

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0258b.1
(207 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
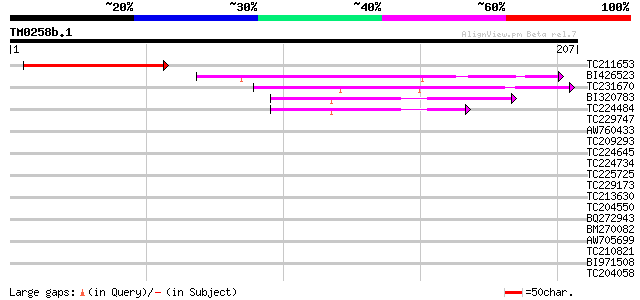
Sequences producing significant alignments: (bits) Value
TC211653 similar to UP|Q38848 (Q38848) LRP1, partial (27%) 83 8e-17
BI426523 similar to GP|10177429|dbj zinc finger protein SHI-like... 74 5e-14
TC231670 similar to UP|Q9LQZ5 (Q9LQZ5) F10A5.26, partial (15%) 64 5e-11
BI320783 53 1e-07
TC224484 similar to UP|Q38848 (Q38848) LRP1, partial (20%) 52 2e-07
TC229747 32 0.27
AW760433 similar to PIR|H86398|H863 protein F17L21.10 [imported]... 30 0.59
TC209293 homologue to UP|RN12_HUMAN (Q9NVW2) RING finger protein... 30 0.59
TC224645 homologue to UP|Q41707 (Q41707) Extensin class 1 protei... 30 0.78
TC224734 homologue to UP|Q09085 (Q09085) Hydroxyproline-rich gly... 30 0.78
TC225725 similar to UP|Q9FR29 (Q9FR29) Transcription factor WRKY... 30 0.78
TC229173 30 1.0
TC213630 30 1.0
TC204550 UP|Q8L3Y5 (Q8L3Y5) Receptor-like kinase RHG1, complete 29 1.3
BQ272943 29 1.3
BM270082 29 1.7
AW705699 29 1.7
TC210821 similar to UP|O22265 (O22265) Expressed protein (At2g47... 29 1.7
BI971508 28 2.3
TC204058 similar to UP|Q39835 (Q39835) Extensin, partial (37%) 28 2.3
>TC211653 similar to UP|Q38848 (Q38848) LRP1, partial (27%)
Length = 515
Score = 83.2 bits (204), Expect = 8e-17
Identities = 33/53 (62%), Positives = 39/53 (73%)
Frame = +3
Query: 6 GQEVKGSRCQECGNQAKKGCAYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHR 58
G G+ CQ+CGNQAKK CA RCRTCC ++GF C THV+STW+P RRR R
Sbjct: 291 GTSSGGTTCQDCGNQAKKDCANRRCRTCCKSRGFDCPTHVKSTWVPAARRRER 449
>BI426523 similar to GP|10177429|dbj zinc finger protein SHI-like
{Arabidopsis thaliana}, partial (16%)
Length = 421
Score = 73.9 bits (180), Expect = 5e-14
Identities = 54/142 (38%), Positives = 68/142 (47%), Gaps = 8/142 (5%)
Frame = +3
Query: 69 NNPHHLHEDIPQSHN----QNPFTSLEELKFPEAMSSMAVFSSVRVRSMDDSVNEMAYQT 124
N P H D S P T LE +FP +S+ A+F VRV ++D S + AYQT
Sbjct: 3 NIPKRHHPDTTTSTQLASAPQPVTGLELGQFPAEVSTSALFRCVRVSAVDASDEQYAYQT 182
Query: 125 SVNIGGHRFSGILYDQGPEQQSLNA----SPLDQHQNLNLTTIHSHDGATMAPPSSSATA 180
SVNIGGH F G LYDQGPE A S + Q L T AT A S ++
Sbjct: 183 SVNIGGHVFKGFLYDQGPESSYTGAAAEGSSGGEPQQLGFITA----AATTATTSGNSPF 350
Query: 181 AHELFFPQPRSLASFRSGVPYF 202
L+ P L +F +G +F
Sbjct: 351 DPSLY---PAPLNAFMAGTQFF 407
>TC231670 similar to UP|Q9LQZ5 (Q9LQZ5) F10A5.26, partial (15%)
Length = 672
Score = 63.9 bits (154), Expect = 5e-11
Identities = 41/122 (33%), Positives = 60/122 (48%), Gaps = 5/122 (4%)
Frame = +1
Query: 90 LEELKFPEAMSSMAVFSSVRVRSMDDSVNE-MAYQTSVNIGGHRFSGILYDQGPEQQSLN 148
LEE+ P + S A F VRV SMD+ E AY T+VNI GH F GILYD GPE + N
Sbjct: 136 LEEVNCPAVVRSAAEFRCVRVSSMDEEAEEEYAYSTAVNIAGHVFKGILYDYGPEGMNTN 315
Query: 149 ----ASPLDQHQNLNLTTIHSHDGATMAPPSSSATAAHELFFPQPRSLASFRSGVPYFSH 204
+ + + + ++ GA ++ P ++ +P P + SG +F H
Sbjct: 316 YMDAVAAAGESSSTGVGALNLTTGAIVSEPLGVDPSS---LYPAPLNSFMPGSGTQFFPH 486
Query: 205 TK 206
+
Sbjct: 487 PR 492
>BI320783
Length = 421
Score = 52.8 bits (125), Expect = 1e-07
Identities = 33/91 (36%), Positives = 47/91 (51%), Gaps = 1/91 (1%)
Frame = +3
Query: 96 PEAMSSMAVFSSVRVRSMDDS-VNEMAYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQ 154
P + + AVF VRV ++DD +E AYQ V IGGH F G LYDQG E D+
Sbjct: 51 PCQVRAPAVFKCVRVTAVDDGGEDEYAYQAVVKIGGHVFKGFLYDQGVE---------DK 203
Query: 155 HQNLNLTTIHSHDGATMAPPSSSATAAHELF 185
NL+ +H G ++ S +H+++
Sbjct: 204 EGYPNLSELHLGGGNGVSSSSPMVDPSHDVY 296
>TC224484 similar to UP|Q38848 (Q38848) LRP1, partial (20%)
Length = 574
Score = 51.6 bits (122), Expect = 2e-07
Identities = 32/74 (43%), Positives = 39/74 (52%), Gaps = 1/74 (1%)
Frame = -3
Query: 96 PEAMSSMAVFSSVRVRSMDDS-VNEMAYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQ 154
P + + AVF VRV S+DD +E AYQ V IGGH F G LYDQG E D+
Sbjct: 524 PSQVRAPAVFKCVRVTSVDDGGEDEYAYQAVVKIGGHVFKGFLYDQGVE---------DK 372
Query: 155 HQNLNLTTIHSHDG 168
NL+ +H G
Sbjct: 371 EGYPNLSELHLSGG 330
>TC229747
Length = 1327
Score = 31.6 bits (70), Expect = 0.27
Identities = 20/81 (24%), Positives = 33/81 (40%), Gaps = 2/81 (2%)
Frame = +3
Query: 109 RVRSMDDSVNEMAYQTSVNIGGHRFSGILYDQGPEQQSLNASPLDQHQN--LNLTTIHSH 166
R+ DD++ MA+ S GG G Y +G E P D+ +++ +
Sbjct: 903 RISKTDDNLLSMAHTFSKGDGGFMLMGHNYGKGDESIVSMGQPFDKGDGNFISMGQSYEK 1082
Query: 167 DGATMAPPSSSATAAHELFFP 187
+ + +S T HE F P
Sbjct: 1083EDGNLISLGTSYTKVHESFIP 1145
>AW760433 similar to PIR|H86398|H863 protein F17L21.10 [imported] -
Arabidopsis thaliana, partial (55%)
Length = 358
Score = 30.4 bits (67), Expect = 0.59
Identities = 14/39 (35%), Positives = 20/39 (50%)
Frame = +1
Query: 56 RHRLMEHQPPPTTNNPHHLHEDIPQSHNQNPFTSLEELK 94
+HR H PP +T P +LH +P +Q+ T L K
Sbjct: 127 QHRYQAHLPPFSTXPPLNLHHQLPTLAHQHQHTHLRXQK 243
>TC209293 homologue to UP|RN12_HUMAN (Q9NVW2) RING finger protein 12 (LIM
domain interacting RING finger protein) (RING finger LIM
domain-binding protein) (R-LIM) (NY-REN-43 antigen),
partial (4%)
Length = 957
Score = 30.4 bits (67), Expect = 0.59
Identities = 14/23 (60%), Positives = 15/23 (64%), Gaps = 1/23 (4%)
Frame = -1
Query: 54 RRRHRLMEHQP-PPTTNNPHHLH 75
R HRL+EHQP PP N HH H
Sbjct: 384 RACHRLLEHQP*PPNHPNIHHHH 316
>TC224645 homologue to UP|Q41707 (Q41707) Extensin class 1 protein precursor,
partial (58%)
Length = 856
Score = 30.0 bits (66), Expect = 0.78
Identities = 16/45 (35%), Positives = 18/45 (39%), Gaps = 4/45 (8%)
Frame = +3
Query: 42 QTHVRSTWIPVDRRRHRLMEHQPPPTT----NNPHHLHEDIPQSH 82
QTH H L H PP TT ++PHH H SH
Sbjct: 540 QTHHHHLLTTTSLHLHHLHHHHPPTTTSHLLHHPHHHHLHTTTSH 674
>TC224734 homologue to UP|Q09085 (Q09085) Hydroxyproline-rich glycoprotein
(HRGP) (Fragment), partial (48%)
Length = 990
Score = 30.0 bits (66), Expect = 0.78
Identities = 16/45 (35%), Positives = 18/45 (39%), Gaps = 4/45 (8%)
Frame = +3
Query: 42 QTHVRSTWIPVDRRRHRLMEHQPPPTT----NNPHHLHEDIPQSH 82
QTH H L H PP TT ++PHH H SH
Sbjct: 87 QTHHHHLLTTTSLHLHHLHHHHPPTTTSHLLHHPHHHHLHTTTSH 221
>TC225725 similar to UP|Q9FR29 (Q9FR29) Transcription factor WRKY4, partial
(51%)
Length = 1012
Score = 30.0 bits (66), Expect = 0.78
Identities = 11/21 (52%), Positives = 15/21 (71%)
Frame = +2
Query: 70 NPHHLHEDIPQSHNQNPFTSL 90
NPH LHE++P+ +N F SL
Sbjct: 2 NPHRLHEELPKKEVENNFLSL 64
>TC229173
Length = 924
Score = 29.6 bits (65), Expect = 1.0
Identities = 12/25 (48%), Positives = 14/25 (56%)
Frame = +2
Query: 64 PPPTTNNPHHLHEDIPQSHNQNPFT 88
PPP T++ HH H P H Q P T
Sbjct: 179 PPPPTSHWHHFHHLKPCQHPQTPTT 253
>TC213630
Length = 847
Score = 29.6 bits (65), Expect = 1.0
Identities = 10/24 (41%), Positives = 16/24 (66%)
Frame = -3
Query: 66 PTTNNPHHLHEDIPQSHNQNPFTS 89
P +NPHH+ ++ Q+HN + F S
Sbjct: 317 PLISNPHHIGNNVGQTHNNHCFVS 246
>TC204550 UP|Q8L3Y5 (Q8L3Y5) Receptor-like kinase RHG1, complete
Length = 2659
Score = 29.3 bits (64), Expect = 1.3
Identities = 14/46 (30%), Positives = 21/46 (45%)
Frame = -1
Query: 26 AYSRCRTCCNNKGFQCQTHVRSTWIPVDRRRHRLMEHQPPPTTNNP 71
A S +CC + C++H V + L++HQPP T P
Sbjct: 1825 ALSHDLSCCTQEIISCKSHWSIKMD*VPLQPPHLLQHQPPATGGTP 1688
>BQ272943
Length = 420
Score = 29.3 bits (64), Expect = 1.3
Identities = 18/45 (40%), Positives = 25/45 (55%)
Frame = +2
Query: 45 VRSTWIPVDRRRHRLMEHQPPPTTNNPHHLHEDIPQSHNQNPFTS 89
VR+T P +RH+L E + TT NPHH H+ ++ FTS
Sbjct: 11 VRATPNP---QRHQLHESRTRFTTTNPHHHHDFCDLNNCCERFTS 136
>BM270082
Length = 421
Score = 28.9 bits (63), Expect = 1.7
Identities = 16/54 (29%), Positives = 25/54 (45%)
Frame = +1
Query: 154 QHQNLNLTTIHSHDGATMAPPSSSATAAHELFFPQPRSLASFRSGVPYFSHTKP 207
Q +L+LT ++H + PP +S EL Q S +G P+ H+ P
Sbjct: 103 QSNHLSLTLTYTHI*TRIVPPMASKRILKELKDLQKDPPTSCSAGTPFLLHSTP 264
>AW705699
Length = 238
Score = 28.9 bits (63), Expect = 1.7
Identities = 13/41 (31%), Positives = 21/41 (50%)
Frame = +1
Query: 54 RRRHRLMEHQPPPTTNNPHHLHEDIPQSHNQNPFTSLEELK 94
R +H+ H PP +T P +LH +P +Q+ L + K
Sbjct: 1 RFQHQCQAHLPPFSTVPPLNLHRQLPXLQHQHQHARLRQQK 123
>TC210821 similar to UP|O22265 (O22265) Expressed protein
(At2g47450/T30B22.25), partial (29%)
Length = 639
Score = 28.9 bits (63), Expect = 1.7
Identities = 26/85 (30%), Positives = 35/85 (40%), Gaps = 17/85 (20%)
Frame = -3
Query: 51 PVDRRRHR-----LMEHQPPPTTNNP---HHLHEDIPQSHNQNPFTSLEE---------L 93
P RRR R L + PP T+ +P HH + N +PF S+ E L
Sbjct: 295 PPQRRRTRGQPRSLSPNPPPQTSPSPTRWHHRPSTAQDTPNSHPFLSVPESPGPPRTRTL 116
Query: 94 KFPEAMSSMAVFSSVRVRSMDDSVN 118
P+ S + V S VR S S +
Sbjct: 115 PLPKP*SHLPVQSCVRTASDSPSAS 41
>BI971508
Length = 713
Score = 28.5 bits (62), Expect = 2.3
Identities = 20/62 (32%), Positives = 34/62 (54%)
Frame = -2
Query: 49 WIPVDRRRHRLMEHQPPPTTNNPHHLHEDIPQSHNQNPFTSLEELKFPEAMSSMAVFSSV 108
W+ + R L+ +Q P T+ ++ ++ + +NQNP SL+E FP S A+ SS
Sbjct: 391 WLLLMRGSLCLL*YQNIPPTSERLYIPKNSQRMYNQNPIISLDE--FP----SQALSSSQ 230
Query: 109 RV 110
R+
Sbjct: 229 RI 224
>TC204058 similar to UP|Q39835 (Q39835) Extensin, partial (37%)
Length = 852
Score = 28.5 bits (62), Expect = 2.3
Identities = 31/123 (25%), Positives = 48/123 (38%), Gaps = 18/123 (14%)
Frame = +1
Query: 62 HQPPPTTNNPHHLHEDIPQSHNQNPFTSL---EELKFPEAMSSMAVFSSVRVRSMDDSVN 118
H PT + HH H+ SH+ +P ++L LK P ++ + S N
Sbjct: 202 HLRSPTNTHHHHHHQFTNTSHHHHPTSTLLLHHHLKSPT--------NTHLLHLQFTSTN 357
Query: 119 EMAYQTSVNIGGHRF------------SGILYDQ--GPEQQSLNASPLDQHQNL-NLTTI 163
+ TS ++ H + +L Q L+ S L HQ+L N T+I
Sbjct: 358 HHPHHTSTHLLHHHLTSTHLHHPQCTSTSLLLHQITSTSHLHLHTSTLHLHQHLRNHTSI 537
Query: 164 HSH 166
H H
Sbjct: 538 HLH 546
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.129 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,023,754
Number of Sequences: 63676
Number of extensions: 228433
Number of successful extensions: 2127
Number of sequences better than 10.0: 95
Number of HSP's better than 10.0 without gapping: 2056
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2111
length of query: 207
length of database: 12,639,632
effective HSP length: 93
effective length of query: 114
effective length of database: 6,717,764
effective search space: 765825096
effective search space used: 765825096
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0258b.1


