

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0258a.2
(167 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
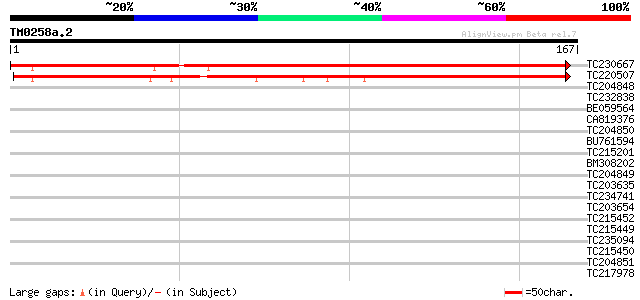
Score E
Sequences producing significant alignments: (bits) Value
TC230667 weakly similar to UP|AGA1_YEAST (P32323) A-agglutinin a... 180 2e-46
TC220507 similar to UP|Q7Q3J9 (Q7Q3J9) AgCP11343 (Fragment), par... 171 1e-43
TC204848 similar to UP|Q6ICZ8 (Q6ICZ8) At5g13850, partial (89%) 38 0.002
TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {A... 35 0.013
BE059564 35 0.022
CA819376 similar to GP|22654971|gb| At1g80930/F23A5_23 {Arabidop... 34 0.037
TC204850 similar to GB|AAM16178.1|20334844|AY094022 AT3g12390/T2... 33 0.064
BU761594 similar to GP|22654971|gb At1g80930/F23A5_23 {Arabidops... 33 0.083
TC215201 similar to UP|Q70Z19 (Q70Z19) Nucleosome assembly prote... 32 0.14
BM308202 32 0.19
TC204849 similar to GB|AAM16178.1|20334844|AY094022 AT3g12390/T2... 32 0.19
TC203635 similar to PIR|T46225|T46225 alpha NAC-like protein - A... 32 0.19
TC234741 weakly similar to UP|Q93ZH7 (Q93ZH7) AT5g03040/F15A17_7... 32 0.19
TC203654 similar to PIR|T46225|T46225 alpha NAC-like protein - A... 31 0.24
TC215452 similar to UP|P93488 (P93488) NAP1Ps, partial (29%) 31 0.32
TC215449 similar to UP|P93488 (P93488) NAP1Ps, partial (97%) 31 0.32
TC235094 similar to UP|PRT3_SCYCA (P30258) Protamine Z3 (Scyllio... 30 0.41
TC215450 similar to UP|P93488 (P93488) NAP1Ps, partial (30%) 30 0.41
TC204851 similar to GB|AAM16178.1|20334844|AY094022 AT3g12390/T2... 30 0.54
TC217978 weakly similar to UP|Q8VYJ8 (Q8VYJ8) AT4g27310/M4I22_12... 30 0.54
>TC230667 weakly similar to UP|AGA1_YEAST (P32323) A-agglutinin attachment
subunit precursor, partial (5%)
Length = 865
Score = 180 bits (457), Expect = 2e-46
Identities = 103/181 (56%), Positives = 121/181 (65%), Gaps = 16/181 (8%)
Frame = -1
Query: 1 MENQP---MQFLLLEQVQFLKVQDDSLLQWELVDVVDAEEEQEQ-----------EPDQN 46
MENQ MQFL+LEQVQFLKV DDSLLQWELVDVVDAEEE + E D +
Sbjct: 796 MENQASSSMQFLILEQVQFLKVNDDSLLQWELVDVVDAEEEIDGFGSSDSSVSWFESDSS 617
Query: 47 QQDAAPVEEIK--IIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAYEDDEEQVVDDEEDD 104
DA P+EEI+ ++H + D V E D ++ H + + DD++ DD ++D
Sbjct: 616 PCDA-PIEEIRHLLLHPDVGLQDRHDGVDESEQDSGLENHHHYHYHHDDDDMGGDDNDED 440
Query: 105 DDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKNSKRSPYLFVRPGCVGGKHGLGLK 164
DD LDDELVPW V NK GRQRM+KLGKRA SKM SK+SPYLFVRPGCV GKHGLGLK
Sbjct: 439 DDGYGLDDELVPWNVSNKFGRQRMRKLGKRAFSKMHGSKKSPYLFVRPGCVRGKHGLGLK 260
Query: 165 H 165
H
Sbjct: 259 H 257
>TC220507 similar to UP|Q7Q3J9 (Q7Q3J9) AgCP11343 (Fragment), partial (3%)
Length = 829
Score = 171 bits (433), Expect = 1e-43
Identities = 103/186 (55%), Positives = 123/186 (65%), Gaps = 22/186 (11%)
Frame = +3
Query: 2 ENQP--MQFLLLEQVQFLKVQDDSLLQWELVDVVDAEEEQE---QEPDQN-----QQDAA 51
ENQ MQFL+LEQVQFLKV DDSLLQWELVDVVDAEEE + D + + D++
Sbjct: 60 ENQTSSMQFLILEQVQFLKVNDDSLLQWELVDVVDAEEEIDGFGSSSDSSSVSWFESDSS 239
Query: 52 PVEEIKIIHQIRPVLPDPDA------VAVVEIDRQQYHYL---HGYAYED-DEEQVVDDE 101
P++ I +IR +L PD + E + +HY H Y ++D D++ V D
Sbjct: 240 PIDAS--IEEIRQLLLHPDVGLQDPRESEQESGLENHHYRYHQHHYHHDDVDDDDVGGDN 413
Query: 102 ED--DDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKNSKRSPYLFVRPGCVGGKH 159
+D DDD LDDELVPW V NK RQRM+KLGKRA SKM NSK+SPYLFVRPGCV GKH
Sbjct: 414 DDGEDDDGYGLDDELVPWNVSNKFERQRMRKLGKRAFSKMHNSKKSPYLFVRPGCVRGKH 593
Query: 160 GLGLKH 165
GLGLKH
Sbjct: 594 GLGLKH 611
>TC204848 similar to UP|Q6ICZ8 (Q6ICZ8) At5g13850, partial (89%)
Length = 916
Score = 38.1 bits (87), Expect = 0.002
Identities = 20/70 (28%), Positives = 37/70 (52%)
Frame = +3
Query: 70 DAVAVVEIDRQQYHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMK 129
+ + ++++Q+ H DDE V DD+EDD+D D DD+ V + + + R +
Sbjct: 96 EELLAADLEQQKIH--------DDEPVVEDDDEDDEDDDDEDDDNVEGDASGRSKQTRSE 251
Query: 130 KLGKRACSKM 139
K ++A K+
Sbjct: 252 KKSRKAMLKL 281
>TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;} , partial (46%)
Length = 902
Score = 35.4 bits (80), Expect = 0.013
Identities = 27/103 (26%), Positives = 48/103 (46%), Gaps = 2/103 (1%)
Frame = +2
Query: 13 QVQFLKVQDDSLLQW--ELVDVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPD 70
+V+ L +DD + E+VD + +E++E++ D ++ D EE PD
Sbjct: 263 EVKVLVEEDDEEYESVDEVVDDAEDDEDEEEDDDDDEGDDNDDEEDDA--------PDGG 418
Query: 71 AVAVVEIDRQQYHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDE 113
E D ++ G +DD+ DD +DD+D + D+E
Sbjct: 419 DDDDDEDDEEEGDVQRGGEPDDDDNDSDDDSDDDEDEDEEDEE 547
Score = 29.3 bits (64), Expect = 0.92
Identities = 12/59 (20%), Positives = 33/59 (55%)
Frame = +2
Query: 55 EIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDE 113
++K ++ + + + +VE D ++Y + + ++++ ++++DDD+ D DDE
Sbjct: 218 KMKATKELSEQVEENEVKVLVEEDDEEYESVDEVVDDAEDDEDEEEDDDDDEGDDNDDE 394
>BE059564
Length = 406
Score = 34.7 bits (78), Expect = 0.022
Identities = 15/30 (50%), Positives = 22/30 (73%)
Frame = +3
Query: 87 GYAYEDDEEQVVDDEEDDDDRCDLDDELVP 116
G +++DD++ DDEE+DDD D D+E VP
Sbjct: 180 GLSHDDDDDDETDDEEEDDDD-DEDEEYVP 266
>CA819376 similar to GP|22654971|gb| At1g80930/F23A5_23 {Arabidopsis
thaliana}, partial (10%)
Length = 449
Score = 33.9 bits (76), Expect = 0.037
Identities = 25/84 (29%), Positives = 39/84 (45%)
Frame = +1
Query: 30 VDVVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYA 89
V V E + ++ DQ + + EEI + PDP+ + + + + G
Sbjct: 139 VSAVRPELDLAEQEDQITHEVSLDEEIDPEISLDIFKPDPNFLENEKCYEELKKSMLGEE 318
Query: 90 YEDDEEQVVDDEEDDDDRCDLDDE 113
+EDDEE +D E DDDD + DE
Sbjct: 319 FEDDEEG-LDAESDDDDEDEESDE 387
>TC204850 similar to GB|AAM16178.1|20334844|AY094022 AT3g12390/T2E22_130
{Arabidopsis thaliana;} , partial (85%)
Length = 878
Score = 33.1 bits (74), Expect = 0.064
Identities = 17/63 (26%), Positives = 35/63 (54%)
Frame = +1
Query: 77 IDRQQYHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRAC 136
+++Q+ H +DD+E+V DD++D+DD DE + + + + R +K ++A
Sbjct: 91 LEQQKIHDDEPVVEDDDDEEVDDDDDDEDDD---HDEGLEGDASGRSKQTRSEKKSRKAM 261
Query: 137 SKM 139
K+
Sbjct: 262 LKL 270
>BU761594 similar to GP|22654971|gb At1g80930/F23A5_23 {Arabidopsis
thaliana}, partial (13%)
Length = 397
Score = 32.7 bits (73), Expect = 0.083
Identities = 22/78 (28%), Positives = 36/78 (45%), Gaps = 6/78 (7%)
Frame = +1
Query: 42 EPDQNQQDAAPVEEIKIIHQIRPVL------PDPDAVAVVEIDRQQYHYLHGYAYEDDEE 95
E D +Q+ E+ + +I P + PDP+ + + + + G EDDEE
Sbjct: 31 ELDLVEQEDQITHEVSLDEEIDPEISLDIFKPDPNFLENEKRYEELKKSMLGEESEDDEE 210
Query: 96 QVVDDEEDDDDRCDLDDE 113
+ + +DDDD D DE
Sbjct: 211 GLDSESDDDDDEDDESDE 264
>TC215201 similar to UP|Q70Z19 (Q70Z19) Nucleosome assembly protein 1-like
protein 1, partial (55%)
Length = 904
Score = 32.0 bits (71), Expect = 0.14
Identities = 21/58 (36%), Positives = 31/58 (53%), Gaps = 1/58 (1%)
Frame = +3
Query: 90 YEDDEEQVVDDEE-DDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKNSKRSP 146
+ED +E DDEE D+D+ D D+E E K G+ R K G+ K + ++R P
Sbjct: 495 FEDIDEVDYDDEEGDEDEDEDEDEEEEEEEEEEKKGKSRSKSGGRVKPGKDQPTERPP 668
>BM308202
Length = 433
Score = 31.6 bits (70), Expect = 0.19
Identities = 27/86 (31%), Positives = 34/86 (39%), Gaps = 18/86 (20%)
Frame = +2
Query: 49 DAAPVEEIKIIHQIRPVL-----PDPDAVAVVEIDRQQYHYLHGYAYE------------ 91
DA +I I R VL P V ++ +D Q Y YE
Sbjct: 74 DAINYSDIATIPVDRCVLDFAAEPTDSFVGLITMDDQDEMYASARIYEIGRRRPTDDDSD 253
Query: 92 -DDEEQVVDDEEDDDDRCDLDDELVP 116
DD E +DE+DDDD D+D L P
Sbjct: 254 PDDAESEEEDEDDDDDDPDVDPLLGP 331
>TC204849 similar to GB|AAM16178.1|20334844|AY094022 AT3g12390/T2E22_130
{Arabidopsis thaliana;} , partial (96%)
Length = 883
Score = 31.6 bits (70), Expect = 0.19
Identities = 18/49 (36%), Positives = 26/49 (52%)
Frame = +2
Query: 91 EDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKM 139
EDD+ D+EEDDDD D DD+ G+ GR + + K++ M
Sbjct: 143 EDDD----DEEEDDDDDDDEDDDHAEGLEGDASGRSKQTRSEKKSRKAM 277
>TC203635 similar to PIR|T46225|T46225 alpha NAC-like protein - Arabidopsis
thaliana {Arabidopsis thaliana;} , partial (71%)
Length = 1007
Score = 31.6 bits (70), Expect = 0.19
Identities = 28/108 (25%), Positives = 48/108 (43%)
Frame = +1
Query: 32 VVDAEEEQEQEPDQNQQDAAPVEEIKIIHQIRPVLPDPDAVAVVEIDRQQYHYLHGYAYE 91
VVDA E + E D A ++I P + DA V ++ +
Sbjct: 70 VVDASPEVDAEQQLPSVDDATQKKI-------PQPQEDDAPLVEDL-------------K 189
Query: 92 DDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKM 139
DD++ D+++DDDD D +D+ G+K + R +K ++A K+
Sbjct: 190 DDDKDDDDEDDDDDDEDDKEDDAQGGAEGSK--QSRSEKKSRKAMLKL 327
>TC234741 weakly similar to UP|Q93ZH7 (Q93ZH7) AT5g03040/F15A17_70, partial
(10%)
Length = 423
Score = 31.6 bits (70), Expect = 0.19
Identities = 24/86 (27%), Positives = 41/86 (46%), Gaps = 1/86 (1%)
Frame = -2
Query: 22 DSLLQWELVDVVDAEEEQEQEPDQNQQDAAPVEEIKI-IHQIRPVLPDPDAVAVVEIDRQ 80
DSL W + D + Q+Q+P + QQ P + + +H P L D + +VE+ Q
Sbjct: 293 DSLFHWNEACL*DKQPLQQQQPRRPQQ*PKPKKRLACSVHYSLPHLLVWD-LFLVEVKEQ 117
Query: 81 QYHYLHGYAYEDDEEQVVDDEEDDDD 106
+ L + D+ + ++ DDDD
Sbjct: 116 RQLVLLLDPHSDESQNILTLTYDDDD 39
>TC203654 similar to PIR|T46225|T46225 alpha NAC-like protein - Arabidopsis
thaliana {Arabidopsis thaliana;} , partial (76%)
Length = 1139
Score = 31.2 bits (69), Expect = 0.24
Identities = 15/49 (30%), Positives = 29/49 (58%)
Frame = +3
Query: 91 EDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKM 139
+DD+++ +DE++DDD D +D+ G K + R +K ++A K+
Sbjct: 219 KDDDKEETEDEDEDDDDDDKEDDAQGGAEGGK--QSRSEKKSRKAMLKL 359
>TC215452 similar to UP|P93488 (P93488) NAP1Ps, partial (29%)
Length = 609
Score = 30.8 bits (68), Expect = 0.32
Identities = 21/56 (37%), Positives = 26/56 (45%)
Frame = +3
Query: 91 EDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKNSKRSP 146
ED EE +DEEDDDD D DD+ E K KK G+ + +R P
Sbjct: 156 EDIEEDDDEDEEDDDDDDDDDDD--EEESKTKKKSSAPKKSGRAQLGDGQQGERPP 317
Score = 26.6 bits (57), Expect = 6.0
Identities = 11/23 (47%), Positives = 16/23 (68%)
Frame = +3
Query: 91 EDDEEQVVDDEEDDDDRCDLDDE 113
EDDE+ +E+DD+D D DD+
Sbjct: 138 EDDEDDEDIEEDDDEDEEDDDDD 206
>TC215449 similar to UP|P93488 (P93488) NAP1Ps, partial (97%)
Length = 1495
Score = 30.8 bits (68), Expect = 0.32
Identities = 20/57 (35%), Positives = 28/57 (49%), Gaps = 1/57 (1%)
Frame = +2
Query: 91 EDDEE-QVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKNSKRSP 146
EDDE+ + DDEE+++D D DDE E K KK G+ + +R P
Sbjct: 1052 EDDEDIEEDDDEEEEEDEDDDDDEDDEEESKTKKKSSAPKKSGRAQLGDGQQGERPP 1222
>TC235094 similar to UP|PRT3_SCYCA (P30258) Protamine Z3 (Scylliorhinine Z3),
partial (76%)
Length = 466
Score = 30.4 bits (67), Expect = 0.41
Identities = 19/55 (34%), Positives = 32/55 (57%), Gaps = 4/55 (7%)
Frame = -2
Query: 89 AYEDDEEQVVDDEEDDDDRCDLDDELVPWEVG--NKLGRQRMK--KLGKRACSKM 139
A EDDE +DE+++++ D DD++V E ++L QR K L +R S++
Sbjct: 357 AEEDDEIDEPEDEDEEEEEDDDDDDVVSQEQSPLSRLREQRSKLETLSRRLASEL 193
>TC215450 similar to UP|P93488 (P93488) NAP1Ps, partial (30%)
Length = 672
Score = 30.4 bits (67), Expect = 0.41
Identities = 18/56 (32%), Positives = 28/56 (49%)
Frame = +3
Query: 91 EDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRACSKMKNSKRSP 146
EDD+E+ +DE+DDDD DDE + + + KK G+ + +R P
Sbjct: 240 EDDDEEEEEDEDDDDDE---DDE--------EESKTKKKKSGRAQLGDGQQGERPP 374
>TC204851 similar to GB|AAM16178.1|20334844|AY094022 AT3g12390/T2E22_130
{Arabidopsis thaliana;} , partial (86%)
Length = 868
Score = 30.0 bits (66), Expect = 0.54
Identities = 16/63 (25%), Positives = 34/63 (53%)
Frame = +2
Query: 77 IDRQQYHYLHGYAYEDDEEQVVDDEEDDDDRCDLDDELVPWEVGNKLGRQRMKKLGKRAC 136
+++Q+ H A +D++E+ DD+++DD+ DD+ G+ GR + + K++
Sbjct: 113 LEQQKIHDDEPVAEDDNDEEEDDDDDEDDE----DDDHADGLEGDASGRSKQTRSEKKSR 280
Query: 137 SKM 139
M
Sbjct: 281 KAM 289
>TC217978 weakly similar to UP|Q8VYJ8 (Q8VYJ8) AT4g27310/M4I22_120, partial
(42%)
Length = 1064
Score = 30.0 bits (66), Expect = 0.54
Identities = 14/31 (45%), Positives = 20/31 (64%), Gaps = 2/31 (6%)
Frame = +1
Query: 89 AYEDDEEQVV--DDEEDDDDRCDLDDELVPW 117
A E+ EE DDE D +D D+D+++VPW
Sbjct: 463 AAEETEESEAGNDDEPDTEDDDDVDNQVVPW 555
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.137 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,341,492
Number of Sequences: 63676
Number of extensions: 95653
Number of successful extensions: 1221
Number of sequences better than 10.0: 102
Number of HSP's better than 10.0 without gapping: 640
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 908
length of query: 167
length of database: 12,639,632
effective HSP length: 90
effective length of query: 77
effective length of database: 6,908,792
effective search space: 531976984
effective search space used: 531976984
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0258a.2


