

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
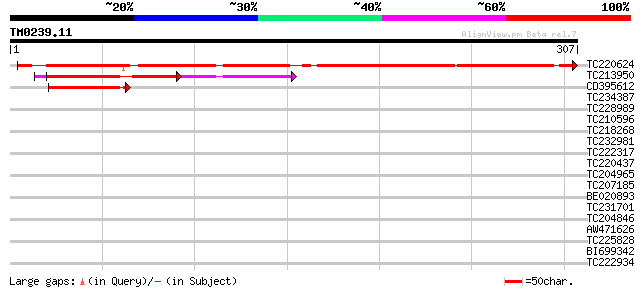
Query= TM0239.11
(307 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC220624 similar to UP|SCWA_YEAST (Q04951) Probable family 17 gl... 356 6e-99
TC213950 UP|Q6U679 (Q6U679) Kunitz trypsin inhibitor 5, partial ... 85 4e-17
CD395612 55 4e-08
TC234387 33 0.21
TC228989 similar to UP|Q9LSS5 (Q9LSS5) Myosin heavy chain-like, ... 33 0.21
TC210596 similar to UP|Q9AXJ6 (Q9AXJ6) Phosphatase-like protein ... 31 0.62
TC218268 30 1.8
TC232981 UP|Q9LLR0 (Q9LLR0) Carboxyl transferase alpha subunit ... 29 2.3
TC222317 UP|Q9LLQ8 (Q9LLQ8) Carboxyl transferase alpha subunit ... 29 2.3
TC220437 29 2.3
TC204965 homologue to UP|HMDH_CAMAC (P48021) 3-hydroxy-3-methylg... 29 3.1
TC207185 similar to UP|Q9MAX9 (Q9MAX9) Peroxidase precursor , p... 29 3.1
BE020893 homologue to GP|14486705|gb| phosphatidylinositol trans... 29 3.1
TC231701 28 4.0
TC204846 GB|AAP47437.1|32331819|AY164863 RTNLB47 {Glycine max;} ... 28 4.0
AW471626 similar to SP|P11827|GLCX Beta-conglycinin alpha' chai... 28 4.0
TC225828 weakly similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like ara... 28 4.0
BI699342 28 4.0
TC222934 28 5.2
>TC220624 similar to UP|SCWA_YEAST (Q04951) Probable family 17 glucosidase
SCW10 precursor (Soluble cell wall protein 10) ,
partial (5%)
Length = 985
Score = 356 bits (914), Expect = 6e-99
Identities = 193/305 (63%), Positives = 231/305 (75%), Gaps = 2/305 (0%)
Frame = +2
Query: 5 QQKAKEFFSNHGGAPTPKATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLF--LH 62
++KAKEFF TP+ F+LLTIT+LSLLLPLSF LLASLS A++YLQT TLF +
Sbjct: 2 REKAKEFF-------TPREVFYLLTITILSLLLPLSFLLLASLSGAQYYLQTLTLFQSYY 160
Query: 63 SPQQQQQPFSFLFTLALQINPCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPSL 122
SPQ PF +LF LAL INPCILY LVSIVS+ TLI GL GKITL ++ P +
Sbjct: 161 SPQ----PFPYLFHLALNINPCILYFLVSIVSMGTLIQGLTGKITLFSEL---PSKSSRI 319
Query: 123 YIAWILLCAFQFCVGLGIEASIAAGIFEFDNSSSSSSSSFGESAAERSLLSKVFFLLGLH 182
AWILLCAFQ CVGLGIE SIAAG+++ D SSFG ERSLLS++ FLLGLH
Sbjct: 320 STAWILLCAFQVCVGLGIEGSIAAGLYDDD------VSSFG---TERSLLSRMIFLLGLH 472
Query: 183 ETTQVWYRVVVRPVVDDTVFGGAKKEKWIERVAVAASLGVLWWWRLREEVESLVVMAEAK 242
ETT VW R+VVRPVVDDTVFGGA++E+W+ERV +AA LG LWWWRLREEVE+LV++A+
Sbjct: 473 ETTHVWCRMVVRPVVDDTVFGGARRERWVERVGLAAGLGTLWWWRLREEVETLVLVAQV- 649
Query: 243 KSEQFGELELKDFIGWWLYYVTVTIGMVRIVKGLMWMFMVALCRRRVTEVPVSPVVESSL 302
KSEQ ++ + DF+GWWLYY+ VTIGMVRIVKGL+W+FM++LCRRR T + VES
Sbjct: 650 KSEQLMDVGIGDFVGWWLYYLIVTIGMVRIVKGLIWIFMISLCRRRPTTTTIE--VESIQ 823
Query: 303 NDDKV 307
NDDKV
Sbjct: 824 NDDKV 838
>TC213950 UP|Q6U679 (Q6U679) Kunitz trypsin inhibitor 5, partial (5%)
Length = 403
Score = 85.1 bits (209), Expect = 4e-17
Identities = 45/73 (61%), Positives = 53/73 (71%)
Frame = +3
Query: 21 PKATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQPFSFLFTLALQ 80
P+ +LLTIT+LSLLLPLSF LLA+LS A++YLQ T + QPF +LF LAL
Sbjct: 24 PREVLYLLTITILSLLLPLSFLLLATLSGAQYYLQALTYY------SSQPFPYLFHLALN 185
Query: 81 INPCILYVLVSIV 93
INPCILY LVSIV
Sbjct: 186INPCILYFLVSIV 224
Score = 77.8 bits (190), Expect = 6e-15
Identities = 57/142 (40%), Positives = 73/142 (51%)
Frame = +2
Query: 14 NHGGAPTPKATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQPFSF 73
+H P P F L+ +L +LP S LL +L T +L P Q QP
Sbjct: 50 HHNPQPPPPFVFLTLSHSLRCPILPSSPHLL--------FLTTLSLSF-PPCPQYQPMYS 202
Query: 74 LFTLALQINPCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPSLYIAWILLCAFQ 133
LF+ V ++ TLI GLMGKITLL++ P + AWILLCAFQ
Sbjct: 203 LFSR------------VHCKTVGTLIQGLMGKITLLSEL---PSKSSRISTAWILLCAFQ 337
Query: 134 FCVGLGIEASIAAGIFEFDNSS 155
CVGLGIE SIAAG+++ D+ S
Sbjct: 338 VCVGLGIEGSIAAGLYDDDDVS 403
>CD395612
Length = 432
Score = 55.1 bits (131), Expect = 4e-08
Identities = 29/44 (65%), Positives = 33/44 (74%)
Frame = +2
Query: 22 KATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSPQ 65
K FHL T TLL+LLLPLSF LLA S A++YLQT T + HSPQ
Sbjct: 254 KEIFHLFTFTLLTLLLPLSFLLLAKFSDAQYYLQTLTGY-HSPQ 382
>TC234387
Length = 755
Score = 32.7 bits (73), Expect = 0.21
Identities = 41/139 (29%), Positives = 62/139 (44%), Gaps = 6/139 (4%)
Frame = -2
Query: 9 KEFFSNHGGAPTPKATFHLLTITLLSLLLPLS---FFLLASLSAAKHYLQTFTLFLHSPQ 65
K FFS H +P+ F L ++ LL LS FFLL + QT L P
Sbjct: 505 KHFFS-HSLSPS----FSLFSLFLLYPQDSLSLHYFFLLLFMLLLVQLFQTKGTILPLP- 344
Query: 66 QQQQPFSFLFTLALQINPCILYVLVS---IVSIATLINGLMGKITLLNDSSTSPVLQPSL 122
P + + L I ILY+ +S + + L+ L+ I +L+ P+L L
Sbjct: 343 ----PLTIVLVLFSFIY--ILYMSLSQRGFILLVLLLYQLLKLIHILSIFIELPLL---L 191
Query: 123 YIAWILLCAFQFCVGLGIE 141
Y W+++ +FCVG +E
Sbjct: 190 YDGWVIVIVLEFCVGESVE 134
>TC228989 similar to UP|Q9LSS5 (Q9LSS5) Myosin heavy chain-like, partial
(23%)
Length = 884
Score = 32.7 bits (73), Expect = 0.21
Identities = 45/176 (25%), Positives = 72/176 (40%), Gaps = 4/176 (2%)
Frame = -1
Query: 27 LLTITLLSLLLPLSFFLLASLSAA-KHYLQTFTLFLHSPQQQQQPFSFLFT---LALQIN 82
LL ++ S+ +P F+ AS SAA T L P FS F+ L ++
Sbjct: 599 LLLLSASSITIPS--FMRASCSAALAPSTSAATSSLALPLLLISDFSISFSSDMLCSVVS 426
Query: 83 PCILYVLVSIVSIATLINGLMGKITLLNDSSTSPVLQPSLYIAWILLCAFQFCVGLGIEA 142
I+ S A+L+ L+ + L+D +S + + + + + L +
Sbjct: 425 LSIISAFSSATHSASLLISLLSLCSFLDDMLSSSFRFKLSFSSTMPCNSVSLSIRLAFNS 246
Query: 143 SIAAGIFEFDNSSSSSSSSFGESAAERSLLSKVFFLLGLHETTQVWYRVVVRPVVD 198
SI+A F F NS+SSS S +S S S F + + WY V + D
Sbjct: 245 SISA--FFFFNSASSSHSLLPDSDFICSTNSYAFLICTMLRICSSWYLVSAVSIAD 84
>TC210596 similar to UP|Q9AXJ6 (Q9AXJ6) Phosphatase-like protein Psc923,
partial (27%)
Length = 570
Score = 31.2 bits (69), Expect = 0.62
Identities = 17/50 (34%), Positives = 29/50 (58%)
Frame = -2
Query: 120 PSLYIAWILLCAFQFCVGLGIEASIAAGIFEFDNSSSSSSSSFGESAAER 169
P+L+ A ++L F+ +G+GI IA G FE ++ + ++ G AER
Sbjct: 494 PTLFAAGVVLVGFEIGIGIGI--IIAGGAFEIAEGANFALAAGGGFLAER 351
>TC218268
Length = 853
Score = 29.6 bits (65), Expect = 1.8
Identities = 22/58 (37%), Positives = 29/58 (49%)
Frame = +1
Query: 119 QPSLYIAWILLCAFQFCVGLGIEASIAAGIFEFDNSSSSSSSSFGESAAERSLLSKVF 176
Q L I + + C GL AS++A IF F+N+ S SS F R LL+ VF
Sbjct: 514 QTGLQFQQIYIYIYIQCQGLRKCASVSAVIFIFNNTLSGQSSQF----VARLLLALVF 675
>TC232981 UP|Q9LLR0 (Q9LLR0) Carboxyl transferase alpha subunit , complete
Length = 2799
Score = 29.3 bits (64), Expect = 2.3
Identities = 18/40 (45%), Positives = 25/40 (62%)
Frame = -3
Query: 141 EASIAAGIFEFDNSSSSSSSSFGESAAERSLLSKVFFLLG 180
E S+A+ F S+S+SSS+ G A +RSL + FLLG
Sbjct: 1690 EDSLASKTCCFSFSTSTSSSASGILAIDRSLFFMLAFLLG 1571
>TC222317 UP|Q9LLQ8 (Q9LLQ8) Carboxyl transferase alpha subunit , complete
Length = 2521
Score = 29.3 bits (64), Expect = 2.3
Identities = 18/40 (45%), Positives = 25/40 (62%)
Frame = -1
Query: 141 EASIAAGIFEFDNSSSSSSSSFGESAAERSLLSKVFFLLG 180
E S+A+ F S+S+SSS+ G A +RSL + FLLG
Sbjct: 1717 EDSLASKTCCFSFSTSTSSSASGILAIDRSLFFMLAFLLG 1598
>TC220437
Length = 369
Score = 29.3 bits (64), Expect = 2.3
Identities = 15/44 (34%), Positives = 25/44 (56%)
Frame = -1
Query: 74 LFTLALQINPCILYVLVSIVSIATLINGLMGKITLLNDSSTSPV 117
L T++LQIN CI Y+ +S + + + + L KI +L P+
Sbjct: 357 LLTISLQINACI-YISISALIVK*VTH*LQNKINMLQTHQNEPM 229
>TC204965 homologue to UP|HMDH_CAMAC (P48021)
3-hydroxy-3-methylglutaryl-coenzyme A reductase
(HMG-CoA reductase) , partial (15%)
Length = 690
Score = 28.9 bits (63), Expect = 3.1
Identities = 20/63 (31%), Positives = 28/63 (43%)
Frame = -1
Query: 22 KATFHLLTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSPQQQQQPFSFLFTLALQI 81
K T ++ + LLS L + A + Q+ +FLH P Q +FLF L I
Sbjct: 483 KGTGNMWSRILLSSLQIIKDLHRQQCKAKRG*QQSNLMFLHFPSQISLTLNFLFILESTI 304
Query: 82 NPC 84
N C
Sbjct: 303 NDC 295
>TC207185 similar to UP|Q9MAX9 (Q9MAX9) Peroxidase precursor , partial (77%)
Length = 1072
Score = 28.9 bits (63), Expect = 3.1
Identities = 13/28 (46%), Positives = 18/28 (63%)
Frame = -1
Query: 24 TFHLLTITLLSLLLPLSFFLLASLSAAK 51
T H+ I+L LL+PL FFL++ L K
Sbjct: 823 TCHMTMISLHHLLIPLQFFLISPLDPVK 740
>BE020893 homologue to GP|14486705|gb| phosphatidylinositol transfer-like
protein III {Lotus japonicus}, partial (10%)
Length = 410
Score = 28.9 bits (63), Expect = 3.1
Identities = 14/32 (43%), Positives = 18/32 (55%)
Frame = +2
Query: 19 PTPKATFHLLTITLLSLLLPLSFFLLASLSAA 50
P P+A H+LT + LL PL F +A S A
Sbjct: 89 PAPRARLHILTTRAILLLPPLGFSHIAECSFA 184
>TC231701
Length = 764
Score = 28.5 bits (62), Expect = 4.0
Identities = 13/39 (33%), Positives = 18/39 (45%)
Frame = -3
Query: 45 ASLSAAKHYLQTFTLFLHSPQQQQQPFSFLFTLALQINP 83
+SL A HY FLHSP + F F +++ P
Sbjct: 240 SSLLLAPHYFSGVVAFLHSPHSENHHFLF*CPWLVEVEP 124
>TC204846 GB|AAP47437.1|32331819|AY164863 RTNLB47 {Glycine max;} , complete
Length = 1113
Score = 28.5 bits (62), Expect = 4.0
Identities = 29/89 (32%), Positives = 41/89 (45%), Gaps = 3/89 (3%)
Frame = +2
Query: 137 GLGIEASIAAGIFEFDNSSSSSSSSFGESAAERSLLSKVFFLLGLHETTQVWYRVVVRPV 196
G + I+ I + D+SSSS S + SA++ SL SKVF L G +P+
Sbjct: 116 GESLLEKISGKIHDHDSSSSSDSDNEKTSASD-SLKSKVFRLFGRE-----------KPI 259
Query: 197 VDDTVFGGAKKEK---WIERVAVAASLGV 222
V GG K W + A++LGV
Sbjct: 260 --HHVLGGGKPADVFLWRNKKISASTLGV 340
>AW471626 similar to SP|P11827|GLCX Beta-conglycinin alpha' chain precursor.
[Soybean] {Glycine max}, partial (26%)
Length = 517
Score = 28.5 bits (62), Expect = 4.0
Identities = 19/57 (33%), Positives = 26/57 (45%), Gaps = 5/57 (8%)
Frame = -1
Query: 28 LTITLLSLLLPLSFFLLASLSAAKHYLQTFTLFLHSPQQ-----QQQPFSFLFTLAL 79
L + L L+LPL L S+ ++ TF LFL + + PF FL L L
Sbjct: 514 LPLVLFLLVLPLVLLLFLSMRLVCAWVFTFILFLFTSLSSVFFLHEMPFVFLLLLVL 344
>TC225828 weakly similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like
arabinogalactan-protein 2, partial (41%)
Length = 1108
Score = 28.5 bits (62), Expect = 4.0
Identities = 18/43 (41%), Positives = 22/43 (50%), Gaps = 3/43 (6%)
Frame = -1
Query: 151 FDNSSSSSSSSFGESAAERS---LLSKVFFLLGLHETTQVWYR 190
F +S SS S G SAA S L+ FFL GL ++ W R
Sbjct: 673 F*SSESSDGPSAGASAASESSLPFLAFFFFLAGLGDSAGAWSR 545
>BI699342
Length = 325
Score = 28.5 bits (62), Expect = 4.0
Identities = 19/57 (33%), Positives = 29/57 (50%)
Frame = +3
Query: 122 LYIAWILLCAFQFCVGLGIEASIAAGIFEFDNSSSSSSSSFGESAAERSLLSKVFFL 178
LY+ +++ +F V I IA +SSSSSSSS S++ S +V+ L
Sbjct: 117 LYLIYVIAISFVLHV*KNIVKLIAFSSSSSSSSSSSSSSSSSSSSSSSSSAFEVYHL 287
>TC222934
Length = 982
Score = 28.1 bits (61), Expect = 5.2
Identities = 20/55 (36%), Positives = 29/55 (52%), Gaps = 4/55 (7%)
Frame = -3
Query: 27 LLTITLLSLLLPLSFFLLASLSAA----KHYLQTFTLFLHSPQQQQQPFSFLFTL 77
LL + L+ ++ L FF AS+ A KH+ LF S + Q+ F F+FTL
Sbjct: 794 LLVVVPLAFIIQL-FFCFASVQFARVSDKHWQYACMLFAISQSKIQE*FVFIFTL 633
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.137 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,507,576
Number of Sequences: 63676
Number of extensions: 287427
Number of successful extensions: 3338
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 3006
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3237
length of query: 307
length of database: 12,639,632
effective HSP length: 97
effective length of query: 210
effective length of database: 6,463,060
effective search space: 1357242600
effective search space used: 1357242600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0239.11


