

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0252a.5
(150 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
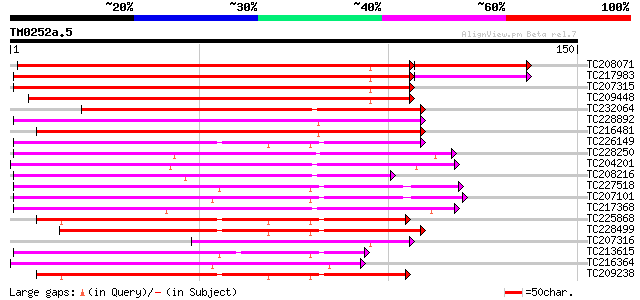
Score E
Sequences producing significant alignments: (bits) Value
TC208071 UP|Q8GRZ5 (Q8GRZ5) Phosphoenolpyruvate carboxylase kina... 161 5e-42
TC217983 UP|Q8GSL0 (Q8GSL0) Phosphoenolpyruvate carboxylase kina... 165 6e-42
TC207315 UP|Q8H1P2 (Q8H1P2) Phosphoenolpyruvate carboxylase kina... 147 2e-36
TC209448 UP|Q6ULS3 (Q6ULS3) Phosphoenolpyruvate carboxylase kina... 134 1e-32
TC232064 similar to GB|AAP31952.1|30387573|BT006608 At1g12680 {A... 98 2e-21
TC228892 UP|Q9XF25 (Q9XF25) SNF-1-like serine/threonine protein ... 87 2e-18
TC216481 homologue to UP|P93113 (P93113) SNF1-related protein ki... 85 1e-17
TC226149 similar to UP|Q8H2C2 (Q8H2C2) Serine/threonine kinase, ... 85 1e-17
TC228250 UP|Q84P29 (Q84P29) Seed calcium dependent protein kinas... 84 3e-17
TC204201 similar to UP|Q8W559 (Q8W559) Calcium/calmodulin-depend... 82 1e-16
TC208216 similar to UP|O48565 (O48565) Calcium-dependent protein... 82 1e-16
TC227518 similar to UP|Q93VD3 (Q93VD3) At1g30270/F12P21_6 (CBL-i... 81 2e-16
TC207101 similar to UP|O24342 (O24342) Serine/threonine kinase ,... 81 2e-16
TC217368 similar to UP|Q93YF4 (Q93YF4) Calcium-dependent protein... 80 4e-16
TC225868 UP|Q8LK24 (Q8LK24) SOS2-like protein kinase, complete 80 4e-16
TC228499 homologue to UP|Q8LK24 (Q8LK24) SOS2-like protein kinas... 80 4e-16
TC207316 homologue to UP|Q8H1P2 (Q8H1P2) Phosphoenolpyruvate car... 80 4e-16
TC213615 similar to UP|Q9SUL7 (Q9SUL7) SNF1 like protein kinase ... 79 1e-15
TC216364 similar to UP|Q9MAI7 (Q9MAI7) F12M16.4, partial (58%) 78 1e-15
TC209238 similar to UP|Q8LK24 (Q8LK24) SOS2-like protein kinase,... 78 2e-15
>TC208071 UP|Q8GRZ5 (Q8GRZ5) Phosphoenolpyruvate carboxylase kinase, complete
Length = 1188
Score = 161 bits (407), Expect(2) = 5e-42
Identities = 80/124 (64%), Positives = 89/124 (71%), Gaps = 19/124 (15%)
Frame = +3
Query: 3 PIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRR 62
P E + ASL+K LLEA+ HCHRLGVAH D+KPDN+LF + +LKLADFGSAEWFGDGR
Sbjct: 318 PFSESQAASLIKNLLEAVAHCHRLGVAHRDIKPDNILFDSADNLKLADFGSAEWFGDGRS 497
Query: 63 MSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIWGN 103
MSGVVGTPYYVAPEVL+G E EKVDVWSCGV IFEAV+ N
Sbjct: 498 MSGVVGTPYYVAPEVLLGREYDEKVDVWSCGVILYIMLAGIPPFYGDSAAEIFEAVVKAN 677
Query: 104 LRFP 107
LRFP
Sbjct: 678 LRFP 689
Score = 26.2 bits (56), Expect(2) = 5e-42
Identities = 18/31 (58%), Positives = 20/31 (64%)
Frame = +1
Query: 108 LESSGMFLLLLRIS*GR*FAGTLLTESLQNK 138
LESSG+ RIS* +* GTLL S QNK
Sbjct: 691 LESSGLCPPPPRIS*EK*SLGTLLEGSPQNK 783
>TC217983 UP|Q8GSL0 (Q8GSL0) Phosphoenolpyruvate carboxylase kinase, complete
Length = 1203
Score = 165 bits (418), Expect(2) = 6e-42
Identities = 82/125 (65%), Positives = 91/125 (72%), Gaps = 19/125 (15%)
Frame = +3
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
GPI E + A+LMK LLEA+ HCHRLGVAH D+KPDN+LF + +LKLADFGSAEWFGDGR
Sbjct: 339 GPIQESQAAALMKNLLEAVAHCHRLGVAHRDIKPDNILFDSADNLKLADFGSAEWFGDGR 518
Query: 62 RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIWG 102
MSGVVGTPYYVAPEVL+G E EKVDVWSCGV IFEAV+
Sbjct: 519 SMSGVVGTPYYVAPEVLLGREYDEKVDVWSCGVILYIMLAGIPPFYGDSAAEIFEAVVRA 698
Query: 103 NLRFP 107
NLRFP
Sbjct: 699 NLRFP 713
Score = 21.6 bits (44), Expect(2) = 6e-42
Identities = 16/31 (51%), Positives = 17/31 (54%)
Frame = +1
Query: 108 LESSGMFLLLLRIS*GR*FAGTLLTESLQNK 138
LESSG RIS +* G LL S QNK
Sbjct: 715 LESSGQCPPPPRISCEK*SVGILLEGSPQNK 807
>TC207315 UP|Q8H1P2 (Q8H1P2) Phosphoenolpyruvate carboxylase kinase, complete
Length = 1063
Score = 147 bits (371), Expect = 2e-36
Identities = 73/125 (58%), Positives = 85/125 (67%), Gaps = 19/125 (15%)
Frame = +3
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
GP+ E ASL+KQLLEA+ HCH G+AH D+KP+N+LF G LKL+DFGSAEW G+G
Sbjct: 336 GPLTEPHAASLLKQLLEAVAHCHAQGLAHRDIKPENILFDEGNKLKLSDFGSAEWLGEGS 515
Query: 62 RMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIWG 102
MSGVVGTPYYVAPEV+MG E EKVDVWS GV IFE+V+
Sbjct: 516 SMSGVVGTPYYVAPEVIMGREYDEKVDVWSSGVILYAMLAGFPPFYGESAPEIFESVLRA 695
Query: 103 NLRFP 107
NLRFP
Sbjct: 696 NLRFP 710
>TC209448 UP|Q6ULS3 (Q6ULS3) Phosphoenolpyruvate carboxylase kinase 4,
complete
Length = 1185
Score = 134 bits (338), Expect = 1e-32
Identities = 71/121 (58%), Positives = 81/121 (66%), Gaps = 19/121 (15%)
Frame = +3
Query: 6 EVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRRMSG 65
E E AS+M QL++A+ HCHRLGVAH DVKPDN+LF LKLADFGSA+ F +G MSG
Sbjct: 327 EPEAASVMWQLMQAVAHCHRLGVAHRDVKPDNILFDEENRLKLADFGSADTFKEGEPMSG 506
Query: 66 VVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------------IFEAVIWGNLRF 106
VVGTP+YVAPEVL G + EKVDVWS GV IFEAV+ NLRF
Sbjct: 507 VVGTPHYVAPEVLAGRDYNEKVDVWSAGVVLYQMLAGFLPFRGDSPVEIFEAVLRANLRF 686
Query: 107 P 107
P
Sbjct: 687 P 689
>TC232064 similar to GB|AAP31952.1|30387573|BT006608 At1g12680 {Arabidopsis
thaliana;} , partial (34%)
Length = 770
Score = 97.8 bits (242), Expect = 2e-21
Identities = 45/91 (49%), Positives = 63/91 (68%)
Frame = +2
Query: 20 ITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRRMSGVVGTPYYVAPEVLM 79
+ +CH +GV H D+KP+N+L A G +KLADFG A +G+ ++GV G+P YVAPEVL+
Sbjct: 5 VKYCHDMGVVHRDIKPENILLTAAGKIKLADFGLAIRISEGQNLTGVAGSPAYVAPEVLL 184
Query: 80 GWENGEKVDVWSCGVIFEAVIWGNLRFPLES 110
G EKVD+WS GV+ A++ G L F +S
Sbjct: 185 G-RYSEKVDIWSSGVLLHALLVGGLPFKGDS 274
>TC228892 UP|Q9XF25 (Q9XF25) SNF-1-like serine/threonine protein kinase,
complete
Length = 1931
Score = 87.4 bits (215), Expect = 2e-18
Identities = 44/110 (40%), Positives = 65/110 (59%), Gaps = 1/110 (0%)
Frame = +1
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + E E +Q++ + +CHR V H D+KP+N+L + ++K+ADFG + DG
Sbjct: 415 GRLQEDEARHFFQQIISGVEYCHRNMVVHRDLKPENLLLDSKFNIKIADFGLSNIMRDGH 594
Query: 62 RMSGVVGTPYYVAPEVLMG-WENGEKVDVWSCGVIFEAVIWGNLRFPLES 110
+ G+P Y APEV+ G G +VDVWSCGVI A++ G L F E+
Sbjct: 595 FLKTSCGSPNYAAPEVISGKLYAGPEVDVWSCGVILYALLCGTLPFDDEN 744
>TC216481 homologue to UP|P93113 (P93113) SNF1-related protein kinase ,
partial (77%)
Length = 1596
Score = 85.1 bits (209), Expect = 1e-17
Identities = 42/104 (40%), Positives = 63/104 (60%), Gaps = 1/104 (0%)
Frame = +3
Query: 8 EVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRRMSGVV 67
E + +Q++ + +CHR V H D+KP+N+L + ++K+ADFG + DG +
Sbjct: 6 EARNFFQQIISGVEYCHRNMVVHRDLKPENLLLDSKCNVKIADFGLSNIMRDGHFLKTSC 185
Query: 68 GTPYYVAPEVLMG-WENGEKVDVWSCGVIFEAVIWGNLRFPLES 110
G+P Y APEV+ G G +VDVWSCGVI A++ G L F E+
Sbjct: 186 GSPNYAAPEVISGKLYAGPEVDVWSCGVILYALLCGTLPFDDEN 317
>TC226149 similar to UP|Q8H2C2 (Q8H2C2) Serine/threonine kinase, partial
(49%)
Length = 676
Score = 85.1 bits (209), Expect = 1e-17
Identities = 49/115 (42%), Positives = 68/115 (58%), Gaps = 6/115 (5%)
Frame = +1
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGR 61
G + E +QL+ A+ CH GV H D+KP+N+L DLK++DFG + + R
Sbjct: 268 GKLTEDLARKYFQQLISAVDFCHSRGVTHRDLKPENLLLDQNEDLKVSDFGLST-LPEQR 444
Query: 62 RMSGVV----GTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRFPLES 110
R G++ GTP YVAPEVL G++ G K D+WSCGVI A++ G L F E+
Sbjct: 445 RADGMLVTPCGTPAYVAPEVLKKKGYD-GSKADIWSCGVILFALLCGYLPFQGEN 606
>TC228250 UP|Q84P29 (Q84P29) Seed calcium dependent protein kinase a,
complete
Length = 1843
Score = 83.6 bits (205), Expect = 3e-17
Identities = 51/121 (42%), Positives = 67/121 (55%), Gaps = 4/121 (3%)
Frame = +3
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGA---GGDLKLADFGSAEWFG 58
G E + A L+K ++E + CH LGV H D+KP+N LF LK DFG + ++
Sbjct: 504 GHYSERQAARLIKTIVEVVEACHSLGVMHRDLKPENFLFDTIDEDAKLKATDFGLSVFYK 683
Query: 59 DGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWGNLRFPLESS-GMFLLL 117
G VVG+PYYVAPEVL G + DVWS GVI ++ G F ES G+F +
Sbjct: 684 PGESFCDVVGSPYYVAPEVLRKL-YGPESDVWSAGVILYILLSGVPPFWAESEPGIFRQI 860
Query: 118 L 118
L
Sbjct: 861 L 863
>TC204201 similar to UP|Q8W559 (Q8W559) Calcium/calmodulin-dependent protein
kinase CaMK3, partial (75%)
Length = 1757
Score = 82.0 bits (201), Expect = 1e-16
Identities = 46/123 (37%), Positives = 69/123 (55%), Gaps = 4/123 (3%)
Frame = +3
Query: 1 GGPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFG---AGGDLKLADFGSAEWF 57
GG E + ++M Q+L + CH GV H D+KP+N L+ +LK DFG +++
Sbjct: 288 GGKYSEDDAKAVMVQILNVVAFCHLQGVVHRDLKPENFLYAKKDESSELKAIDFGLSDFV 467
Query: 58 GDGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWGNLRF-PLESSGMFLL 116
R++ +VG+ YYVAPEVL G + DVWS GVI ++ G+ F SG+F
Sbjct: 468 RPDERLNDIVGSAYYVAPEVLHR-SYGTEADVWSIGVIAYILLCGSRPFWARTESGIFRA 644
Query: 117 LLR 119
+L+
Sbjct: 645 VLK 653
>TC208216 similar to UP|O48565 (O48565) Calcium-dependent protein kinase,
partial (42%)
Length = 1316
Score = 81.6 bits (200), Expect = 1e-16
Identities = 42/104 (40%), Positives = 62/104 (59%), Gaps = 3/104 (2%)
Frame = +3
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGD---LKLADFGSAEWFG 58
G E A++ K ++E + CH+ GV H D+KP+N LF + LK DFG + +F
Sbjct: 735 GHYTERAAAAVTKTIVEVVQMCHKQGVMHRDLKPENFLFANKKETAALKAIDFGLSVFFK 914
Query: 59 DGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWG 102
G + + +VG+PYY+APEVL G +VD+WS GVI ++ G
Sbjct: 915 PGEKFNEIVGSPYYMAPEVLKR-NYGPEVDIWSAGVILYILLCG 1043
>TC227518 similar to UP|Q93VD3 (Q93VD3) At1g30270/F12P21_6 (CBL-interacting
protein kinase 23), partial (86%)
Length = 1492
Score = 80.9 bits (198), Expect = 2e-16
Identities = 48/124 (38%), Positives = 71/124 (56%), Gaps = 5/124 (4%)
Frame = +1
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSA---EWFG 58
G + E E +QL+ A+ +CH GV H D+KP+N+L A G LK++DFG + +
Sbjct: 316 GRLKEDEARKYFQQLICAVDYCHSRGVFHRDLKPENLLLDANGVLKVSDFGLSALPQQVR 495
Query: 59 DGRRMSGVVGTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRFPLESSGMFLL 116
+ + GTP YVAPEV+ G++ G K D+WSCGVI ++ G L P E + + L
Sbjct: 496 EDGLLHTTCGTPNYVAPEVINNKGYD-GAKADLWSCGVILFVLMAGYL--PFEETNLSAL 666
Query: 117 LLRI 120
+I
Sbjct: 667 YKKI 678
>TC207101 similar to UP|O24342 (O24342) Serine/threonine kinase , partial
(98%)
Length = 1719
Score = 80.9 bits (198), Expect = 2e-16
Identities = 48/125 (38%), Positives = 70/125 (55%), Gaps = 5/125 (4%)
Frame = +1
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFG---SAEWFG 58
G + E E +QL+ A+ +CH GV H D+KP+N+L G+LK++DFG ++
Sbjct: 499 GRMSENEARRYFQQLINAVDYCHSRGVYHRDLKPENLLLDTYGNLKVSDFGLSALSQQVR 678
Query: 59 DGRRMSGVVGTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRFPLESSGMFLL 116
D + GTP YVAPEVL G++ G D+WSCGVI ++ G L P + + L
Sbjct: 679 DDGLLHTTCGTPNYVAPEVLNDRGYD-GATADLWSCGVILFVLVAGYL--PFDDPNLMNL 849
Query: 117 LLRIS 121
+IS
Sbjct: 850 YKKIS 864
>TC217368 similar to UP|Q93YF4 (Q93YF4) Calcium-dependent protein kinase 2,
partial (66%)
Length = 1349
Score = 80.1 bits (196), Expect = 4e-16
Identities = 51/122 (41%), Positives = 67/122 (54%), Gaps = 4/122 (3%)
Frame = +1
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLF---GAGGDLKLADFGSAEWFG 58
G E + A L K ++ + CH LGV H D+KP+N LF LK DFG + +F
Sbjct: 52 GHYTERQAAELTKTIVGVVEACHSLGVMHRDLKPENFLFINQHEDSLLKTIDFGLSVFFK 231
Query: 59 DGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGVIFEAVIWGNLRFPLES-SGMFLLL 117
G + VVG+PYYVAPEVL G + DVWS GVI ++ G F E+ G+F +
Sbjct: 232 PGDIFNDVVGSPYYVAPEVLRK-RYGPEADVWSAGVILYILLSGVPPFWAENEQGIFEQV 408
Query: 118 LR 119
LR
Sbjct: 409 LR 414
>TC225868 UP|Q8LK24 (Q8LK24) SOS2-like protein kinase, complete
Length = 2261
Score = 80.1 bits (196), Expect = 4e-16
Identities = 47/106 (44%), Positives = 66/106 (61%), Gaps = 7/106 (6%)
Frame = +2
Query: 8 EVASL-MKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRRMSGV 66
E+A L +QL+ A+ CH GV H D+KP+N+L G+LK+ DFG + F + R G+
Sbjct: 779 EMARLYFQQLISAVDFCHSRGVYHRDLKPENLLLDDDGNLKVTDFGLST-FSEHLRHDGL 955
Query: 67 V----GTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRF 106
+ GTP YVAPEV+ G++ G K D+WSCGVI ++ G L F
Sbjct: 956 LHTTCGTPAYVAPEVIGKRGYD-GAKADIWSCGVILYVLLAGFLPF 1090
>TC228499 homologue to UP|Q8LK24 (Q8LK24) SOS2-like protein kinase, partial
(72%)
Length = 1411
Score = 80.1 bits (196), Expect = 4e-16
Identities = 45/103 (43%), Positives = 64/103 (61%), Gaps = 6/103 (5%)
Frame = +1
Query: 14 KQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRRMSGVV----GT 69
+QL+ A+ CH GV H D+KP+N+L G+LK+ DFG + F + R G++ GT
Sbjct: 1 QQLISAVDFCHSRGVFHRDLKPENLLLDDDGNLKVTDFGLST-FSEHLRHDGLLHTTCGT 177
Query: 70 PYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRFPLES 110
P YVAPEV+ G++ G K D+WSCGVI ++ G L F E+
Sbjct: 178 PAYVAPEVIGKRGYD-GAKADIWSCGVILYVLLAGFLPFQDEN 303
>TC207316 homologue to UP|Q8H1P2 (Q8H1P2) Phosphoenolpyruvate carboxylase
kinase, partial (46%)
Length = 603
Score = 80.1 bits (196), Expect = 4e-16
Identities = 42/78 (53%), Positives = 47/78 (59%), Gaps = 19/78 (24%)
Frame = +2
Query: 49 ADFGSAEWFGDGRRMSGVVGTPYYVAPEVLMGWENGEKVDVWSCGV-------------- 94
+DF EW G+G MSGVVGTPYYVAPEV+MG E EKVDVWS GV
Sbjct: 5 SDFXXXEWLGEGSSMSGVVGTPYYVAPEVIMGREYDEKVDVWSSGVILYAMLAGFPPFYG 184
Query: 95 -----IFEAVIWGNLRFP 107
IFE+V+ NLRFP
Sbjct: 185 ESAPEIFESVLRANLRFP 238
>TC213615 similar to UP|Q9SUL7 (Q9SUL7) SNF1 like protein kinase
(CBL-interacting protein kinase 4), partial (45%)
Length = 674
Score = 78.6 bits (192), Expect = 1e-15
Identities = 44/99 (44%), Positives = 59/99 (59%), Gaps = 5/99 (5%)
Frame = +2
Query: 2 GPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSA---EWFG 58
G +PE QL+ A+ CHR GVAH D+KP N+L A G+LK++DFG + E
Sbjct: 377 GRLPEPLARRYFAQLVSALRFCHRHGVAHRDLKPQNLLLDAAGNLKVSDFGLSALPEHLH 556
Query: 59 DGRRMSGVVGTPYYVAPEVL--MGWENGEKVDVWSCGVI 95
DG + GTP + APE+L +G++ G D WSCGVI
Sbjct: 557 DG-LLHTACGTPAFTAPEILRRVGYD-GSXADAWSCGVI 667
>TC216364 similar to UP|Q9MAI7 (Q9MAI7) F12M16.4, partial (58%)
Length = 2224
Score = 78.2 bits (191), Expect = 1e-15
Identities = 38/96 (39%), Positives = 58/96 (59%), Gaps = 2/96 (2%)
Frame = +1
Query: 1 GGPIPEVEVASLMKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFG-SAEWFGD 59
G P+ E+ +A +++ LL A+ + H G H D+K N+L GD+K+ADFG SA+
Sbjct: 130 GPPLDEMSIACILRDLLHAVDYLHSEGKIHRDIKAANILLSENGDVKVADFGVSAQLTRT 309
Query: 60 GRRMSGVVGTPYYVAPEVLMGWEN-GEKVDVWSCGV 94
R VGTP+++APEV+ + EK D+WS G+
Sbjct: 310 ISRRKTFVGTPFWMAPEVIQNTDGYNEKADIWSLGI 417
>TC209238 similar to UP|Q8LK24 (Q8LK24) SOS2-like protein kinase, partial
(53%)
Length = 926
Score = 77.8 bits (190), Expect = 2e-15
Identities = 45/106 (42%), Positives = 66/106 (61%), Gaps = 7/106 (6%)
Frame = +2
Query: 8 EVASL-MKQLLEAITHCHRLGVAHHDVKPDNVLFGAGGDLKLADFGSAEWFGDGRRMSGV 66
+VA L +QL+ A+ CH GV H D+KP+N+L G+LK++DFG F + + G+
Sbjct: 536 DVARLYFQQLISAVDFCHSRGVYHRDLKPENLLLDEHGNLKVSDFGLTA-FSEHLKEDGL 712
Query: 67 V----GTPYYVAPEVL--MGWENGEKVDVWSCGVIFEAVIWGNLRF 106
+ GTP YV+PEV+ G++ G K D+WSCGVI ++ G L F
Sbjct: 713 LHTTCGTPAYVSPEVIAKKGYD-GAKADIWSCGVILYVLLAGFLPF 847
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.334 0.150 0.505
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,547,636
Number of Sequences: 63676
Number of extensions: 100355
Number of successful extensions: 1670
Number of sequences better than 10.0: 631
Number of HSP's better than 10.0 without gapping: 1374
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1397
length of query: 150
length of database: 12,639,632
effective HSP length: 89
effective length of query: 61
effective length of database: 6,972,468
effective search space: 425320548
effective search space used: 425320548
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.5 bits)
S2: 54 (25.4 bits)
Lotus: description of TM0252a.5


