

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0260.10
(751 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
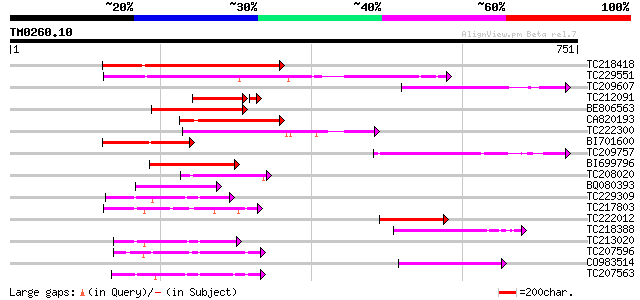
Score E
Sequences producing significant alignments: (bits) Value
TC218418 homologue to UP|Q6X4V5 (Q6X4V5) ABC transporter, partia... 202 4e-52
TC229551 weakly similar to UP|Q6X4V5 (Q6X4V5) ABC transporter, p... 196 3e-50
TC209607 136 3e-32
TC212091 similar to UP|Q9LFG8 (Q9LFG8) ABC transporter-like prot... 126 1e-30
BE806563 similar to GP|11994752|db ABC transporter-like protein ... 126 4e-29
CA820193 similar to PIR|T04229|T04 ABC-type transport protein F1... 125 9e-29
TC222300 similar to GB|AAP37786.1|30725528|BT008427 At1g51560 {A... 115 9e-26
BI701600 107 2e-23
TC209757 82 7e-16
BI699796 similar to PIR|G85167|G85 ABC transporter like protein ... 79 7e-15
TC208020 similar to UP|Q76CU2 (Q76CU2) PDR-type ABC transporter ... 78 2e-14
BQ080393 75 8e-14
TC229309 homologue to UP|Q9LJX0 (Q9LJX0) P-glycoprotein; multi-d... 74 3e-13
TC217803 similar to UP|Q9AT00 (Q9AT00) At1g65410/T8F5_19, partia... 74 3e-13
TC222012 similar to UP|Q9FLX5 (Q9FLX5) ABC transporter-like prot... 73 5e-13
TC218388 similar to UP|Q9LMU4 (Q9LMU4) F2H15.7 protein, partial ... 70 4e-12
TC213020 weakly similar to UP|Q8GU71 (Q8GU71) MDR-like ABC trans... 69 6e-12
TC207596 similar to UP|Q7GBY1 (Q7GBY1) Tonoplast ABC transporter... 69 1e-11
CO983514 69 1e-11
TC207563 similar to UP|Q9ZRG2 (Q9ZRG2) P-glycoprotein, partial (... 66 6e-11
>TC218418 homologue to UP|Q6X4V5 (Q6X4V5) ABC transporter, partial (49%)
Length = 1220
Score = 202 bits (514), Expect = 4e-52
Identities = 113/242 (46%), Positives = 159/242 (65%), Gaps = 1/242 (0%)
Frame = +1
Query: 124 TKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGR-LKGTLALNGEALESRL 182
T+ +L ++G A G A++G SGSGKSTL+DAL++R+A L GT+ LNG +++L
Sbjct: 289 TQNVLEGLTGYAEPGTFTALMGPSGSGKSTLLDALSSRLAANAFLSGTILLNGR--KAKL 462
Query: 183 LKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKT 242
+AYV QDD L LTV ET++++A RLP + + K+A V++ I +GL++ A T
Sbjct: 463 SFGTAAYVTQDDNLIGTLTVRETISYSARLRLPDNMPWADKRALVESTIVAMGLQDCADT 642
Query: 243 VIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGS 302
VIG+ RG+SGGE+RRVSI ++I+ P LLFLDEPTSGLDS SAF V + L+ +A+ G
Sbjct: 643 VIGNWHLRGISGGEKRRVSIALEILMRPRLLFLDEPTSGLDSASAFFVTQTLRALARDGR 822
Query: 303 IVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFALD 362
VI SIHQPS + L D++ LS G+TVY G ++ FFA+ G P P N ++ L
Sbjct: 823 TVIASIHQPSSEVFELFDQLYLLSSGKTVYFGQASEAYEFFAQAGFPCPALRNPSDHFLR 1002
Query: 363 LI 364
I
Sbjct: 1003CI 1008
>TC229551 weakly similar to UP|Q6X4V5 (Q6X4V5) ABC transporter, partial (34%)
Length = 1436
Score = 196 bits (498), Expect = 3e-50
Identities = 149/477 (31%), Positives = 234/477 (48%), Gaps = 16/477 (3%)
Frame = +3
Query: 125 KTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKG-RLKGTLALNGEALESRLL 183
K +L+ ++G A+ G ++A++G SGSGKSTL+DALA R+ + G + +NG E L
Sbjct: 69 KLILHGLTGYAQPGRLLAIIGPSGSGKSTLLDALAGRLTSNIKQTGKILINGHKQE--LA 242
Query: 184 KVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTV 243
S YV QDD + LT ETL ++A + P T+S +KK R + ++GL++A T
Sbjct: 243 YGTSGYVTQDDAMLSCLTAGETLYYSAMLQFPNTMSVEEKKERADMTLREMGLQDAINTR 422
Query: 244 IGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSI 303
+G +G+SGG+RRR+SI I+I+ P LLFLDEPTSGLDS +++ V+ + + Q I
Sbjct: 423 VGGWNCKGLSGGQRRRLSICIEILTHPKLLFLDEPTSGLDSAASYYVMSGIANLIQRDGI 602
Query: 304 ---VIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGHPLPDSDNRTEFA 360
++ S+HQPS + L + LS G+TVY G + FFA G P P N ++
Sbjct: 603 QRTIVASVHQPSSEVFQLFHDLFLLSSGETVYFGPASDANQFFASNGFPCPPLYNPSDHY 782
Query: 361 LDLI-RDL-----EG--SPGGTKSLVEFNKSWQSMTKVHSHSVSSQPERPNGMSLKEAIS 412
L +I +D EG + TK LV KS + V S S+ + K+ I
Sbjct: 783 LRIINKDFNQDADEGITTEEATKILVNSYKSSEFSNHVQSEIAKSETDF-GACGKKKKIH 959
Query: 413 ASISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTLSKRSFTDSRRKPELFGIRLGAV 472
A+ F + + L +R+ R RL
Sbjct: 960 AA----------------------------FITQCLILIRRASLQIYRDTNNHRARLVVF 1055
Query: 473 MVTGFILATMFWNLDNSP-KGVQERLGFFAFAMS-TTFYTTADALPVFLQERYIFMRETA 530
+ + ++F++ + + +R F +S TF T + ++E +F RE
Sbjct: 1056IFISLSVGSIFYHSGGPDLRSIMDRGSLLCFFLSVVTFMTLVGGISPLIEEMNVFKRERL 1235
Query: 531 YNAYRRSSYLISHALVSLPPLAFLSLAF--AVITFWAVGLDGGASGFLFYFLIIFAS 585
Y +++LIS+ + S P FL L AV+T+ + GL G F+F ++FA+
Sbjct: 1236NGHYGITAFLISN-IFSAVPYNFLMLIIPGAVVTYLS-GLHKGVDNFVFLISVLFAT 1400
>TC209607
Length = 828
Score = 136 bits (343), Expect = 3e-32
Identities = 75/224 (33%), Positives = 119/224 (52%), Gaps = 1/224 (0%)
Frame = +1
Query: 520 QERYIFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYF 579
QER I M+ET+ +YR SSY I++ V LP L L++ F++ +W VGL+ FL +
Sbjct: 1 QEREILMKETSCGSYRVSSYAIANGXVYLPFLLILAILFSMPLYWLVGLNRNFLAFLHFL 180
Query: 580 LIIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHY 639
L+I+ + NS V S +VP+ ++G +++ ++ F L SG+FI + IP+YWI+ HY
Sbjct: 181 LLIWLILYTANSVVVCFSALVPNFIVGNSVIAGVIGSFFLFSGYFISKQEIPNYWIFMHY 360
Query: 640 LSLVKYPYEAVLQNEFDDPVKCFVRGVQIFDNSPLRSVPYELKLKLLGSMSGTLGTNITA 699
+SL KYP+E L NEF + KC L M G
Sbjct: 361 ISLFKYPFEGFLINEFSNSGKC------------------------LEYMFG-------- 444
Query: 700 TTCLTTGADILQQNGV-TELSKWNCLWITVAWGFFFRFLFYLAL 742
C+ +G D+L++ G E ++W + +TV + +RF+ Y+ L
Sbjct: 445 -ACIKSGEDVLKEEGYGGESNRWKNVGVTVCFILVYRFISYVIL 573
>TC212091 similar to UP|Q9LFG8 (Q9LFG8) ABC transporter-like protein, partial
(12%)
Length = 477
Score = 126 bits (316), Expect(2) = 1e-30
Identities = 64/73 (87%), Positives = 67/73 (91%)
Frame = +3
Query: 243 VIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGS 302
VIGD GHR VSGGERRRVSIG DIIH+PI+LFLDEPTS LDSTSAFMVVKVLQRIAQSG+
Sbjct: 9 VIGDNGHRSVSGGERRRVSIGTDIIHNPIVLFLDEPTSNLDSTSAFMVVKVLQRIAQSGN 188
Query: 303 IVIMSIHQPSYRI 315
I IMSIH PSYRI
Sbjct: 189 IFIMSIH*PSYRI 227
Score = 26.2 bits (56), Expect(2) = 1e-30
Identities = 11/16 (68%), Positives = 13/16 (80%)
Frame = +1
Query: 318 LLDRMIFLSRGQTVYS 333
LLD +IFLS G TV+S
Sbjct: 229 LLDHLIFLSHGNTVFS 276
>BE806563 similar to GP|11994752|db ABC transporter-like protein {Arabidopsis
thaliana}, partial (20%)
Length = 428
Score = 126 bits (316), Expect = 4e-29
Identities = 65/129 (50%), Positives = 94/129 (72%), Gaps = 2/129 (1%)
Frame = +1
Query: 189 YVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGD-- 246
+V QDD+ +P LTV ETLT+AA RLP++LS+ +KK + +I +LGL + +G
Sbjct: 28 FVPQDDVHYPHLTVLETLTYAALLRLPKSLSREEKKEHAEMVIAELGLTRCRNSPVGGCM 207
Query: 247 EGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIM 306
RG+SGGER+RVSIG +++ +P LLF+DEPTSGLDST+A ++V VL +A++G V+
Sbjct: 208 ALFRGISGGERKRVSIGQEMLVNPSLLFVDEPTSGLDSTTAQLIVSVLHGLARAGRTVVA 387
Query: 307 SIHQPSYRI 315
+IHQPS R+
Sbjct: 388 TIHQPSSRL 414
>CA820193 similar to PIR|T04229|T04 ABC-type transport protein F14M19.30 -
Arabidopsis thaliana, partial (23%)
Length = 423
Score = 125 bits (313), Expect = 9e-29
Identities = 65/141 (46%), Positives = 97/141 (68%), Gaps = 1/141 (0%)
Frame = +1
Query: 225 ARVQALIDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDS 284
A V +L+ +L L + + T + RG+SGGERRRVSIG+ ++HDP +L LDEPTSGLDS
Sbjct: 7 AIVSSLLSELRLTHLSNTRLA----RGLSGGERRRVSIGLCLLHDPAVLLLDEPTSGLDS 174
Query: 285 TSAFMVVKVLQRIAQS-GSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFF 343
TSAF V+++L++ S +I+SIHQPS++IL +DR++ LS+GQ V+ GS L +F
Sbjct: 175 TSAFKVMRILKQTCVSRNRTIILSIHQPSFKILACIDRILLLSKGQVVHHGSVATLQAFL 354
Query: 344 AEFGHPLPDSDNRTEFALDLI 364
G +P N E+A++++
Sbjct: 355 HSNGFTVPHQLNALEYAMEIL 417
>TC222300 similar to GB|AAP37786.1|30725528|BT008427 At1g51560 {Arabidopsis
thaliana;} , partial (37%)
Length = 801
Score = 115 bits (287), Expect = 9e-26
Identities = 84/279 (30%), Positives = 127/279 (45%), Gaps = 18/279 (6%)
Frame = +1
Query: 229 ALIDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAF 288
A I ++GL++ +IG RG+SGGE++R+SI + I+ P LLFLDEPTSGLDS SAF
Sbjct: 13 ATIIEMGLQDCDDRLIGKWHLRGISGGEKKRLSIAL*ILTRPRLLFLDEPTSGLDSASAF 192
Query: 289 MVVKVLQRIAQSGSIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSGSPTQLPSFFAEFGH 348
VV+ L+ +A+ VI SIHQP + L D + LS G+TVY G FF E G
Sbjct: 193 FVVQTLRNVARDERTVIYSIHQPDSEVFALYDDLFLLSGGETVYFGEAKSAIEFFDEAGF 372
Query: 349 PLPDSDNRTEFALDLIR--------DLEGS------PGGTKSLVEFNKSWQSMTKVHSHS 394
P P N + L I ++GS P + + T V +
Sbjct: 373 PCPRKRNPYDHFLRCINYDFDIVTATIKGSQRIHDVPNSADPFMNLATAEIKATLVEKYK 552
Query: 395 VSSQPERPNG----MSLKEAISASISRGKLVSGATNTATTTPSSMVPTFANPFWIEMVTL 450
S+ R +S E + G S +W +++TL
Sbjct: 553 RSTYARRAKNRIQELSTDEGLHPPTQHGSQAS--------------------WWKQLLTL 672
Query: 451 SKRSFTDSRRKPELFGIRLGAVMVTGFILATMFWNLDNS 489
+KRSF + R + +R+ ++ + T ++++ S
Sbjct: 673 TKRSFVNMCRDVGYYWLRIIVYIIVSICV*TDYFDVGYS 789
>BI701600
Length = 426
Score = 107 bits (267), Expect = 2e-23
Identities = 59/123 (47%), Positives = 80/123 (64%)
Frame = +3
Query: 123 RTKTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALESRL 182
+ +T+L ++G A+ GEI+AVLG SGSGKSTL+ ALA R+ L GT+ N L +
Sbjct: 60 KERTILKGVTGIAQPGEILAVLGPSGSGKSTLLHALAGRLHGPGLTGTILANSSKLTKPV 239
Query: 183 LKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKT 242
L+ + +V QDD+L+P LTV ETL F A RLPR L +S+K A +A I +LGL T
Sbjct: 240 LRR-TGFVTQDDILYPHLTVRETLVFCAMLRLPRALLRSEKLAAAEAAIAELGLGKCENT 416
Query: 243 VIG 245
+IG
Sbjct: 417 IIG 425
>TC209757
Length = 1116
Score = 82.4 bits (202), Expect = 7e-16
Identities = 62/262 (23%), Positives = 109/262 (40%), Gaps = 1/262 (0%)
Frame = +3
Query: 482 MFWNLDNSPKGVQERLGFFAF-AMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSYL 540
M+W+ D + +Q+RLG F ++ + + +++ F QER IFM+E A Y SSY
Sbjct: 6 MWWHSDY--RNIQDRLGLLFFISIFWGVFPSFNSVFAFPQERAIFMKERASGMYTLSSYF 179
Query: 541 ISHALVSLPPLAFLSLAFAVITFWAVGLDGGASGFLFYFLIIFASFWAGNSFVTFLSGVV 600
++ + LP L F ++T+W GL FL L++ L +
Sbjct: 180 MARIVGDLPMELILPTIFLIVTYWMGGLKPDLWAFLLTLLVVLGYVMVSQGLGLALGAAI 359
Query: 601 PHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYWIWFHYLSLVKYPYEAVLQNEFDDPVK 660
T+ + F+L G+++ ++PS W Y+S Y Y + + +++D K
Sbjct: 360 MDAKQASTVAAVTMLAFVLTGGYYV--HKVPSCMAWIKYISTTFYCYRLLTRIQYEDGKK 533
Query: 661 CFVRGVQIFDNSPLRSVPYELKLKLLGSMSGTLGTNITATTCLTTGADILQQNGVTELSK 720
+ Y LLG G G G ++++ V ++
Sbjct: 534 ----------------ISY-----LLGCYHGDKG-----------GCRFVEEDVVGQIGT 617
Query: 721 WNCLWITVAWGFFFRFLFYLAL 742
C+ + + F+R L YLAL
Sbjct: 618 LGCIGVLLFMFVFYRLLAYLAL 683
>BI699796 similar to PIR|G85167|G85 ABC transporter like protein [imported] -
Arabidopsis thaliana, partial (13%)
Length = 421
Score = 79.0 bits (193), Expect = 7e-15
Identities = 37/119 (31%), Positives = 73/119 (61%)
Frame = +1
Query: 186 ISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIG 245
+S Y Q+D+ P +TVEE++ F+A RLP + K V +I + L +++G
Sbjct: 55 VSGYCEQNDIHSPNITVEESVMFSAWLRLPSQIDAKTKAEFVNEVIHTIELDGIKDSLVG 234
Query: 246 DEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIV 304
G+S +R+R++I ++++ +P ++F+DEPT+GLD+ +A +V++ ++ + +G V
Sbjct: 235 MPNISGLSTEQRKRLTIAVELVANPSIIFMDEPTTGLDARAAAVVMRAVKNVVGTGRTV 411
>TC208020 similar to UP|Q76CU2 (Q76CU2) PDR-type ABC transporter 1, partial
(14%)
Length = 765
Score = 77.8 bits (190), Expect = 2e-14
Identities = 42/126 (33%), Positives = 74/126 (58%), Gaps = 5/126 (3%)
Frame = +2
Query: 227 VQALIDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTS 286
V L++ LRN+ ++G G G+S +R+R++I ++++ +P ++F+DEPTSGLD+ +
Sbjct: 173 VMELVELNPLRNS---LVGLPGVSGLSTEQRKRLTIAVELVANPSIIFMDEPTSGLDARA 343
Query: 287 AFMVVKVLQRIAQSGSIVIMSIHQPSYRILGLLDRMIFLSR-GQTVYSG----SPTQLPS 341
A +V++ ++ +G V+ +IHQPS I D + + R GQ +Y G T L
Sbjct: 344 AAIVMRTVRNTVDTGRTVVCTIHQPSIDIFEAFDELFLMKRGGQEIYVGPLGRHSTHLIK 523
Query: 342 FFAEFG 347
+F G
Sbjct: 524 YFESIG 541
>BQ080393
Length = 347
Score = 75.5 bits (184), Expect = 8e-14
Identities = 35/114 (30%), Positives = 68/114 (58%)
Frame = +2
Query: 167 LKGTLALNGEALESRLLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKAR 226
++G + ++G +S Y Q D+ P +T+ E+L ++A RLP+ +SK +K
Sbjct: 5 IEGDIRISGFPKNQETFARVSGYCEQTDIHSPQVTIRESLLYSAYLRLPKEVSKDEKIQF 184
Query: 227 VQALIDQLGLRNAAKTVIGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTS 280
V ++D + L N ++G G G+S +R+R++I ++++ +P ++F+DEPTS
Sbjct: 185 VDQVMDLVELDNLKDAIVGLPGVTGLSTEQRKRLTIAVELVANPSIIFMDEPTS 346
>TC229309 homologue to UP|Q9LJX0 (Q9LJX0) P-glycoprotein; multi-drug
resistance related; ABC transporter-like protein,
partial (22%)
Length = 841
Score = 73.6 bits (179), Expect = 3e-13
Identities = 52/174 (29%), Positives = 95/174 (53%), Gaps = 3/174 (1%)
Frame = +1
Query: 127 LLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALESRLLKVI 186
+ D++ R G+ A++GASGSGKS++I AL R + G + ++G+ + LK +
Sbjct: 322 VFKDLNLRIRAGQSQALVGASGSGKSSVI-ALIERFYDP-IAGKVMVDGKDIRKLNLKSL 495
Query: 187 S---AYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTV 243
V Q+ LF ++ E + + E + ++ + A V + GL KT
Sbjct: 496 RLKIGLVQQEPALFAA-SIFENIAYGKEGATEAEVIEAARAANVHGFVS--GLPEGYKTP 666
Query: 244 IGDEGHRGVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRI 297
+G+ G + +SGG+++R++I ++ DP +L LDE TS LD+ S ++ + L+R+
Sbjct: 667 VGERGVQ-LSGGQKQRIAIARAVLKDPTILLLDEATSALDAESECVLQEALERL 825
>TC217803 similar to UP|Q9AT00 (Q9AT00) At1g65410/T8F5_19, partial (81%)
Length = 1563
Score = 73.6 bits (179), Expect = 3e-13
Identities = 63/232 (27%), Positives = 112/232 (48%), Gaps = 22/232 (9%)
Frame = +3
Query: 125 KTLLNDISGEARDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGE-----ALE 179
K +LN +S + + GE + ++G SG+GKST++ +A +A KG + + G+ +
Sbjct: 378 KKILNGVSFKIKHGEAVGIIGPSGTGKSTVLKIIAGLLAPD--KGEVYIRGKKRVGLVSD 551
Query: 180 SRLLKVISAYVMQDDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNA 239
+ + V Q LF LTV E + F ++S+ + V + +GL+
Sbjct: 552 DDISGLRIGLVFQSAALFDSLTVRENVGFLLYEH--SSMSEDQISELVTETLAAVGLKG- 722
Query: 240 AKTVIGDEGHRGVSGGERRRVSIGIDIIHD-------PILLFLDEPTSGLDSTSAFMVVK 292
+ D +SGG ++RV++ II D P +L DEPT+GLD ++ +V
Sbjct: 723 ----VEDRLPSELSGGMKKRVALARSIICDTTKESIEPEVLLYDEPTAGLDPIASTVVED 890
Query: 293 VLQRIAQSG----------SIVIMSIHQPSYRILGLLDRMIFLSRGQTVYSG 334
+++ + G S ++ HQ S I +DR++FL +G+ V+ G
Sbjct: 891 LIRSVHIKGQDARGKPGNISSYVVVTHQHS-TIKRAIDRLLFLHKGKIVWEG 1043
>TC222012 similar to UP|Q9FLX5 (Q9FLX5) ABC transporter-like protein, partial
(24%)
Length = 503
Score = 72.8 bits (177), Expect = 5e-13
Identities = 33/91 (36%), Positives = 58/91 (63%)
Frame = +1
Query: 491 KGVQERLGFFAFAMSTTFYTTADALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPP 550
+G+++R G FAF ++ +T + LP+F+ ER I +RET+ YR SSYLI++ LV LP
Sbjct: 229 EGIEKRFGLFAFTLTFLLSSTTETLPIFINERPILLRETSSGVYRLSSYLIANTLVFLPY 408
Query: 551 LAFLSLAFAVITFWAVGLDGGASGFLFYFLI 581
L +++ +++ ++ VGL F ++ L+
Sbjct: 409 LFVVAVIYSIPVYFLVGLCASWLSFAYFVLV 501
>TC218388 similar to UP|Q9LMU4 (Q9LMU4) F2H15.7 protein, partial (38%)
Length = 933
Score = 69.7 bits (169), Expect = 4e-12
Identities = 48/178 (26%), Positives = 92/178 (50%), Gaps = 2/178 (1%)
Frame = +1
Query: 509 YTTADALPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGL 568
+ + P F+++ +F RE Y +S++IS+ L ++P L ++ I ++ V L
Sbjct: 7 FMSIGGFPSFVEDMKVFQRERLNGHYGVTSFVISNTLSAMPFLILITFLSGTICYFMVRL 186
Query: 569 DGGASGFLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRD 628
G +LF+ L ++AS S + ++ +VP+ ++G I I F+L+SG+F
Sbjct: 187 HPGFWHYLFFVLCLYASVTVVESLMMAIASIVPNFLMGIIIGAGIQGIFMLVSGYFRLPH 366
Query: 629 RIPSYWIWFHYLSLVKYPYEAVLQNEFDDPVKCFVRGVQIFDNS--PLRSVPYELKLK 684
IP +W + +S + + + A LQ ++ + +RG+ +FDN L +P E L+
Sbjct: 367 DIPKP-VWRYPMSYISFHFWA-LQGQYQND----LRGL-VFDNQTPDLPKIPGEYILE 519
>TC213020 weakly similar to UP|Q8GU71 (Q8GU71) MDR-like ABC transporter,
partial (18%)
Length = 864
Score = 69.3 bits (168), Expect = 6e-12
Identities = 54/174 (31%), Positives = 89/174 (51%), Gaps = 4/174 (2%)
Frame = +1
Query: 138 GEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEA----LESRLLKVISAYVMQD 193
G+ +AV+G SGSGKST+I + L L E L R L++ V Q+
Sbjct: 28 GKSLAVVGQSGSGKSTVISLVMRFYDPD---SGLVLVDECDIKNLNLRSLRLRIGLVQQE 198
Query: 194 DLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGDEGHRGVS 253
LF TV E + + E + K+ K A I ++ KT +G+ G + +S
Sbjct: 199 PALFST-TVYENIKYGKEEASEIEVMKAAKAANAHEFISRMP--EGYKTEVGERGVQ-LS 366
Query: 254 GGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIMS 307
GG+++RV+I I+ DP +L LDE TS LD+ S +V + L ++ + + ++++
Sbjct: 367 GGQKQRVAIARAILKDPSILLLDEATSALDTVSERLVQEALDKLMEGRTTILVA 528
>TC207596 similar to UP|Q7GBY1 (Q7GBY1) Tonoplast ABC transporter IDI7,
partial (35%)
Length = 1076
Score = 68.6 bits (166), Expect = 1e-11
Identities = 62/209 (29%), Positives = 107/209 (50%), Gaps = 7/209 (3%)
Frame = +2
Query: 138 GEIMAVLGASGSGKSTLIDALANRIAK--GRLKGTLALNG----EALESRLLKVISAYVM 191
G +A++G SG GKST+ AN I + KG + LNG E L + IS V
Sbjct: 20 GSKVALVGPSGGGKSTI----ANLIERFYDPTKGKILLNGVPLVEISHKHLHRKISI-VS 184
Query: 192 QDDLLFPMLTVEETLTFAAEFRLPRT-LSKSKKKARVQALIDQLGLRNAAKTVIGDEGHR 250
Q+ LF ++EE + + + ++ + + K A I + + +T +G+ G R
Sbjct: 185 QEPTLFNC-SIEENIAYGFDGKVNDVDIENAAKMANAHEFISKFPEKY--QTFVGERGVR 355
Query: 251 GVSGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIMSIHQ 310
+SGG+++R++I ++ DP +L LDE TS LD+ S ++V ++ + G V++ H+
Sbjct: 356 -LSGGQKQRIAIARALLMDPKILLLDEATSALDAESEYLVQDAMESL-MKGRTVLVIAHR 529
Query: 311 PSYRILGLLDRMIFLSRGQTVYSGSPTQL 339
S + D + +S GQ V G+ +L
Sbjct: 530 LS--TVKTADTVAVISDGQVVERGNHEEL 610
>CO983514
Length = 575
Score = 68.6 bits (166), Expect = 1e-11
Identities = 35/143 (24%), Positives = 65/143 (44%)
Frame = -2
Query: 515 LPVFLQERYIFMRETAYNAYRRSSYLISHALVSLPPLAFLSLAFAVITFWAVGLDGGASG 574
LP ER + RE Y +Y + L+ +P + ++ + +IT+ + D A
Sbjct: 538 LPYVATERTVLYRERFAGMYSPWAYSFAQVLIEVPYIFIQAVVYVIITYPMLSYDWSAYK 359
Query: 575 FLFYFLIIFASFWAGNSFVTFLSGVVPHVMLGYTIVVAILAYFLLLSGFFIDRDRIPSYW 634
+ F +F + N + + P+V L + + L SG+F+ R RIP +W
Sbjct: 358 IFWSFFSMFCNILYYNYLGMLIVSLTPNVQLAAIVASSSYTMLNLFSGYFVPRLRIPKWW 179
Query: 635 IWFHYLSLVKYPYEAVLQNEFDD 657
IW +YL + + +L +++ D
Sbjct: 178 IWMYYLCPMSWALNGMLTSQYGD 110
>TC207563 similar to UP|Q9ZRG2 (Q9ZRG2) P-glycoprotein, partial (24%)
Length = 1164
Score = 65.9 bits (159), Expect = 6e-11
Identities = 58/207 (28%), Positives = 102/207 (49%), Gaps = 3/207 (1%)
Frame = +1
Query: 136 RDGEIMAVLGASGSGKSTLIDALANRIAKGRLKGTLALNGEALESRLLKVISAYVM---Q 192
R G+ +A++G SG GKS++I AL R G + ++G+ + LK + ++ Q
Sbjct: 244 RAGKTLALVGPSGCGKSSII-ALIQRFYDPT-SGRVMIDGKDIRKYNLKSLRRHISVVPQ 417
Query: 193 DDLLFPMLTVEETLTFAAEFRLPRTLSKSKKKARVQALIDQLGLRNAAKTVIGDEGHRGV 252
+ LF T+ E + + E + ++ A I GL + KT +G+ G + +
Sbjct: 418 EPCLFAT-TIYENIAYGHESATEAEIIEAATLANAHKFIS--GLPDGYKTFVGERGVQ-L 585
Query: 253 SGGERRRVSIGIDIIHDPILLFLDEPTSGLDSTSAFMVVKVLQRIAQSGSIVIMSIHQPS 312
SGG+++R+++ + L+ LDE TS LD+ S V + L R A SG I+ H+ S
Sbjct: 586 SGGQKQRIAVARAFVRKAELMLLDEATSALDAESERSVQEALDR-ASSGKTTIIVAHRLS 762
Query: 313 YRILGLLDRMIFLSRGQTVYSGSPTQL 339
+ + + + G+ GS +QL
Sbjct: 763 --TIRNANLIAVIDDGKVAEQGSHSQL 837
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.137 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,218,606
Number of Sequences: 63676
Number of extensions: 392468
Number of successful extensions: 2789
Number of sequences better than 10.0: 132
Number of HSP's better than 10.0 without gapping: 2714
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2757
length of query: 751
length of database: 12,639,632
effective HSP length: 104
effective length of query: 647
effective length of database: 6,017,328
effective search space: 3893211216
effective search space used: 3893211216
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0260.10


