

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
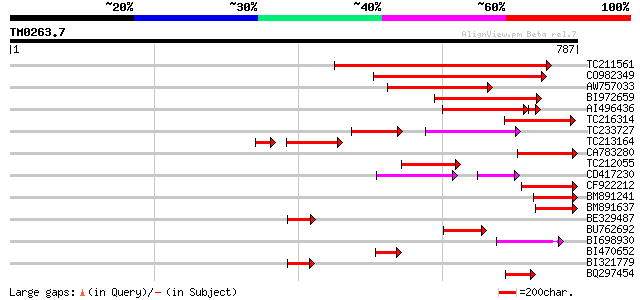
Query= TM0263.7
(787 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC211561 weakly similar to UP|O22675 (O22675) Reverse transcript... 355 3e-98
CO982349 262 4e-70
AW757033 165 6e-41
BI972659 137 1e-32
AI496436 114 9e-28
TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {A... 109 5e-24
TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR ret... 72 1e-21
TC213164 94 5e-21
CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polym... 94 2e-19
TC212055 92 1e-18
CD417230 weakly similar to GP|2244915|emb| reverse transcriptase... 72 8e-17
CF922212 73 4e-13
BM891241 70 5e-12
BM891637 67 4e-11
BE329487 weakly similar to PIR|T14619|T146 reverse transcriptase... 67 4e-11
BU762692 65 1e-10
BI698930 65 2e-10
BI470652 similar to PIR|T17102|T171 RNA-directed DNA polymerase ... 55 2e-07
BI321779 52 1e-06
BQ297454 49 9e-06
>TC211561 weakly similar to UP|O22675 (O22675) Reverse transcriptase
(Fragment) , partial (59%)
Length = 1077
Score = 355 bits (912), Expect = 3e-98
Identities = 170/303 (56%), Positives = 226/303 (74%), Gaps = 1/303 (0%)
Frame = +1
Query: 451 LDSVVVANEVVEEAKRRKKECFVFKVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIKGC 510
L S ++ANEV++EAKR K C VFKVD+EKAYDSV WNFL YMM R+ FS KWI WI+ C
Sbjct: 1 LHSALIANEVIDEAKRSNKSCLVFKVDYEKAYDSVSWNFLMYMMWRMDFSPKWIKWIEEC 180
Query: 511 IQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGYRV 570
++SAS SVLVNGSPT EF RGLRQGDPL PFLF IVAEGLNGLMR+AV++ L+ Y V
Sbjct: 181 VKSASISVLVNGSPTAEFLPQRGLRQGDPLTPFLFNIVAEGLNGLMRRAVEENLYKPYLV 360
Query: 571 GGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGISMVSR 630
G + V IS+LQ+ADDT+FFGEA+ +N+ +K ILR FE +S L++NF KS + +
Sbjct: 361 GANGVPISILQYADDTIFFGEAAKENVEAIKVILRSFELVSNLRINFAKSCFGVFGVTDQ 540
Query: 631 ETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSLSFGGRI 690
Q A L+C ++ PF YLGIP+GAN R+ W+ +I+K +RKLS K + +SFGGR+
Sbjct: 541 WKQEAANYLHCSLLAFPFIYLGIPIGANSRRCHMWDSIIRKCERKLSKWK*RHISFGGRV 720
Query: 691 CLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLW-GGEEGRGVAWVKWEEICKKKEE 749
LIKSVL+S+P++FFSFF+ P+ ++D+ KI R FLW GG + + ++W++WE++C KE
Sbjct: 721 TLIKSVLTSIPIYFFSFFRVPQSVVDKLVKI*RTFLWGGGPDQKKISWIRWEKLCLPKER 900
Query: 750 GGL 752
GG+
Sbjct: 901 GGI 909
>CO982349
Length = 795
Score = 262 bits (670), Expect = 4e-70
Identities = 126/241 (52%), Positives = 173/241 (71%), Gaps = 1/241 (0%)
Frame = +3
Query: 505 HWIKGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKL 564
HWI+GC+ SAS S+LVN SP EF+ RGLRQGDPLAP LF IVAEGL GLMR+AV +K
Sbjct: 66 HWIRGCLHSASVSILVNESPINEFSPQRGLRQGDPLAPLLFNIVAEGLTGLMREAVDRKR 245
Query: 565 FTGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAG 624
F + VG ++ +S+LQ+ADDT+FF EA+ +N+ V+K+ILR FE SGLK+NF +S+
Sbjct: 246 FNSFLVGKNKEPVSILQYADDTIFFEEATMENMKVIKTILRCFELASGLKINFAESRFGA 425
Query: 625 ISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSL 684
I + A LNC ++ +PF+YLGIP+ ANPR +P+I+K + KL+ K + +
Sbjct: 426 IWKPDHWCKEAAEFLNCSMLSMPFSYLGIPIEANPRCREI*DPIIRKCETKLARWKQRHI 605
Query: 685 SFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLW-GGEEGRGVAWVKWEEI 743
S GGR+ LI ++L++LP++FFSFF+AP ++ + IQRKFLW GG R +AWVKWE +
Sbjct: 606 SLGGRVTLINAILTALPIYFFSFFRAPSKVVQKLVNIQRKFLWGGGSXQRRIAWVKWETV 785
Query: 744 C 744
C
Sbjct: 786 C 788
>AW757033
Length = 441
Score = 165 bits (418), Expect = 6e-41
Identities = 78/146 (53%), Positives = 107/146 (72%)
Frame = -2
Query: 525 TEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGYRVGGDEVEISLLQFAD 584
T EF+ RGLRQGDPLAP LF I AEGL GLMR+AV + + + VG + +S+LQ+AD
Sbjct: 440 TREFSPHRGLRQGDPLAPLLFNIAAEGLTGLMREAVARNHYKSFAVGKYKKPVSILQYAD 261
Query: 585 DTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGISMVSRETQIFAAMLNCKVM 644
DT+FFGEA+ +N+ V+K+ILR FE SGLK+NF KS+ IS+ + A +NC ++
Sbjct: 260 DTIFFGEATMENVRVIKTILRGFELASGLKINFAKSRFGAISVPD*WRKEAAEFMNCSLL 81
Query: 645 GVPFTYLGIPVGANPRKAATWEPVIK 670
+PF+YLGIP+GANPR+ TW+P+I+
Sbjct: 80 SLPFSYLGIPIGANPRRRETWDPIIR 3
>BI972659
Length = 453
Score = 137 bits (346), Expect = 1e-32
Identities = 66/150 (44%), Positives = 96/150 (64%), Gaps = 1/150 (0%)
Frame = +1
Query: 590 GEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGISMVSRETQIFAAMLNCKVMGVPFT 649
GEAS NI+V+KS+LR FE +SGL++N+ KS+ + A +LNC+ + +PF
Sbjct: 4 GEASWDNIIVLKSMLRGFEMVSGLRINYAKSQFGIVGFQPNWAHDAAQLLNCRQLDIPFH 183
Query: 650 YLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSLSFGGRICLIKSVLSSLPLFFFSFFK 709
YLG+P+ WEP+I K + KLS K LS GGR+ LIKSVL++LP++ SFFK
Sbjct: 184 YLGMPIAVKASSRMVWEPLINKFKAKLSKWNQKYLSMGGRVTLIKSVLNALPIYLLSFFK 363
Query: 710 APKCIIDQCNKIQRKFLWGGEEGRG-VAWV 738
P+ I+D+ +QR F+WGG + ++WV
Sbjct: 364 IPQRIVDKLVSLQRTFMWGGNQHHNRISWV 453
>AI496436
Length = 414
Score = 114 bits (286), Expect(2) = 9e-28
Identities = 56/118 (47%), Positives = 79/118 (66%)
Frame = +1
Query: 602 SILRWFEAMSGLKVNFFKSKLAGISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRK 661
+ILR FE SGLK+NF KS+ I + A LN V+ +PF YLGIP+GAN R
Sbjct: 1 TILRCFELSSGLKINFAKSQFGTICKSDIWRKEAADYLNSSVLSMPFVYLGIPIGANSRH 180
Query: 662 AATWEPVIKKMQRKLSL*KHKSLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCN 719
+ WEPV++K +RKL+ K K +SFGGR+ LI SVL++LP++ SFF+ P ++D+ +
Sbjct: 181 SDVWEPVVRKCERKLASWKQKYVSFGGRVTLINSVLTALPIYLLSFFRIPDKVVDKAD 354
Score = 28.1 bits (61), Expect(2) = 9e-28
Identities = 12/18 (66%), Positives = 15/18 (82%), Gaps = 1/18 (5%)
Frame = +2
Query: 721 IQRKFLWGGE-EGRGVAW 737
IQR+FLWGG+ E R +AW
Sbjct: 359 IQRRFLWGGDLE*RKIAW 412
>TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;} , partial (18%)
Length = 759
Score = 109 bits (272), Expect = 5e-24
Identities = 50/100 (50%), Positives = 67/100 (67%), Gaps = 1/100 (1%)
Frame = +2
Query: 687 GGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEEGRG-VAWVKWEEICK 745
GGR+ LIKSVLS+LP+ SFFK P+ I+D+ +QR+FLWGG + + WVKW +IC
Sbjct: 5 GGRVTLIKSVLSALPIXXLSFFKIPQRIVDKLVTLQRQFLWGGTQHHNRIPWVKWADICN 184
Query: 746 KKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLE 785
K +GGLG+KDL +FN AL +W W L LW ++ E
Sbjct: 185 PKIDGGLGIKDLSNFNAALRGRWIWGLASNHNQLWARLAE 304
>TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR retroelement
reverse transcriptase, partial (7%)
Length = 956
Score = 72.0 bits (175), Expect(2) = 1e-21
Identities = 31/71 (43%), Positives = 50/71 (69%)
Frame = -3
Query: 475 KVDFEKAYDSVDWNFLFYMMSRIGFSHKWIHWIKGCIQSASSSVLVNGSPTEEFNMGRGL 534
K+D KA+D++D +FL ++++ G+SH + +WI+ + SA S+ VNG P F+ RG+
Sbjct: 756 KIDIVKAFDTLDRDFLISVLNKFGYSHTFCNWIRVILLSAKLSISVNGEPVGFFSCQRGV 577
Query: 535 RQGDPLAPFLF 545
RQGDPL+P L+
Sbjct: 576 RQGDPLSPSLY 544
Score = 50.1 bits (118), Expect(2) = 1e-21
Identities = 33/131 (25%), Positives = 65/131 (49%)
Frame = -2
Query: 578 SLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSKLAGISMVSRETQIFAA 637
S + ADD + F + +++N+ + + + ++ G ++ KSK+ S+ + ++ +
Sbjct: 451 SHVMHADDIMIFCKGTSRNVHNLINFFQLYKDSFGQVISKHKSKIFIGSIPGQRLRVISD 272
Query: 638 MLNCKVMGVPFTYLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSLSFGGRICLIKSVL 697
+L +PF Y G+P+ V K++ KL+ K LS GRI L+ SV+
Sbjct: 271 LLQYFHSDIPFNYFGVPIFKGKPTRRHLYCVADKIKCKLASWKGSILSIMGRIQLVNSVI 92
Query: 698 SSLPLFFFSFF 708
+ L+ FS +
Sbjct: 91 HGMLLYSFSVY 59
>TC213164
Length = 446
Score = 94.4 bits (233), Expect(2) = 5e-21
Identities = 45/78 (57%), Positives = 60/78 (76%)
Frame = +3
Query: 385 LRGGNASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKLLSMRLREVLSKVIDDRQTAF 444
L+ NA F+ LIPKV +P L Y+PISL+GC+YKIVAKLLS R ++VL +ID+RQT F
Sbjct: 189 LKRSNAFFLALIPKVHDP*SLNEYKPISLIGCIYKIVAKLLSGRHKKVLPTIIDERQTVF 368
Query: 445 IGGRFMLDSVVVANEVVE 462
+ GR L +VV+ANE++E
Sbjct: 369 MEGRHTLHNVVIANEIME 422
Score = 25.8 bits (55), Expect(2) = 5e-21
Identities = 15/28 (53%), Positives = 17/28 (60%)
Frame = +1
Query: 342 GSVGMTKARDRTGLISGSSRILGR*SKE 369
G VGM +A DR IS SSR G SK+
Sbjct: 61 GIVGMRRAPDRMD*ISSSSRNFGIFSKQ 144
>CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polymerase
homolog T24M8.8 - Arabidopsis thaliana, partial (4%)
Length = 456
Score = 94.4 bits (233), Expect = 2e-19
Identities = 43/84 (51%), Positives = 57/84 (67%), Gaps = 1/84 (1%)
Frame = +2
Query: 705 FSFFKAPKCIIDQCNKIQRKFLWGGE-EGRGVAWVKWEEICKKKEEGGLGVKDLESFNKA 763
FSFF P II + +QR+FLWGGE + R +AWV W+ +C K +GGLG+KDL +FN
Sbjct: 2 FSFFSLP*KIIAKLESLQRRFLWGGEADSRKIAWVNWKTVCLPKAKGGLGIKDLRTFNTT 181
Query: 764 LLVKWRWRLLREREWLWCQVLESK 787
LL KWRW L ++ W +VL+SK
Sbjct: 182 LLGKWRWDLFYIQQEPWAKVLQSK 253
>TC212055
Length = 776
Score = 91.7 bits (226), Expect = 1e-18
Identities = 42/82 (51%), Positives = 62/82 (75%)
Frame = +1
Query: 544 LFLIVAEGLNGLMRQAVQQKLFTGYRVGGDEVEISLLQFADDTLFFGEASTQNILVVKSI 603
LF IVAEGL GLMR+A+ + L++ VG +++ +++LQ+ADDT+F GEA+ QN++ +KS+
Sbjct: 1 LFNIVAEGLTGLMREALDKSLYSSLMVGKNKIPVNILQYADDTIFLGEATMQNVMTIKSM 180
Query: 604 LRWFEAMSGLKVNFFKSKLAGI 625
LR FE SGLK++F KS I
Sbjct: 181 LRVFELASGLKIHFAKSSFGAI 246
>CD417230 weakly similar to GP|2244915|emb| reverse transcriptase like
protein {Arabidopsis thaliana}, partial (7%)
Length = 688
Score = 72.0 bits (175), Expect(2) = 8e-17
Identities = 40/112 (35%), Positives = 62/112 (54%)
Frame = -3
Query: 510 CIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPFLFLIVAEGLNGLMRQAVQQKLFTGYR 569
C+ + + S+L NG + F+ G+RQ DP+AP+LF++ E L+ L+ + +KL +
Sbjct: 668 CMSTTTMSLLWNGEVLDHFSPNIGVRQRDPIAPYLFVLCLERLSHLINLVMNKKL*RSIQ 489
Query: 570 VGGDEVEISLLQFADDTLFFGEASTQNILVVKSILRWFEAMSGLKVNFFKSK 621
V + IS L F DD + F EA+ + V+ L F SG KVN K+K
Sbjct: 488 VNRGGLLISHLAFVDDLILFVEANMDQVDVINLTLELFSDSSGAKVNVDKTK 333
Score = 33.9 bits (76), Expect(2) = 8e-17
Identities = 19/58 (32%), Positives = 31/58 (52%)
Frame = -2
Query: 650 YLGIPVGANPRKAATWEPVIKKMQRKLSL*KHKSLSFGGRICLIKSVLSSLPLFFFSF 707
YLG+P+ + T++ +I K+ ++ S K LS GR+ L K V+ +LP F
Sbjct: 243 YLGVPIFHKNPTSHTFDFIIDKVNKRFSRWKSNLLSMVGRLTLTK*VV*ALPSHIMQF 70
>CF922212
Length = 445
Score = 73.2 bits (178), Expect = 4e-13
Identities = 35/78 (44%), Positives = 50/78 (63%), Gaps = 1/78 (1%)
Frame = -2
Query: 711 PKCIIDQCNKIQRKFLWGGEEG-RGVAWVKWEEICKKKEEGGLGVKDLESFNKALLVKWR 769
P ++D+ ++Q+ FLWGG G + +AWV W+ +C KE+ GLG +DL FN ALL K R
Sbjct: 438 PNKVLDKLVRLQQWFLWGGGLGQKKIAWVNWDSVCLPKEKEGLGNRDLRKFNYALLGKRR 259
Query: 770 WRLLREREWLWCQVLESK 787
W L + L +VL+SK
Sbjct: 258 WNLFHHQGELGARVLDSK 205
>BM891241
Length = 407
Score = 69.7 bits (169), Expect = 5e-12
Identities = 27/60 (45%), Positives = 40/60 (66%)
Frame = -2
Query: 728 GGEEGRGVAWVKWEEICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
G + + + WVKWE +C K +GGLG+KD+ FN ALL KW W L +++ LW +++ SK
Sbjct: 403 GDHDHKKIPWVKWEVVCLPKTKGGLGIKDISKFNIALLGKWIWALASDQQQLWARIINSK 224
>BM891637
Length = 427
Score = 66.6 bits (161), Expect = 4e-11
Identities = 24/57 (42%), Positives = 38/57 (66%)
Frame = -1
Query: 731 EGRGVAWVKWEEICKKKEEGGLGVKDLESFNKALLVKWRWRLLREREWLWCQVLESK 787
EG+ +AW+ W + C + GGLG+KD++ N ALL+KW+W + + LW ++L SK
Sbjct: 424 EGKKIAWISWRQCCASGDVGGLGIKDIKILNNALLIKWKWLMFHQPHQLWNRILISK 254
>BE329487 weakly similar to PIR|T14619|T146 reverse transcriptase - beet
retrotransposon (fragment), partial (15%)
Length = 376
Score = 66.6 bits (161), Expect = 4e-11
Identities = 29/39 (74%), Positives = 36/39 (91%)
Frame = -2
Query: 386 RGGNASFITLIPKVENPQGLGNYRPISLVGCVYKIVAKL 424
+GGNASFI LIPKV++PQ L ++RPISL+GCVYKIVAK+
Sbjct: 228 KGGNASFIALIPKVKHPQALNDFRPISLIGCVYKIVAKI 112
>BU762692
Length = 423
Score = 65.1 bits (157), Expect = 1e-10
Identities = 30/59 (50%), Positives = 38/59 (63%)
Frame = -3
Query: 603 ILRWFEAMSGLKVNFFKSKLAGISMVSRETQIFAAMLNCKVMGVPFTYLGIPVGANPRK 661
ILR FE +SGLK+NF KS M R A+ LNC ++ +PF Y GIP+GANPR+
Sbjct: 181 ILRTFELVSGLKINFAKSCFGAFGMTDRWKTEAASCLNCSLLAIPFVYQGIPIGANPRR 5
>BI698930
Length = 384
Score = 64.7 bits (156), Expect = 2e-10
Identities = 41/95 (43%), Positives = 56/95 (58%), Gaps = 2/95 (2%)
Frame = +3
Query: 676 LSL*KHKSLSFGGRICLIKSVLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGGEEGRGV 735
L+L KH+ LSF GR CLI S+LSSLPLF FSFFK P + + I R F+ E +
Sbjct: 81 LALWKHEVLSFAGRTCLINSMLSSLPLFSFSFFKVPLNVGMRFISI*RNFVGEWRES*MI 260
Query: 736 AWVKWEEICKKK--EEGGLGVKDLESFNKALLVKW 768
WV W+++ K+ E GG + F+ L+VK+
Sbjct: 261 PWVNWDKV*VKRVCEFGGKQI-----FSCKLVVKF 350
>BI470652 similar to PIR|T17102|T171 RNA-directed DNA polymerase (EC
2.7.7.49) (clone AmLi2) - garden snapdragon (fragment),
partial (22%)
Length = 111
Score = 54.7 bits (130), Expect = 2e-07
Identities = 25/36 (69%), Positives = 27/36 (74%)
Frame = +1
Query: 508 KGCIQSASSSVLVNGSPTEEFNMGRGLRQGDPLAPF 543
K C+ SAS S+LVNGSPTEEF RGLRQG P PF
Sbjct: 1 KECLSSASISILVNGSPTEEFKPXRGLRQGGPFGPF 108
>BI321779
Length = 421
Score = 52.0 bits (123), Expect = 1e-06
Identities = 25/38 (65%), Positives = 28/38 (72%)
Frame = +2
Query: 386 RGGNASFITLIPKVENPQGLGNYRPISLVGCVYKIVAK 423
RG N SFI LI K +NP +G Y PISLVGC+YKIV K
Sbjct: 257 RGTNRSFIALIAKCDNPLDIGQYCPISLVGCLYKIV*K 370
>BQ297454
Length = 135
Score = 48.9 bits (115), Expect = 9e-06
Identities = 22/42 (52%), Positives = 32/42 (75%), Gaps = 1/42 (2%)
Frame = -1
Query: 689 RICLIKS-VLSSLPLFFFSFFKAPKCIIDQCNKIQRKFLWGG 729
R+ LI S L+S+ ++FFSFF+ PK ++D+ +IQRKFLW G
Sbjct: 135 RVTLIPSWXLTSISIYFFSFFRVPKKVVDKLARIQRKFLWEG 10
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.341 0.150 0.502
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 36,089,649
Number of Sequences: 63676
Number of extensions: 544749
Number of successful extensions: 4562
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 4413
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4549
length of query: 787
length of database: 12,639,632
effective HSP length: 105
effective length of query: 682
effective length of database: 5,953,652
effective search space: 4060390664
effective search space used: 4060390664
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0263.7


