

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0253.9
(677 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
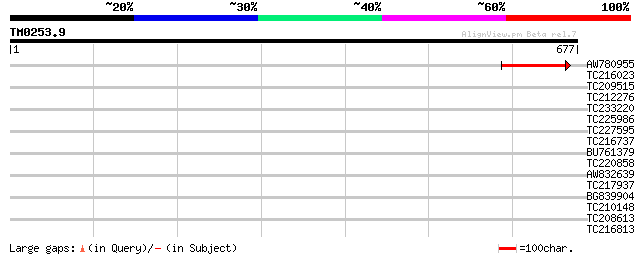
Score E
Sequences producing significant alignments: (bits) Value
AW780955 120 2e-27
TC216023 similar to GB|AAM78036.1|21928015|AY125526 At2g01100/F2... 35 0.083
TC209515 similar to UP|Q8VWH9 (Q8VWH9) Stress responsive gene 6,... 35 0.083
TC212276 similar to UP|Q94A31 (Q94A31) AT5g02520/T22P11_110, par... 35 0.14
TC233220 similar to UP|Q9M9T8 (Q9M9T8) F6A14.22 protein, partial... 34 0.24
TC225986 weakly similar to UP|Q9W1I6 (Q9W1I6) CG18426-PA (LD1203... 33 0.54
TC227595 homologue to UP|Q43451 (Q43451) G-box binding factor (F... 32 0.92
TC216737 homologue to UP|Q8GZD2 (Q8GZD2) Expansin, partial (97%) 31 1.6
BU761379 similar to PIR|T18277|T18 kinesin heavy chain - slime m... 31 2.0
TC220858 31 2.0
AW832639 similar to GP|22135832|gb At1g34430/F7P12_2 {Arabidopsi... 30 2.7
TC217937 similar to GB|AAP68233.1|31711754|BT008794 At4g15680 {A... 30 2.7
BG839904 30 4.6
TC210148 29 6.0
TC208613 similar to GB|AAP31934.1|30387537|BT006590 At2g35860 {A... 29 7.8
TC216813 similar to UP|WR11_ARATH (Q9SV15) Probable WRKY transcr... 29 7.8
>AW780955
Length = 295
Score = 120 bits (302), Expect = 2e-27
Identities = 59/83 (71%), Positives = 70/83 (84%), Gaps = 1/83 (1%)
Frame = +3
Query: 588 TNEGSEQCLQEERDGSRAPVE-QKRVKSRSQKDTKSKRLAKRQSLAAAGTSWTSGIRRST 646
+++ E CLQE+ DGS PV QKRVKS QK +K+K+L+ R SLAAAGTSWTSG+RRST
Sbjct: 9 SDDDPEHCLQEKTDGSMVPVNGQKRVKSLVQKVSKNKKLSNRHSLAAAGTSWTSGLRRST 188
Query: 647 RIRTRPLEYWKGERLVYGRIHQS 669
RIRTRPLEYWKGERLVYGR+H+S
Sbjct: 189 RIRTRPLEYWKGERLVYGRVHKS 257
>TC216023 similar to GB|AAM78036.1|21928015|AY125526 At2g01100/F23H14.7
{Arabidopsis thaliana;} , partial (15%)
Length = 1437
Score = 35.4 bits (80), Expect = 0.083
Identities = 46/195 (23%), Positives = 71/195 (35%), Gaps = 14/195 (7%)
Frame = +3
Query: 51 LRSMALRSPAKLVEQTKAILEENPGLFNSQDDERNDVVDVDVDVAEDGQDFPRKRRPGLG 110
LR RS + + T L + Q + D D +G +KR G+
Sbjct: 576 LRDRGSRSDSSNYDSTADELSKRKRQHKRQRPSKFSGSDYSTD---EGDSLVQKRNHGMH 746
Query: 111 LKRARFSLKPSTSVSLESLLPNLDIENLKDPVEFFKAHERMENARREI---------QKQ 161
KR R S S ES L + D + K + K H+R N + Q+
Sbjct: 747 SKRHR------QSESSESDLSSDDAPHKKGHRKHHKHHKRSHNVELRLSDSDYRSHGQRS 908
Query: 162 MGVSVEQNQDDVS-TRPRQRRPGL----RGNDQRSIKYRHRYSTEISDKNDHAMASQEAF 216
S+E+N +D S R ++ G R + R +RH E + H+
Sbjct: 909 RSKSLEENSEDESKKRSMDKKSGRHHHHRHHHHRHHHHRHHLDDERNYMQQHSPKFNGKH 1088
Query: 217 GSDSLDPDTEKIENG 231
G + +TEK + G
Sbjct: 1089GEELDKTETEKKDGG 1133
>TC209515 similar to UP|Q8VWH9 (Q8VWH9) Stress responsive gene 6, Srg6
(Stress responsive gene 6 protein, Srg6), partial (55%)
Length = 1386
Score = 35.4 bits (80), Expect = 0.083
Identities = 46/214 (21%), Positives = 86/214 (39%), Gaps = 6/214 (2%)
Frame = +2
Query: 447 GKSSNKL-DAPLIEDILAVSESCLIEDAVMNSTSTSQMPMEDNSMEPEFDANVDRNEPHA 505
GK KL D + ED L ++ L ED ++ E+ + EFD++ D++EP
Sbjct: 23 GKRLTKLLDDEIQEDELFWNQDALKEDEEDDNYQ------EEPEIADEFDSDFDQDEPVP 184
Query: 506 DMDVDNGGSGMGETVMDDTVAKPNIEVNEQCQFEGKILT-----ENTDAFTASMPTDDTD 560
+ + D + E + ++ F GK L + T + S P ++ D
Sbjct: 185 EEEEDPNKNDDDE----------RMHKKKRLIFPGKTLAKKKKKKKTISKLESSPKENED 334
Query: 561 IDVVNPLADQSSPGVIQANSIDKRTTCTNEGSEQCLQEERDGSRAPVEQKRVKSRSQKDT 620
++ S V++ ++ + S Q ERD RA ++ + +K+
Sbjct: 335 --------EEHSGKVVEDEGGERMIRKSTRTSVIVRQAERDAIRAALQATIKPVKRKKEG 490
Query: 621 KSKRLAKRQSLAAAGTSWTSGIRRSTRIRTRPLE 654
+ KR+ + + L A + +R R+ R E
Sbjct: 491 EEKRMTQEEMLLEAAQTEIMNLRNLERVLAREEE 592
>TC212276 similar to UP|Q94A31 (Q94A31) AT5g02520/T22P11_110, partial (9%)
Length = 1009
Score = 34.7 bits (78), Expect = 0.14
Identities = 42/213 (19%), Positives = 84/213 (38%), Gaps = 21/213 (9%)
Frame = +2
Query: 477 STSTSQMPMEDNSMEPEFDANVDRNEPHADMDVDNGGSGMGETVMDDTVAKPNIEVNEQC 536
S ++ + P + PE ++NV + ++ +GG + D I+V +Q
Sbjct: 8 SLASEEAPGDHEKSFPENESNVSKEINGVNVACSSGGKSRSARLHD-------IKVYQQK 166
Query: 537 Q-FEGKILTENTDAFTASMPTDDTDI----DVVNPLADQSS------PGVIQANSIDKRT 585
+ G L + + S+ ++ D+ P+ QSS PG + S K +
Sbjct: 167 KPASGGSLKHPNNENSTSVALENCDVKGLKSPATPIQSQSSRQLSTSPGQVIKKSASKIS 346
Query: 586 TCTNEGSEQCLQEERDGSRAPVEQKR---------VKSRSQKDTKSKRLAKRQSLAA-AG 635
+ +E C +++R V + +K+ +KD +Q ++
Sbjct: 347 RTLSPKTEGCYKKKRVTVETKVVMPKGKLNKSASALKNPREKDLSPLAKGSQQKISTFTP 526
Query: 636 TSWTSGIRRSTRIRTRPLEYWKGERLVYGRIHQ 668
S + RS R+ PLE+W+ + +Y H+
Sbjct: 527 ESLSFRKSRSGRLLLPPLEFWRNQIPIYNADHE 625
>TC233220 similar to UP|Q9M9T8 (Q9M9T8) F6A14.22 protein, partial (19%)
Length = 566
Score = 33.9 bits (76), Expect = 0.24
Identities = 23/80 (28%), Positives = 36/80 (44%)
Frame = +3
Query: 270 GDGAMDLLQERLQIKPVVLEKVSVPDFPDNQPIDLKSLRGSLSKPRKALSNIDNLLNRRK 329
G L E + KP + S+P +P ++ ID K S +K + R+
Sbjct: 120 GTATSALSSEYFKTKPYACDPSSLPVYPPSKEIDAKHREESR---KKISGRVRGTETRKP 290
Query: 330 NKTPLRQDAGSPAEQLASPT 349
++ PL + +PAE LAS T
Sbjct: 291 SRKPLGFNKLAPAEDLASQT 350
>TC225986 weakly similar to UP|Q9W1I6 (Q9W1I6) CG18426-PA (LD12035p), partial
(13%)
Length = 1553
Score = 32.7 bits (73), Expect = 0.54
Identities = 30/134 (22%), Positives = 57/134 (42%), Gaps = 7/134 (5%)
Frame = +2
Query: 146 KAHERMENARREIQKQMGVSVEQNQDDVSTRPRQRRPGLRGNDQRSIKY-------RHRY 198
+ +E+ + RE +++ + ++ +D +R R RR + ++ ++ R +
Sbjct: 203 RQYEKEKEKERERKRRKEILYDEEDEDEDSRKRWRRSVIEEKRKKRLREKEDDLVDRQKE 382
Query: 199 STEISDKNDHAMASQEAFGSDSLDPDTEKIENGEACLTSLENEVTDSSATEGNKICEILD 258
EI++ A Q+ D+L +E + NG EN +TD E I D
Sbjct: 383 EEEIAEAKKKAEEEQKR-QRDALKLLSEHVVNG-----GKENMITDDITNEVKSIVVEQD 544
Query: 259 GLLRCNSEDLEGDG 272
+ + ED GDG
Sbjct: 545 TVADYSREDHIGDG 586
>TC227595 homologue to UP|Q43451 (Q43451) G-box binding factor (Fragment),
complete
Length = 1453
Score = 32.0 bits (71), Expect = 0.92
Identities = 28/93 (30%), Positives = 43/93 (46%), Gaps = 5/93 (5%)
Frame = +3
Query: 559 TDIDVVNPLADQSSPGVIQANSIDKRTTCTNEGSEQCLQEERDGSRAPVEQKR----VKS 614
T +++ NP S +ANS C +E CLQ ER+ R +Q +S
Sbjct: 555 TTLELRNPSTVHS-----KANSTSAPQPCAIVPNETCLQNERELKRERRKQSNRESARRS 719
Query: 615 RSQKDTKSKRLAKR-QSLAAAGTSWTSGIRRST 646
R +K +++ LA++ + L A S S I R T
Sbjct: 720 RLRKQAETEELARKVEMLTAENVSLKSEITRLT 818
>TC216737 homologue to UP|Q8GZD2 (Q8GZD2) Expansin, partial (97%)
Length = 1296
Score = 31.2 bits (69), Expect = 1.6
Identities = 15/41 (36%), Positives = 21/41 (50%)
Frame = -1
Query: 573 PGVIQANSIDKRTTCTNEGSEQCLQEERDGSRAPVEQKRVK 613
PG NS D T C + + C +E +GS E++RVK
Sbjct: 141 PGATGVNSRDSCTCCNKKRDQ*CNVDENNGSHCKRERERVK 19
>BU761379 similar to PIR|T18277|T18 kinesin heavy chain - slime mold
(Dictyostelium discoideum), partial (1%)
Length = 449
Score = 30.8 bits (68), Expect = 2.0
Identities = 24/84 (28%), Positives = 35/84 (41%), Gaps = 4/84 (4%)
Frame = +2
Query: 310 SLSKPRKALSNIDNLLNRRKNKTPLRQDAGSPAEQLASPTPPRSPFAP----LSSLLNHI 365
SLS PRK +N N + TP P L P PP + ++P +SS
Sbjct: 41 SLSSPRKIQTNDAPTANCHVSVTPTTLSIVQPHSLLPKPVPPTTTYSPNDVVVSSSATFA 220
Query: 366 SRLKPSVDPFSAHDIDHLSTTKYS 389
L+ + S+ +ST K+S
Sbjct: 221 KFLRQRRNDLSSAISKGISTLKHS 292
>TC220858
Length = 881
Score = 30.8 bits (68), Expect = 2.0
Identities = 25/102 (24%), Positives = 43/102 (41%), Gaps = 5/102 (4%)
Frame = +3
Query: 497 NVDRNEPHADMDVDNGGSGMGETVMDDTVAKPNIEVNEQCQFEGKILT----ENTDAFTA 552
N RNE ++ D+ G T +T PN++ E F+G T E +D+ +A
Sbjct: 330 NAYRNE-RSESDISESDKSNGNT---ETNENPNVDATEDEMFKGDTQTGETDETSDSSSA 497
Query: 553 SMPTDDTDIDVVNPLADQSSPGVIQANS-IDKRTTCTNEGSE 593
+ D + D ++ V +A + +D NEG +
Sbjct: 498 NETMDSAEQDAIDSSDTHIHEDVAEARTDLDTLPDIRNEGDD 623
>AW832639 similar to GP|22135832|gb At1g34430/F7P12_2 {Arabidopsis
thaliana}, partial (21%)
Length = 307
Score = 30.4 bits (67), Expect = 2.7
Identities = 20/61 (32%), Positives = 34/61 (54%), Gaps = 4/61 (6%)
Frame = +2
Query: 42 DDLDAIHNHLRSMALRSPAKLVEQTKAILEENPGLFN-SQDDER---NDVVDVDVDVAED 97
D LDA++ ++S + L + T L ++PG+ + S+D R N V+++ V VA D
Sbjct: 38 DALDALYKKIKSKGVTMTTLLAKATVLALVKHPGMNSVSRDGNRSTYNIVINIAVGVAID 217
Query: 98 G 98
G
Sbjct: 218G 220
>TC217937 similar to GB|AAP68233.1|31711754|BT008794 At4g15680 {Arabidopsis
thaliana;} , partial (14%)
Length = 906
Score = 30.4 bits (67), Expect = 2.7
Identities = 34/138 (24%), Positives = 60/138 (42%), Gaps = 3/138 (2%)
Frame = +1
Query: 327 RRKNKTPLRQDAGSPAEQLASPTPPRSPFAPLSSLLNHISRLKPSVDPFSAHDIDHLSTT 386
R+ N+ P+R A A+ ++ + A SS S L+P A+D+DH+S++
Sbjct: 229 RKMNQRPVRISAWKLAKLDSNEATKAAAKARASS-----SVLRPISSRPHAYDVDHVSSS 393
Query: 387 KYSPVHKMNQELNVVGSAKPSNELNDHITKDAVAIGETNAVPDTLRNCASASENSKED-- 444
NV G + P + HI D V G + P ++ S+ SKED
Sbjct: 394 ------------NVSGRSSPISNQGFHIKNDTV--GTSRLSPS--KSSYPPSQASKEDID 525
Query: 445 -NSGKSSNKLDAPLIEDI 461
+ S + + +P + ++
Sbjct: 526 ASCQHSMSNISSPQVSNL 579
>BG839904
Length = 712
Score = 29.6 bits (65), Expect = 4.6
Identities = 23/78 (29%), Positives = 35/78 (44%), Gaps = 6/78 (7%)
Frame = -3
Query: 134 DIENLKDPVEFFKAHERMENARREIQKQMGVSVEQNQDDVST------RPRQRRPGLRGN 187
D + + E F + E M N R++ K+ G + + + ST R QR+ +
Sbjct: 518 DCSDAFNTTEVFPSREAMLNWARDVAKENGFVLIILRSETSTTCTRSTRCNQRKTFVIMG 339
Query: 188 DQRSIKYRHRYSTEISDK 205
RS KYR Y E+S K
Sbjct: 338 CDRSGKYRGPYKNELSRK 285
>TC210148
Length = 950
Score = 29.3 bits (64), Expect = 6.0
Identities = 11/19 (57%), Positives = 16/19 (83%)
Frame = +3
Query: 332 TPLRQDAGSPAEQLASPTP 350
TP R +A +PAEQ+++PTP
Sbjct: 603 TPGRDEAATPAEQISTPTP 659
>TC208613 similar to GB|AAP31934.1|30387537|BT006590 At2g35860 {Arabidopsis
thaliana;} , partial (62%)
Length = 1101
Score = 28.9 bits (63), Expect = 7.8
Identities = 22/75 (29%), Positives = 33/75 (43%), Gaps = 6/75 (8%)
Frame = -3
Query: 299 NQPIDLKSLRGSLSKPRKALSNIDNLLNRRKNK--TPLRQD----AGSPAEQLASPTPPR 352
+ P+ LR L++ + + LL RRK PL +D EQ++ P P
Sbjct: 880 SSPVLDNKLRSVLTRLGNGSNPLRILLRRRKQNPINPLNRDPPVCVNIRVEQISRPVPVP 701
Query: 353 SPFAPLSSLLNHISR 367
P+S L NH+ R
Sbjct: 700 EFNGPISFLSNHLMR 656
>TC216813 similar to UP|WR11_ARATH (Q9SV15) Probable WRKY transcription
factor 11 (WRKY DNA-binding protein 11), partial (46%)
Length = 1173
Score = 28.9 bits (63), Expect = 7.8
Identities = 22/101 (21%), Positives = 42/101 (40%)
Frame = +3
Query: 315 RKALSNIDNLLNRRKNKTPLRQDAGSPAEQLASPTPPRSPFAPLSSLLNHISRLKPSVDP 374
R+ L +R N + + P L +PTPP P S + S P++
Sbjct: 147 RRQLKKQPPKASRAWNTSSVSSPINLPTSALTTPTPP----FPTSKNSSPFSTAAPAMPV 314
Query: 375 FSAHDIDHLSTTKYSPVHKMNQELNVVGSAKPSNELNDHIT 415
+AH L T P+H++ + + + S + ++ ++T
Sbjct: 315 SAAHPSLPLPTLSPHPLHRLTRSPSPLRSHRV*PSISQNLT 437
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.309 0.128 0.354
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,095,851
Number of Sequences: 63676
Number of extensions: 297155
Number of successful extensions: 1661
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 1650
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1659
length of query: 677
length of database: 12,639,632
effective HSP length: 104
effective length of query: 573
effective length of database: 6,017,328
effective search space: 3447928944
effective search space used: 3447928944
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0253.9


