

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
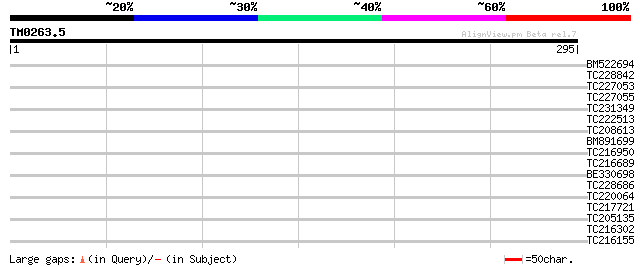
Query= TM0263.5
(295 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BM522694 31 0.58
TC228842 31 0.58
TC227053 similar to GB|AAP21363.1|30102890|BT006555 At4g38290 {A... 31 0.76
TC227055 similar to GB|AAP21363.1|30102890|BT006555 At4g38290 {A... 29 2.9
TC231349 similar to GB|BAA20386.1|2189959|AB001343 DDX10-NUP98 f... 28 3.8
TC222513 28 3.8
TC208613 similar to GB|AAP31934.1|30387537|BT006590 At2g35860 {A... 28 3.8
BM891699 28 4.9
TC216950 28 4.9
TC216689 similar to UP|Q8L8A8 (Q8L8A8) Transcription activator, ... 28 4.9
BE330698 28 6.4
TC228686 weakly similar to PIR|H64231|H64231 protein L homolog -... 28 6.4
TC220064 UP|Q8CEM2 (Q8CEM2) Mus musculus adult male testis cDNA,... 27 8.4
TC217721 similar to GB|AAR24767.1|38604048|BT010989 At5g21280 {A... 27 8.4
TC205135 similar to UP|C862_ARATH (O23066) Cytochrome P450 86A2 ... 27 8.4
TC216302 similar to UP|Q708Y0 (Q708Y0) F-box protein (EIN3-bindi... 27 8.4
TC216155 weakly similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription... 27 8.4
>BM522694
Length = 434
Score = 31.2 bits (69), Expect = 0.58
Identities = 21/69 (30%), Positives = 34/69 (48%), Gaps = 7/69 (10%)
Frame = +3
Query: 193 HESDACGSNSIGSI-------PRPMGREAAKKKNKKKSRDAALEVVNNEWSEYKQLREQE 245
HE A S+S S+ P+G++ ++ KKKS +AL + W Y+ E+E
Sbjct: 24 HEEAASSSSSSSSVGPNDAAKKNPLGKQQVSEEKKKKSHHSALP---SNWDRYED-EEEE 191
Query: 246 LERLEKISS 254
L+ I+S
Sbjct: 192 LDSGSGIAS 218
>TC228842
Length = 1023
Score = 31.2 bits (69), Expect = 0.58
Identities = 23/73 (31%), Positives = 38/73 (51%)
Frame = +2
Query: 217 KKNKKKSRDAALEVVNNEWSEYKQLREQELERLEKISSVQQEANRLEKMKMFLQLSFEEH 276
K +K A+ VV+ E ++LRE EKI++++ E N L K L++ +E+
Sbjct: 410 KMDKAVILSDAVRVVSQLREEAQKLRESTENLQEKINALKDEKNELRDEKQRLKVE-KEN 586
Query: 277 LDDRRKELLKQLS 289
L+ + K L Q S
Sbjct: 587 LEQKVKALSSQPS 625
>TC227053 similar to GB|AAP21363.1|30102890|BT006555 At4g38290 {Arabidopsis
thaliana;} , partial (42%)
Length = 1356
Score = 30.8 bits (68), Expect = 0.76
Identities = 32/120 (26%), Positives = 52/120 (42%), Gaps = 24/120 (20%)
Frame = +1
Query: 197 ACGSNSIG--SIPRPMGREAAKKKNKKKSRDAALE----------VVNNEWSE-YKQL-- 241
A NS G S+P +G+ A K + + A + N W E Y+QL
Sbjct: 658 ASSMNSSGKPSVPISLGKSAKKLAPVESNYVTASSGPTTIGNPKGLKNLHWDERYQQLQM 837
Query: 242 ---------REQELERLEKISSVQQEANRLEKMKMFLQLSFEEHLDDRRKELLKQLSQEL 292
RE+ ++ L +SSV+ + +E K +QLS EE + +R +L L + +
Sbjct: 838 FLRKLDQSDREEYIQMLHSLSSVELSKHAVELEKRSIQLSLEEAKELQRVAVLNVLGKSV 1017
>TC227055 similar to GB|AAP21363.1|30102890|BT006555 At4g38290 {Arabidopsis
thaliana;} , partial (42%)
Length = 613
Score = 28.9 bits (63), Expect = 2.9
Identities = 21/73 (28%), Positives = 36/73 (48%), Gaps = 12/73 (16%)
Frame = +1
Query: 232 NNEWSE-YKQLR-----------EQELERLEKISSVQQEANRLEKMKMFLQLSFEEHLDD 279
N W E Y+QL+ E+ ++ L +SSV+ + +E K +QLS EE +
Sbjct: 139 NLHWEERYQQLQMFLRKLDQSDQEEYIQMLRSLSSVELSKHAVELEKRSIQLSLEEAKEL 318
Query: 280 RRKELLKQLSQEL 292
+R +L L + +
Sbjct: 319 QRVAVLNVLGKSV 357
>TC231349 similar to GB|BAA20386.1|2189959|AB001343 DDX10-NUP98 fusion
protein type 2 {Homo sapiens;} , partial (14%)
Length = 484
Score = 28.5 bits (62), Expect = 3.8
Identities = 11/37 (29%), Positives = 21/37 (56%), Gaps = 4/37 (10%)
Frame = -1
Query: 143 LAKAQDIYTNGK----NSHPFTLMKEWLALCNEPHYC 175
++K Q IY + + ++ PF + + +C EPH+C
Sbjct: 226 ISKCQSIYVDPRIYTYSAIPFAIFLSYRLICYEPHFC 116
>TC222513
Length = 455
Score = 28.5 bits (62), Expect = 3.8
Identities = 18/81 (22%), Positives = 33/81 (40%), Gaps = 7/81 (8%)
Frame = +3
Query: 23 TNMTLGDMTAVNEVTPNEEDSTPRSRKTQSPAWNTEQNLVLISG-------WIKYGTSSV 75
T M L N T + + + P W ++ LVLI G + + T+ +
Sbjct: 66 TWMALEQEVGQNRNTNGVDGGDEGGKAPRLPRWTRQEILVLIQGKRDAENKFRRGRTAGL 245
Query: 76 VGRNQKGETYWSQIAEYCHEH 96
+ + E W+ ++ YC +H
Sbjct: 246 AFGSGQVEPKWASVSSYCRKH 308
>TC208613 similar to GB|AAP31934.1|30387537|BT006590 At2g35860 {Arabidopsis
thaliana;} , partial (62%)
Length = 1101
Score = 28.5 bits (62), Expect = 3.8
Identities = 13/37 (35%), Positives = 18/37 (48%)
Frame = -2
Query: 40 EEDSTPRSRKTQSPAWNTEQNLVLISGWIKYGTSSVV 76
+ ST R R+ +S W T V GW+ S+VV
Sbjct: 131 KSSSTERGRRRRSMPWMTPSGRVTTFGWVTLAWSTVV 21
>BM891699
Length = 421
Score = 28.1 bits (61), Expect = 4.9
Identities = 15/47 (31%), Positives = 21/47 (43%)
Frame = +2
Query: 186 SSGSKRSHESDACGSNSIGSIPRPMGREAAKKKNKKKSRDAALEVVN 232
S+G+KRS E+ C S+G GR A + S + L N
Sbjct: 29 SNGAKRSAEAVGCKKESVGERSALEGRTRASSAGRSGSENVGLSNAN 169
>TC216950
Length = 873
Score = 28.1 bits (61), Expect = 4.9
Identities = 16/57 (28%), Positives = 30/57 (52%), Gaps = 2/57 (3%)
Frame = +2
Query: 213 EAAKKKNKKKSRDAALEVVNNEWSEYKQLREQELERLEKI--SSVQQEANRLEKMKM 267
E ++K ++ R ALE E YK+L E + ++ E + ++E RL ++K+
Sbjct: 464 EENRRKVEEAQRREALEQQRREEERYKELEEMQRQKEEAMRKKKHEEEQERLNQIKL 634
>TC216689 similar to UP|Q8L8A8 (Q8L8A8) Transcription activator, partial
(11%)
Length = 779
Score = 28.1 bits (61), Expect = 4.9
Identities = 16/51 (31%), Positives = 25/51 (48%)
Frame = +1
Query: 152 NGKNSHPFTLMKEWLALCNEPHYCSQVGGNLGSGSSGSKRSHESDACGSNS 202
+G ++ T W+ + E S +GG LG + S S+ SD CG N+
Sbjct: 181 SGASNEANTRQANWIPITWE----SSMGGPLGEVLNLSNNSNASDQCGKNN 321
>BE330698
Length = 186
Score = 27.7 bits (60), Expect = 6.4
Identities = 20/58 (34%), Positives = 31/58 (52%), Gaps = 5/58 (8%)
Frame = +2
Query: 212 REAAKKKNKKKSRDAALEVVNNEWSE-----YKQLREQELERLEKISSVQQEANRLEK 264
R KKKNKKK A + ++++ W E K+ +E+E E SV++E R E+
Sbjct: 17 RRICKKKNKKK---AEVFIISSIWLESNIFFLKKEKEKEN**SENYESVKREGERREE 181
>TC228686 weakly similar to PIR|H64231|H64231 protein L homolog - Mycoplasma
genitalium {Mycoplasma genitalium;} , partial (5%)
Length = 1408
Score = 27.7 bits (60), Expect = 6.4
Identities = 23/73 (31%), Positives = 30/73 (40%)
Frame = +3
Query: 186 SSGSKRSHESDACGSNSIGSIPRPMGREAAKKKNKKKSRDAALEVVNNEWSEYKQLREQE 245
SS SKR SD G R G + KK KSR + E E + EQE
Sbjct: 939 SSHSKRRERSDDEEGGGTGEKKRKKGGKRRKKDKHSKSR------YDTEEPENDMMDEQE 1100
Query: 246 LERLEKISSVQQE 258
+E E + ++E
Sbjct: 1101 MEDEEADINYREE 1139
>TC220064 UP|Q8CEM2 (Q8CEM2) Mus musculus adult male testis cDNA, RIKEN
full-length enriched library, clone:1700123E07
product:protamine 1, full insert sequence, partial (22%)
Length = 461
Score = 27.3 bits (59), Expect = 8.4
Identities = 19/51 (37%), Positives = 23/51 (44%), Gaps = 6/51 (11%)
Frame = +1
Query: 164 EWLALCNE---PHYCSQVGGNLG---SGSSGSKRSHESDACGSNSIGSIPR 208
EW E PH CS+ GNLG S S ++R H + C S PR
Sbjct: 28 EWTLRYRERQVPHGCSKQPGNLG*CSSLPSSAQRQHGRENCRHPSRSRRPR 180
>TC217721 similar to GB|AAR24767.1|38604048|BT010989 At5g21280 {Arabidopsis
thaliana;} , partial (15%)
Length = 1636
Score = 27.3 bits (59), Expect = 8.4
Identities = 32/120 (26%), Positives = 45/120 (36%), Gaps = 23/120 (19%)
Frame = +2
Query: 4 SPSQTPPFNVGVTDFPEFSTNMTLGDMTAVNEVT-------PNEEDSTPRSRKTQ-SPAW 55
SP Q P +VG++ FS+ + + A +VT +E + RS Q W
Sbjct: 1025 SPPQELP-SVGISQGEGFSSLVDTIESNAATQVTGGLHTAMDDEGMAEMRSLGEQYQMEW 1201
Query: 56 NTEQNLVLISGWIKYGTSSVVGRNQKGETY---------------WSQIAEYCHEHCSFD 100
N NLV + W KY ++ R + T W E C E CS D
Sbjct: 1202 NDTMNLVKSACWFKY-LKNMEHRAHEANTVDVAYNNFDQLLEFPAWLNANESCLEQCSVD 1378
>TC205135 similar to UP|C862_ARATH (O23066) Cytochrome P450 86A2 , partial
(32%)
Length = 1096
Score = 27.3 bits (59), Expect = 8.4
Identities = 9/19 (47%), Positives = 11/19 (57%)
Frame = +1
Query: 84 TYWSQIAEYCHEHCSFDPP 102
T WS ++C HC F PP
Sbjct: 118 TVWSTSRQHCLRHCGFIPP 174
>TC216302 similar to UP|Q708Y0 (Q708Y0) F-box protein (EIN3-binding F-box
protein 2), partial (15%)
Length = 1122
Score = 27.3 bits (59), Expect = 8.4
Identities = 14/43 (32%), Positives = 22/43 (50%), Gaps = 3/43 (6%)
Frame = +2
Query: 62 VLISGWIKYGTS---SVVGRNQKGETYWSQIAEYCHEHCSFDP 101
VL+ W++ +S S+V R + W Q+ + CH H F P
Sbjct: 872 VLLLAWLQLSSSVL*SLVFR----DWLWQQVIQVCHNHDLFSP 988
>TC216155 weakly similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor
ATB2, partial (66%)
Length = 1264
Score = 27.3 bits (59), Expect = 8.4
Identities = 27/116 (23%), Positives = 48/116 (41%)
Frame = +1
Query: 174 YCSQVGGNLGSGSSGSKRSHESDACGSNSIGSIPRPMGREAAKKKNKKKSRDAALEVVNN 233
YC G G+ SSGS S + G I R R + +++ ++SR + +
Sbjct: 577 YCMASPGGSGTYSSGSSSLQNSGSEGDRDIME-QRKRKRMLSNRESARRSRIRKQQHLEG 753
Query: 234 EWSEYKQLREQELERLEKISSVQQEANRLEKMKMFLQLSFEEHLDDRRKELLKQLS 289
++ QL+++ + IS Q +E L+ E L +R L + +S
Sbjct: 754 LSAQLDQLKKENAQINTNISITTQMYLNVEAENAILRAQMGE-LSNRLNSLNEMIS 918
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.312 0.128 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,444,327
Number of Sequences: 63676
Number of extensions: 229587
Number of successful extensions: 1282
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 1109
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1211
length of query: 295
length of database: 12,639,632
effective HSP length: 97
effective length of query: 198
effective length of database: 6,463,060
effective search space: 1279685880
effective search space used: 1279685880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0263.5


