

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0247.8
(170 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
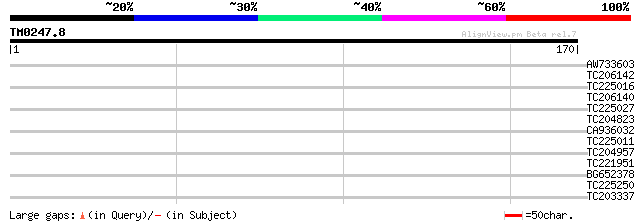
Score E
Sequences producing significant alignments: (bits) Value
AW733603 similar to GP|14289169|dbj pectate lyase {Salix gilgian... 29 0.95
TC206142 similar to UP|Q6U7H9 (Q6U7H9) Pectate lyase, partial (26%) 29 0.95
TC225016 similar to UP|Q39835 (Q39835) Extensin, partial (32%) 29 1.2
TC206140 similar to UP|Q6U7H9 (Q6U7H9) Pectate lyase, partial (25%) 28 1.6
TC225027 similar to UP|Q39835 (Q39835) Extensin, partial (49%) 28 1.6
TC204823 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, part... 28 2.1
CA936032 27 4.7
TC225011 similar to UP|Q09083 (Q09083) Hydroxyproline-rich glyco... 27 4.7
TC204957 homologue to UP|Q8LJU3 (Q8LJU3) Ascorbate oxidase (Frag... 27 6.2
TC221951 UP|NO75_SOYBN (P08297) Early nodulin 75 precursor (N-75... 27 6.2
BG652378 similar to PIR|T10863|T108 extensin precursor - kidney ... 26 8.1
TC225250 weakly similar to GB|AAO64828.1|29028898|BT005893 At3g5... 26 8.1
TC203337 similar to UP|Q9S728 (Q9S728) En/Spm-like transposon pr... 26 8.1
>AW733603 similar to GP|14289169|dbj pectate lyase {Salix gilgiana}, partial
(21%)
Length = 262
Score = 29.3 bits (64), Expect = 0.95
Identities = 12/39 (30%), Positives = 22/39 (55%)
Frame = -2
Query: 77 RSHASVEFHAFHKVSRLSGVSSIHKVSRLSEASRLIPWM 115
+S + ++FH FH +S S ++ S A+ L+PW+
Sbjct: 120 KSPSDLQFHPFHSLSAASSCFVTSLLNLSSGATNLLPWL 4
>TC206142 similar to UP|Q6U7H9 (Q6U7H9) Pectate lyase, partial (26%)
Length = 723
Score = 29.3 bits (64), Expect = 0.95
Identities = 12/39 (30%), Positives = 22/39 (55%)
Frame = -3
Query: 77 RSHASVEFHAFHKVSRLSGVSSIHKVSRLSEASRLIPWM 115
+S + ++FH FH +S S ++ S A+ L+PW+
Sbjct: 187 KSPSDLQFHPFHSLSAASSCFVTSLLNLSSGATNLLPWL 71
>TC225016 similar to UP|Q39835 (Q39835) Extensin, partial (32%)
Length = 973
Score = 28.9 bits (63), Expect = 1.2
Identities = 24/97 (24%), Positives = 36/97 (36%)
Frame = +3
Query: 47 IKRVQPLTMTLRVINTLTKSHASVKFHALTRSHASVEFHAFHKVSRLSGVSSIHKVSRLS 106
++ + +T + + T T H H L RSH ++ H LS + H
Sbjct: 3 LRHLLSITTSTLTLPTFTPPH-----HLLLRSHTTITHHPLLHHLHLSHTITTHHHHHHQ 167
Query: 107 EASRLIPWMANKLLNERRSTNILRTYQTPQHLDPTTL 143
+ + + LL R T I TP HL TL
Sbjct: 168 RRNHTTTTLHHLLLQRRNHTTI-----TPHHLLTLTL 263
>TC206140 similar to UP|Q6U7H9 (Q6U7H9) Pectate lyase, partial (25%)
Length = 683
Score = 28.5 bits (62), Expect = 1.6
Identities = 17/42 (40%), Positives = 23/42 (54%), Gaps = 3/42 (7%)
Frame = -3
Query: 77 RSHASVEFHAFHKVSRLSGVSSIHKVSRL---SEASRLIPWM 115
+S + ++FH FH LSG SS S L S A+ L PW+
Sbjct: 162 KSPSDLQFHPFHS---LSGASSCFVTSLLNLSSGATNLFPWL 46
>TC225027 similar to UP|Q39835 (Q39835) Extensin, partial (49%)
Length = 747
Score = 28.5 bits (62), Expect = 1.6
Identities = 19/54 (35%), Positives = 26/54 (47%)
Frame = +3
Query: 41 RQNTIIIKRVQPLTMTLRVINTLTKSHASVKFHALTRSHASVEFHAFHKVSRLS 94
R+N I LT+TL + TLT + H L RSH S H H+ + L+
Sbjct: 531 RRNHTTITPHHLLTLTLTLTLTLTLPSSITLLHHLLRSHTSTH-HLHHQYTLLT 689
>TC204823 similar to UP|Q8SMH8 (Q8SMH8) RNA-binding protein, partial (66%)
Length = 1596
Score = 28.1 bits (61), Expect = 2.1
Identities = 13/28 (46%), Positives = 18/28 (63%)
Frame = +3
Query: 9 SQTTSPTSLDTSPRVLAKLMSLKAYPKV 36
S ++P SL SPR L L S +A+PK+
Sbjct: 186 SHASTPQSLSPSPRELTALPSSRAWPKL 269
>CA936032
Length = 454
Score = 26.9 bits (58), Expect = 4.7
Identities = 15/43 (34%), Positives = 22/43 (50%)
Frame = +2
Query: 27 LMSLKAYPKVKSILRQNTIIIKRVQPLTMTLRVINTLTKSHAS 69
L+ K K K ++ I +KR P+T TL + + TKS S
Sbjct: 11 LLKKKKKKKKKKKKKKTAIAMKRTNPMTATLMIPPSPTKSSPS 139
>TC225011 similar to UP|Q09083 (Q09083) Hydroxyproline-rich glycoprotein
precursor, partial (20%)
Length = 964
Score = 26.9 bits (58), Expect = 4.7
Identities = 28/107 (26%), Positives = 41/107 (38%), Gaps = 5/107 (4%)
Frame = +3
Query: 44 TIIIKRVQPLTMTLRVINTLTKSHASVKFHALTRSHASVEFHAFHKVSRLSGVSSIHKVS 103
T+ + LT TL TLT ++ H L RSH + H + L + I +
Sbjct: 87 TLTLTHTHTLTHTL----TLTLPMSTTLLHHLLRSHTTTILHHLQSTTTL---TPIFTPT 245
Query: 104 RLSEASRLIPWMANKLLNERRSTNILRTYQTPQH-----LDPTTLLV 145
L+ +L+ +ST L TP H L PT L+
Sbjct: 246 HTLILHHLLRSHTTIILHHLQSTPTLTPILTPTHTLTPILTPTHTLI 386
>TC204957 homologue to UP|Q8LJU3 (Q8LJU3) Ascorbate oxidase (Fragment),
complete
Length = 1535
Score = 26.6 bits (57), Expect = 6.2
Identities = 17/60 (28%), Positives = 26/60 (43%)
Frame = +2
Query: 79 HASVEFHAFHKVSRLSGVSSIHKVSRLSEASRLIPWMANKLLNERRSTNILRTYQTPQHL 138
H +E H H + +HKV ++ + NKL+ R N L TYQ ++L
Sbjct: 1160 HCHIEPH-LHMGMGVIFAEGVHKVGKIPRDALTCGLTGNKLVGNRHY*N*LATYQHQKNL 1336
>TC221951 UP|NO75_SOYBN (P08297) Early nodulin 75 precursor (N-75) (NGM-75),
complete
Length = 1179
Score = 26.6 bits (57), Expect = 6.2
Identities = 26/99 (26%), Positives = 43/99 (43%), Gaps = 2/99 (2%)
Frame = +2
Query: 42 QNTIIIKRVQPLTMTLRVINTLTKSHASVKFHALTRSHASVEFHAFHKVSRLSGVSSIHK 101
QNT ++ R P+ + R N L +SH S + L RSH H + +
Sbjct: 293 QNTNLLMRNHPMRIHHRSTNHLMRSHQST--NHLMRSHHQSMNHLMRNHHQNTNHLMRSH 466
Query: 102 VSRLSEASRLIPWMANKLL-NERRSTN-ILRTYQTPQHL 138
+ R N L+ + +STN ++R++Q+ HL
Sbjct: 467 HQNTNHLMRNHHQNTNHLMRSHHQSTNHLMRSHQSTSHL 583
>BG652378 similar to PIR|T10863|T108 extensin precursor - kidney bean,
partial (14%)
Length = 361
Score = 26.2 bits (56), Expect = 8.1
Identities = 13/41 (31%), Positives = 20/41 (48%)
Frame = +1
Query: 54 TMTLRVINTLTKSHASVKFHALTRSHASVEFHAFHKVSRLS 94
T+TL + TLT ++ H L RSH + H + L+
Sbjct: 211 TLTLTLTLTLTLPMSTTLLHHLLRSHTTTILHHLQSTTTLT 333
>TC225250 weakly similar to GB|AAO64828.1|29028898|BT005893 At3g56480
{Arabidopsis thaliana;} , partial (54%)
Length = 1302
Score = 26.2 bits (56), Expect = 8.1
Identities = 16/57 (28%), Positives = 28/57 (49%)
Frame = -1
Query: 69 SVKFHALTRSHASVEFHAFHKVSRLSGVSSIHKVSRLSEASRLIPWMANKLLNERRS 125
S+K +++ S+ FH FH VS + + S+A++ +PW K + R S
Sbjct: 1278 SIKIQSISTSYTGHSFHQFHNVSSKTFI*C------HSKANKNMPWCW*KKIE*RNS 1126
>TC203337 similar to UP|Q9S728 (Q9S728) En/Spm-like transposon protein
(Protodermal factor 1), partial (46%)
Length = 1410
Score = 26.2 bits (56), Expect = 8.1
Identities = 12/31 (38%), Positives = 17/31 (54%), Gaps = 2/31 (6%)
Frame = -2
Query: 72 FHALTRSHASVEFHAFHKVSRLSG--VSSIH 100
FH RS VE H FH++ + G +S+ H
Sbjct: 434 FHHHLRSGEGVEIHGFHQLEDIDGDWISNFH 342
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.131 0.375
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,481,226
Number of Sequences: 63676
Number of extensions: 94666
Number of successful extensions: 557
Number of sequences better than 10.0: 27
Number of HSP's better than 10.0 without gapping: 552
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 557
length of query: 170
length of database: 12,639,632
effective HSP length: 90
effective length of query: 80
effective length of database: 6,908,792
effective search space: 552703360
effective search space used: 552703360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 55 (25.8 bits)
Lotus: description of TM0247.8


