFQGNPPDTAAPLPPTL-FDHQSVIAPLVILATRTVNDDDIEV---LVQWQGLSPDDATW 364
Query: 1482 EDNEVLAGQFPEFCLEDKAV 1501
E L EF LEDK +
Sbjct: 365 EKWTELC---KEFHLEDKVL 415
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.140 0.434
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 68,990,792
Number of Sequences: 63676
Number of extensions: 1003129
Number of successful extensions: 5582
Number of sequences better than 10.0: 90
Number of HSP's better than 10.0 without gapping: 5321
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5473
length of query: 1543
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1433
effective length of database: 5,635,272
effective search space: 8075344776
effective search space used: 8075344776
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0265.11


