

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
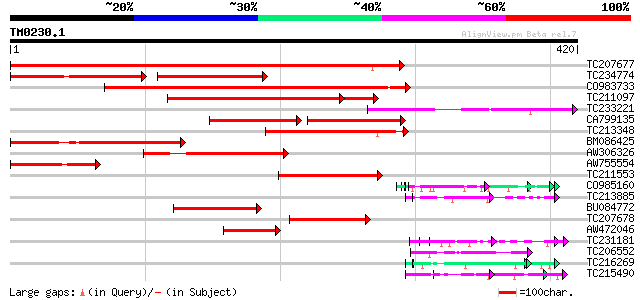
Query= TM0230.1
(420 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC207677 328 3e-90
TC234774 182 5e-89
CO983733 313 9e-86
TC211097 206 1e-60
TC233221 UP|Q9NK49 (Q9NK49) Dac protein (Fragment), partial (5%) 175 4e-44
CA799135 103 7e-39
TC213348 125 3e-29
BM086425 125 3e-29
AW306326 122 2e-28
AW755554 120 1e-27
TC211553 108 4e-24
CO985160 91 1e-18
TC213885 weakly similar to UP|Q9VUB7 (Q9VUB7) CG32133-PA, partia... 85 5e-17
BU084772 85 7e-17
TC207678 75 7e-14
AW472046 homologue to GP|18844910|dbj P0519D04.38 {Oryza sativa ... 59 4e-09
TC231181 weakly similar to UP|Q86PD9 (Q86PD9) RE65015p, partial ... 57 2e-08
TC206552 weakly similar to UP|Q8SX56 (Q8SX56) LD47350p (CG6824-P... 55 5e-08
TC216269 similar to UP|E6_GOSHI (Q01197) Protein E6, partial (17%) 54 1e-07
TC215490 weakly similar to UP|Q8I7C3 (Q8I7C3) Lasp protein, part... 50 1e-06
>TC207677
Length = 1373
Score = 328 bits (841), Expect = 3e-90
Identities = 163/306 (53%), Positives = 219/306 (71%), Gaps = 14/306 (4%)
Frame = +1
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQL-KKRDDGKLTVVSAASPSVPIPHADDS-SLDVLLVD 58
MG + S +AAKFAFFPP PP+Y + + G + A P + IP ++DVL +
Sbjct: 88 MGGVTSSIAAKFAFFPPHPPSYTVVAAAEGGGFSDPPPAPPRLAIPEVPSKDNVDVLKLR 267
Query: 59 TKHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGAS 118
T+ GN+IVA Y+K T+LYSHGNAADLGQ+++LFV+L LR+N+MGYDYSGYG S
Sbjct: 268 TRRGNEIVALYVKYHRPTCTMLYSHGNAADLGQMFELFVELSNRLRLNVMGYDYSGYGQS 447
Query: 119 TGKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHS 178
TGKP+E +TYADI+A Y+CL+ +YGV E +ILYGQSVGSGPTL LA+++ LRGV+LHS
Sbjct: 448 TGKPTECNTYADIDAAYKCLKEQYGVEDEQLILYGQSVGSGPTLDLASRIAELRGVILHS 627
Query: 179 GILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARES 238
ILSGLRVL VK ++ FDIYKNI+K+ VKCPVLVIHGT D+VV+ HG LW++ +
Sbjct: 628 PILSGLRVLYPVKRTYWFDIYKNIDKVGAVKCPVLVIHGTADEVVDVSHGKQLWELCKVK 807
Query: 239 YDPLWIKGGGHCNLELYPDYIRHLCKFIQ------------EMESITTEKRLKKLRQSQG 286
Y+PLW+ GGGHCNLELYP++I+HL KF+Q + +++ +E + K ++S+
Sbjct: 808 YEPLWVSGGGHCNLELYPEFIKHLKKFVQTIGKSKATANGSKKDTVESENQGKASKESES 987
Query: 287 PQSNSS 292
S +S
Sbjct: 988 GTSGTS 1005
>TC234774
Length = 747
Score = 182 bits (461), Expect(2) = 5e-89
Identities = 92/102 (90%), Positives = 97/102 (94%), Gaps = 1/102 (0%)
Frame = +1
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKR-DDGKLTVVSAASPSVPIPHADDSSLDVLLVDT 59
MGCMVSQLAAKFAFFPPSPPTYQLKK +DGKLTVVSAA+P IPHADD+SLDVLLVDT
Sbjct: 205 MGCMVSQLAAKFAFFPPSPPTYQLKKSGEDGKLTVVSAAAP---IPHADDTSLDVLLVDT 375
Query: 60 KHGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKV 101
KHGNKIVAFYL+NPYARLTLLYSHGNAADLGQLYDLFVQLK+
Sbjct: 376 KHGNKIVAFYLRNPYARLTLLYSHGNAADLGQLYDLFVQLKI 501
Score = 164 bits (415), Expect(2) = 5e-89
Identities = 79/82 (96%), Positives = 81/82 (98%)
Frame = +2
Query: 110 YDYSGYGASTGKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLP 169
YDYSGYGASTGKPSESSTYADIEAIYECLE+EYGV QED+ILYGQSVGSGPTLHLAAKLP
Sbjct: 500 YDYSGYGASTGKPSESSTYADIEAIYECLETEYGVSQEDVILYGQSVGSGPTLHLAAKLP 679
Query: 170 RLRGVVLHSGILSGLRVLCHVK 191
RLRGVVLHSGILSGLRVLCHVK
Sbjct: 680 RLRGVVLHSGILSGLRVLCHVK 745
>CO983733
Length = 753
Score = 313 bits (802), Expect = 9e-86
Identities = 143/227 (62%), Positives = 183/227 (79%)
Frame = -1
Query: 71 KNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTGKPSESSTYAD 130
K+P+AR T+LYSHGNAADLGQ++ ++L+ +LRVN+M YDYSGYGASTGKPSE +TY D
Sbjct: 753 KHPFARFTVLYSHGNAADLGQMHXXXIELRAHLRVNIMSYDYSGYGASTGKPSEFNTYCD 574
Query: 131 IEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSGILSGLRVLCHV 190
IEA+Y CL++EYG+ QE++ILYGQSVGSGPTLHLA+KL +LRGVVLHS ILSG+RVL V
Sbjct: 573 IEAVYNCLKNEYGIKQEELILYGQSVGSGPTLHLASKLQKLRGVVLHSAILSGIRVLYPV 394
Query: 191 KFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESYDPLWIKGGGHC 250
K +F FDI+KNI+KI+ V CPV VIHGT DD+V+W HG LW++++E YDPLW+KGGGHC
Sbjct: 393 KMTFWFDIFKNIDKIRHVNCPVFVIHGTNDDIVDWSHGKRLWELSKEKYDPLWVKGGGHC 214
Query: 251 NLELYPDYIRHLCKFIQEMESITTEKRLKKLRQSQGPQSNSSSCFSC 297
NLE +P+YI++L KFI ME ++ K+ K + +Q P S C
Sbjct: 213 NLETFPEYIKYLRKFINAMEKLSLTKQTNK-QLTQNPSITESRHNKC 76
>TC211097
Length = 694
Score = 206 bits (523), Expect(2) = 1e-60
Identities = 93/131 (70%), Positives = 113/131 (85%)
Frame = +3
Query: 118 STGKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLH 177
STGKPSE +TY DIEA+Y+CL+SEYG+ QED+ILYGQSVGSGPT+HLA KLP LRGVVLH
Sbjct: 3 STGKPSEFNTYYDIEAVYDCLKSEYGIKQEDLILYGQSVGSGPTIHLATKLPNLRGVVLH 182
Query: 178 SGILSGLRVLCHVKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARE 237
SGILSG+RVL VK +F FDI+KNI+KI+ V CPVLVIHGT D++V+W HG LW++++E
Sbjct: 183 SGILSGIRVLYPVKVTFWFDIFKNIDKIRHVDCPVLVIHGTNDEIVDWSHGKRLWELSKE 362
Query: 238 SYDPLWIKGGG 248
YDPLW+ G G
Sbjct: 363 KYDPLWVPGWG 395
Score = 46.2 bits (108), Expect(2) = 1e-60
Identities = 17/28 (60%), Positives = 23/28 (81%)
Frame = +1
Query: 246 GGGHCNLELYPDYIRHLCKFIQEMESIT 273
GGGHCNLE +P+YI+HL KF+ ME ++
Sbjct: 388 GGGHCNLEAFPEYIKHLRKFLNAMEKLS 471
>TC233221 UP|Q9NK49 (Q9NK49) Dac protein (Fragment), partial (5%)
Length = 651
Score = 175 bits (443), Expect = 4e-44
Identities = 84/166 (50%), Positives = 99/166 (59%), Gaps = 11/166 (6%)
Frame = +1
Query: 266 IQEMESITTEKRLKKLRQSQGPQSNSSSCFSCSCCRMKCCSNNCTGCSNCSCINPECCNC 325
IQEMES+TTEKRLKK+RQS +++ C CCR+KCC C CSNC
Sbjct: 1 IQEMESMTTEKRLKKIRQSANQSKSNTGCTCSRCCRVKCCRVKCPECSNC---------- 150
Query: 326 SCINPECFNCSCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFK- 384
C C+N S S+ C +CC PSC KCC LPKF +CF S C TK S PSCC
Sbjct: 151 ---------CCCMNVSWSINCPECCWRPSC-KCCRLPKFPSCFRSICRTKWSWPSCCATK 300
Query: 385 ---------CCCNLRCARPSCCCMSCFCWRCCVGKHS-GRDEKQSG 420
CCC + ARPSCCCM+CFCW+CCVGKHS G + KQ+G
Sbjct: 301 CSLSLPSCCCCCYPKRARPSCCCMNCFCWQCCVGKHSDGTNGKQNG 438
>CA799135
Length = 435
Score = 103 bits (256), Expect(2) = 7e-39
Identities = 52/68 (76%), Positives = 58/68 (84%)
Frame = +1
Query: 149 IILYGQSVGSGPTLHLAAKLPRLRGVVLHSGILSGLRVLCHVKFSFCFDIYKNINKIKKV 208
IILYGQSVGSGPTL LAAKLP+LR VVLHS ILSGLRV+ VK S+ FDIYKNI+KI V
Sbjct: 1 IILYGQSVGSGPTLDLAAKLPQLRAVVLHSPILSGLRVMYPVKRSYWFDIYKNIDKIPLV 180
Query: 209 KCPVLVIH 216
CP+L+IH
Sbjct: 181 NCPILIIH 204
Score = 75.9 bits (185), Expect(2) = 7e-39
Identities = 32/73 (43%), Positives = 48/73 (64%)
Frame = +3
Query: 221 DVVNWLHGNGLWKMARESYDPLWIKGGGHCNLELYPDYIRHLCKFIQEMESITTEKRLKK 280
+VV+ HG LW++ +E Y+PLW+KGG HC+LE +P+YIRHL KFI +E T+++ +
Sbjct: 207 EVVDCSHGKQLWELCKEKYEPLWLKGGNHCDLEQFPEYIRHLKKFIATVEKSTSQRYSFR 386
Query: 281 LRQSQGPQSNSSS 293
Q Q S+
Sbjct: 387 RSMEQFEQPRKST 425
>TC213348
Length = 623
Score = 125 bits (315), Expect = 3e-29
Identities = 57/111 (51%), Positives = 80/111 (71%), Gaps = 5/111 (4%)
Frame = +3
Query: 190 VKFSFCFDIYKNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESYDPLWIKGGGH 249
VK ++ FDIYKN++KI VKCPVLVIHGT D+VV+ HG LW++ ++ Y+PLW+KGG H
Sbjct: 6 VKRTYWFDIYKNVDKIPLVKCPVLVIHGTADEVVDCSHGKQLWELCQQKYEPLWLKGGNH 185
Query: 250 CNLELYPDYIRHLCKFIQEMES-----ITTEKRLKKLRQSQGPQSNSSSCF 295
CNLELYP+Y+RHL KFI +E ++ + + ++ QS+G S+ CF
Sbjct: 186 CNLELYPEYLRHLRKFISSVEKPPSQRVSFRRSIDRVEQSRG----STDCF 326
>BM086425
Length = 428
Score = 125 bits (315), Expect = 3e-29
Identities = 63/130 (48%), Positives = 89/130 (68%)
Frame = +1
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAASPSVPIPHADDSSLDVLLVDTK 60
MG + S +AAK AFFPPSPP+Y++ + V+ A P ++ + V+ T+
Sbjct: 61 MGGVTSSMAAKMAFFPPSPPSYEVVEEAATGALVLEA------FPRREN--VRVVKFGTR 216
Query: 61 HGNKIVAFYLKNPYARLTLLYSHGNAADLGQLYDLFVQLKVNLRVNLMGYDYSGYGASTG 120
G++IV Y+ +P A+ T+LYSHGNAAD+G + +L+V L +LRVNL GYDYSGYG S+G
Sbjct: 217 RGSEIVGVYIAHPMAKSTILYSHGNAADIGHMLELYVDLSTHLRVNLFGYDYSGYGQSSG 396
Query: 121 KPSESSTYAD 130
KPSE++TYAD
Sbjct: 397 KPSENNTYAD 426
>AW306326
Length = 425
Score = 122 bits (307), Expect = 2e-28
Identities = 62/107 (57%), Positives = 79/107 (72%)
Frame = +3
Query: 100 KVNLRVNLMGYDYSGYGASTGKPSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSG 159
K+ + YDY GYGAST I+A+Y CL++EYGV QE++I YG+S+GSG
Sbjct: 138 KIRKHSSSCSYDY*GYGAST-----------IKALYNCLKNEYGVKQEELIFYGRSIGSG 284
Query: 160 PTLHLAAKLPRLRGVVLHSGILSGLRVLCHVKFSFCFDIYKNINKIK 206
PTLHLA+KL +LRGVVLHS ILSG+RVL VK +F FDI+KNI+KI+
Sbjct: 285 PTLHLASKLQKLRGVVLHSAILSGIRVLYPVKMTFWFDIFKNIDKIR 425
>AW755554
Length = 436
Score = 120 bits (301), Expect = 1e-27
Identities = 60/67 (89%), Positives = 63/67 (93%)
Frame = +3
Query: 1 MGCMVSQLAAKFAFFPPSPPTYQLKKRDDGKLTVVSAASPSVPIPHADDSSLDVLLVDTK 60
MGCMVSQLAAKFAFFPPSPPTYQLKK +DGKLTVVSAA+ PIPHADD+SLDVLLVDTK
Sbjct: 243 MGCMVSQLAAKFAFFPPSPPTYQLKKNEDGKLTVVSAAA---PIPHADDTSLDVLLVDTK 413
Query: 61 HGNKIVA 67
HGNKIVA
Sbjct: 414 HGNKIVA 434
>TC211553
Length = 647
Score = 108 bits (270), Expect = 4e-24
Identities = 47/77 (61%), Positives = 60/77 (77%)
Frame = +3
Query: 200 KNINKIKKVKCPVLVIHGTEDDVVNWLHGNGLWKMARESYDPLWIKGGGHCNLELYPDYI 259
+NI+KI V CPVLVIHGT DDVV++ HG LW+ ++ Y+PLWIKGG HCNLELYP YI
Sbjct: 69 QNIDKIPLVNCPVLVIHGTADDVVDYSHGKQLWEHCKQKYEPLWIKGGNHCNLELYPQYI 248
Query: 260 RHLCKFIQEMESITTEK 276
+HL KFI +E+ + +K
Sbjct: 249 KHLKKFITAIETSSHKK 299
>CO985160
Length = 777
Score = 90.9 bits (224), Expect = 1e-18
Identities = 53/146 (36%), Positives = 56/146 (38%), Gaps = 25/146 (17%)
Frame = +1
Query: 287 PQSNSSSCFSCSCCRMK--CCSNNCTGC----SNCSC-INPECCNCSCINPECFNCSCIN 339
P C CSCC CC +C C S C C I CC CSC + C+ C C
Sbjct: 310 PDPTICCCCGCSCCNCGSCCCCCSCGSC*S*GSCCCCWICGSCCCCSCASCCCWTCCCCC 489
Query: 340 CSCSLKCTQCCCLPSCIKCCHLPKFSTCF----GSSCCTKCSRPSCCF------------ 383
CS C CC SC CC C GS CC CS SCC
Sbjct: 490 CS*GSCCCCCCS*GSCCCCCCS*GSCCCCCWS*GSCCCCCCS*GSCCCCCWS*GSCCCCM 669
Query: 384 --KCCCNLRCARPSCCCMSCFCWRCC 407
CCC C R CCC C C CC
Sbjct: 670 *GSCCCCCNCGR--CCCWFCIC*CCC 741
Score = 74.3 bits (181), Expect = 9e-14
Identities = 43/118 (36%), Positives = 46/118 (38%), Gaps = 8/118 (6%)
Frame = +1
Query: 294 CFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQ----- 348
C SC CC C S C C C C CC C C C C C SC C
Sbjct: 427 CGSCCCC--SCASCCCWTCCCCCCS*GSCCCCCCS*GSCCCCCCS*GSCCCCCWS*GSCC 600
Query: 349 CCCLP--SCIKCCHL-PKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCCCMSCFC 403
CCC SC CC C SCC C+ C +CCC C CCC+ C C
Sbjct: 601 CCCCS*GSCCCCCWS*GSCCCCM*GSCCCCCN----CGRCCCWF-CIC*CCCCICCCC 759
Score = 65.1 bits (157), Expect = 6e-11
Identities = 40/115 (34%), Positives = 42/115 (35%), Gaps = 18/115 (15%)
Frame = +1
Query: 291 SSSCFSC-SCCRMKCCSNNCT--GCSNCSCINPECCNCSCINPECFNC-----SCINCSC 342
S C SC SCC CC C+ C C C CC C C C C SC C C
Sbjct: 433 SCCCCSCASCCCWTCCCCCCS*GSCCCCCCS*GSCCCCCCS*GSCCCCCWS*GSCCCCCC 612
Query: 343 SLKCTQCCCLP----------SCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCC 387
S CCC SC CC+ G CC C CC CCC
Sbjct: 613 S*GSCCCCCWS*GSCCCCM*GSCCCCCNC-------GRCCCWFCIC*CCCCICCC 756
Score = 49.3 bits (116), Expect = 3e-06
Identities = 29/100 (29%), Positives = 32/100 (32%), Gaps = 5/100 (5%)
Frame = +1
Query: 293 SCFSC-----SCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCT 347
SC C SCC CC + C C C CC C C C +C C C
Sbjct: 532 SCCCCCCS*GSCC---CCCWS*GSCCCCCCS*GSCCCCCWS*GSCCCCM*GSCCCCCNCG 702
Query: 348 QCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCC 387
+CCC CC CC CCC
Sbjct: 703 RCCCWFCIC*CC---------------------CCICCCC 759
Score = 41.2 bits (95), Expect = 9e-04
Identities = 26/67 (38%), Positives = 32/67 (46%), Gaps = 7/67 (10%)
Frame = +1
Query: 296 SCSCCRMKCCS-NNCTGC-----SNCSCI-NPECCNCSCINPECFNCSCINCSCSLKCTQ 348
SC CC CCS +C C S C C+ CC C+C C+ C C C C + C
Sbjct: 592 SCCCC---CCS*GSCCCCCWS*GSCCCCM*GSCCCCCNCGRCCCWFCIC-*CCCCICC-- 753
Query: 349 CCCLPSC 355
CC L +C
Sbjct: 754 CC*LDNC 774
>TC213885 weakly similar to UP|Q9VUB7 (Q9VUB7) CG32133-PA, partial (4%)
Length = 540
Score = 85.1 bits (209), Expect = 5e-17
Identities = 46/120 (38%), Positives = 51/120 (42%), Gaps = 11/120 (9%)
Frame = -1
Query: 299 CCRMKCCSNNCTGCSNCSCINPECCNCS--------CINPECFNCSCINCSCSLKCTQCC 350
CC CC +C C +C C CC+C C C +C C +C C CC
Sbjct: 309 CC---CCGCSCCNCGSCCC----CCSCGSC*S*GSCCCCWSCGSCCCWSCCCCCS*GSCC 151
Query: 351 CL---PSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCCCMSCFCWRCC 407
C SC CC GS CC C SCC CCCN C R CCC C CW CC
Sbjct: 150 CCCS*GSCCCCC-------S*GSCCC--CM*GSCC--CCCN--CGR--CCCWFCICWCCC 16
Score = 51.2 bits (121), Expect = 9e-07
Identities = 25/67 (37%), Positives = 28/67 (41%), Gaps = 1/67 (1%)
Frame = -1
Query: 294 CFSC-SCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCL 352
C+SC SCC CC G C C CC C C C C +C C C +CCC
Sbjct: 216 CWSCGSCCCWSCCCCCS*GSCCCCCS*GSCC-CCCS*GSCCCCM*GSCCCCCNCGRCCCW 40
Query: 353 PSCIKCC 359
CC
Sbjct: 39 FCICWCC 19
Score = 32.7 bits (73), Expect = 0.31
Identities = 18/63 (28%), Positives = 20/63 (31%)
Frame = -1
Query: 297 CSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCI 356
C CC C C C+ CC C C+C C C CCC
Sbjct: 126 CCCCS*GSC---------CCCM*GSCCCC---------CNCGRCCCWFCICWCCC----- 16
Query: 357 KCC 359
CC
Sbjct: 15 -CC 10
Score = 32.0 bits (71), Expect = 0.53
Identities = 11/28 (39%), Positives = 12/28 (42%)
Frame = -1
Query: 379 PSCCFKCCCNLRCARPSCCCMSCFCWRC 406
P+ C CCC C CC C C C
Sbjct: 318 PTIC--CCCGCSCCNCGSCCCCCSCGSC 241
Score = 29.3 bits (64), Expect = 3.5
Identities = 14/32 (43%), Positives = 14/32 (43%), Gaps = 2/32 (6%)
Frame = -1
Query: 296 SCSCCR--MKCCSNNCTGCSNCSCINPECCNC 325
SC CC CC NC C CI CC C
Sbjct: 105 SCCCCM*GSCCCCCNCGRCCCWFCICWCCCCC 10
>BU084772
Length = 403
Score = 84.7 bits (208), Expect = 7e-17
Identities = 42/65 (64%), Positives = 50/65 (76%)
Frame = +1
Query: 122 PSESSTYADIEAIYECLESEYGVGQEDIILYGQSVGSGPTLHLAAKLPRLRGVVLHSGIL 181
PS+ +TY+DI A Y+CLE Y QE II+Y QSVGSGPTL LAA+LP+LR V+LH IL
Sbjct: 178 PSKQNTYSDIVAAYKCLEESYRAKQEGIIVYRQSVGSGPTLDLAARLPQLRVVILHCPIL 357
Query: 182 SGLRV 186
SG RV
Sbjct: 358 SGFRV 372
>TC207678
Length = 691
Score = 74.7 bits (182), Expect = 7e-14
Identities = 31/60 (51%), Positives = 41/60 (67%)
Frame = +2
Query: 208 VKCPVLVIHGTEDDVVNWLHGNGLWKMARESYDPLWIKGGGHCNLELYPDYIRHLCKFIQ 267
VKC V HGT D+VV HG ++ + Y+ LW+ GGGHCNLEL P++I+HL KF+Q
Sbjct: 2 VKCXVXXXHGTADEVVXVSHGKXXXELCKVKYEXLWVSGGGHCNLELXPEFIKHLKKFVQ 181
>AW472046 homologue to GP|18844910|dbj P0519D04.38 {Oryza sativa (japonica
cultivar-group)}, partial (7%)
Length = 127
Score = 58.9 bits (141), Expect = 4e-09
Identities = 30/42 (71%), Positives = 34/42 (80%)
Frame = +2
Query: 159 GPTLHLAAKLPRLRGVVLHSGILSGLRVLCHVKFSFCFDIYK 200
GPTL LA+ +P LRGVVLHS ILSGLRVL VK ++ FDIYK
Sbjct: 2 GPTLDLASLIPELRGVVLHSPILSGLRVLYPVKRTYWFDIYK 127
>TC231181 weakly similar to UP|Q86PD9 (Q86PD9) RE65015p, partial (10%)
Length = 671
Score = 57.0 bits (136), Expect = 2e-08
Identities = 35/112 (31%), Positives = 40/112 (35%), Gaps = 8/112 (7%)
Frame = -3
Query: 304 CCSNNCTGCSNCSCINPECCN---CSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCCH 360
CC N N + P C++ C CSC C CCC C CC
Sbjct: 567 CCDNTAPWPCNVTIACPWSATG*LLLCVDGCCCCIRFSGCSC---CRGCCC*KYCCCCC- 400
Query: 361 LPKFSTCFGSSCCTKCSRPSCCFKCCCNLRCARPSCC-----CMSCFCWRCC 407
TC S CC C+ CCC+ C CC C SC C CC
Sbjct: 399 ---CRTCCWS-CCG*RGSSRLCWSCCCDSPCWNDCCCN*I*CCCSCSCCCCC 256
Score = 49.7 bits (117), Expect = 2e-06
Identities = 35/104 (33%), Positives = 43/104 (40%), Gaps = 1/104 (0%)
Frame = -3
Query: 312 CSNCSCINPECCNCSCINPECFNCSC-INCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGS 370
C+ I P +C C N + C+ I C S C+ C CC + +FS C
Sbjct: 606 CTTGEEIGPSLSDC-CDNTAPWPCNVTIACPWSATG*LLLCVDGC--CCCI-RFSGC--- 448
Query: 371 SCCTKCSRPSCCFKCCCNLRCARPSCCCMSCFCWRCCVGKHSGR 414
SCC C CC K CC CCC CW CC + S R
Sbjct: 447 SCCRGC----CC*KYCC--------CCCCRTCCWSCCG*RGSSR 352
Score = 46.6 bits (109), Expect = 2e-05
Identities = 28/76 (36%), Positives = 33/76 (42%), Gaps = 12/76 (15%)
Frame = -3
Query: 297 CSCCRMKCCSNNCTGCSNCSCI---------NPECCNCSCINPECFNCSCIN---CSCSL 344
CSCCR CC C C +C + C +C C +P C+N C N C CS
Sbjct: 450 CSCCRGCCC*KYCCCCCCRTCCWSCCG*RGSSRLCWSCCCDSP-CWNDCCCN*I*CCCSC 274
Query: 345 KCTQCCCLPSCIKCCH 360
C CCC C CH
Sbjct: 273 SC--CCC---C**YCH 241
Score = 40.0 bits (92), Expect = 0.002
Identities = 31/93 (33%), Positives = 33/93 (35%)
Frame = -3
Query: 295 FSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPS 354
FS C C GC C CC C C C++C C S C CCC
Sbjct: 459 FSGCSC--------CRGC----CC*KYCC-CCCCRTCCWSC-CG*RGSSRLCWSCCCDSP 322
Query: 355 CIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCC 387
C C C CC C SCC CCC
Sbjct: 321 CWNDC------CCN*I*CCCSC---SCC--CCC 256
Score = 33.9 bits (76), Expect = 0.14
Identities = 17/43 (39%), Positives = 21/43 (48%), Gaps = 3/43 (6%)
Frame = -3
Query: 286 GPQSNSSSCFSCSC---CRMKCCSNNCTGCSNCSCINPECCNC 325
G + +S C+SC C C CC N C +CSC CC C
Sbjct: 372 G*RGSSRLCWSCCCDSPCWNDCCCN*I*CCCSCSC----CCCC 256
>TC206552 weakly similar to UP|Q8SX56 (Q8SX56) LD47350p (CG6824-PB), partial
(3%)
Length = 1401
Score = 55.5 bits (132), Expect = 5e-08
Identities = 31/94 (32%), Positives = 40/94 (41%), Gaps = 4/94 (4%)
Frame = -1
Query: 298 SCCRMKCCSNNC---TGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCL-P 353
+C C NC +G NC CC C+C +P C C+NC C + CCC
Sbjct: 714 TCPEGNCLP*NC*CPSG*ENCI*----CCPCNCPSP-CI-ADCLNCCC*ARIWWCCCC*G 553
Query: 354 SCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCC 387
+C+ C CC CS CC+ CCC
Sbjct: 552 NCMNFCR-----------CCCCCSTGGCCWVCCC 484
>TC216269 similar to UP|E6_GOSHI (Q01197) Protein E6, partial (17%)
Length = 1139
Score = 54.3 bits (129), Expect = 1e-07
Identities = 33/98 (33%), Positives = 38/98 (38%), Gaps = 10/98 (10%)
Frame = -2
Query: 299 CCRMKCCSNNCTGCSNCSCINPECCNCSC-INPECFNCS--CINCSCSLKCTQCCCLPSC 355
CC K C+ C C NC C CC C C + S C + SC C CCC+
Sbjct: 613 CCCCKYCTCYC*HC-NCCC----CCRSICHWQHRCCSTSGCCSDRSCDTCCCCCCCILLL 449
Query: 356 IKCCHLPKFSTCFGSSCC-------TKCSRPSCCFKCC 386
CHL + CC T C R CC CC
Sbjct: 448 CTLCHLTHIGYPHKNPCCCCNRFRRTLCYRHHCCNTCC 335
Score = 49.7 bits (117), Expect = 2e-06
Identities = 36/126 (28%), Positives = 46/126 (35%), Gaps = 18/126 (14%)
Frame = -2
Query: 300 CRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCC 359
C S++CT +C C C C+C C +C+C CCC C C
Sbjct: 658 CHSFLASHHCTR*RHCCC----CKYCTCY--------C*HCNC------CCC---CRSIC 542
Query: 360 HLPKFSTCFGSSCCTKCSRPSCCFKCC-CNLRCA------------RPSCCC-----MSC 401
H + C S CC+ S +CC CC L C P CCC C
Sbjct: 541 HW-QHRCCSTSGCCSDRSCDTCCCCCCCILLLCTLCHLTHIGYPHKNPCCCCNRFRRTLC 365
Query: 402 FCWRCC 407
+ CC
Sbjct: 364 YRHHCC 347
Score = 45.4 bits (106), Expect = 5e-05
Identities = 29/97 (29%), Positives = 37/97 (37%), Gaps = 5/97 (5%)
Frame = -2
Query: 294 CFSCSCCRMKC-----CSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQ 348
C C CCR C C + CS+ SC + CC C CI C C + K
Sbjct: 574 CNCCCCCRSICHWQHRCCSTSGCCSDRSC-DTCCCCCCCILLLCTLCHLTHIGYPHK-NP 401
Query: 349 CCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKC 385
CC CC+ + + C+ CC C S C
Sbjct: 400 CC-------CCNRFRRTLCYRHHCCNTCCWSSQTRSC 311
>TC215490 weakly similar to UP|Q8I7C3 (Q8I7C3) Lasp protein, partial (7%)
Length = 715
Score = 50.4 bits (119), Expect = 1e-06
Identities = 29/69 (42%), Positives = 32/69 (46%), Gaps = 3/69 (4%)
Frame = -1
Query: 294 CFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCINPECFNCSCINCSCSLKCTQCCCLP 353
C S C CCSN C GC +CSC C NC C C +CSC C+ CCC
Sbjct: 499 CGSVCWCWCWCCSN*CCGC-DCSCWC--CSNC---------CCCCDCSCWC-CSNCCCNK 359
Query: 354 ---SCIKCC 359
SC CC
Sbjct: 358 YPGSCTCCC 332
Score = 48.9 bits (115), Expect = 4e-06
Identities = 27/86 (31%), Positives = 34/86 (39%), Gaps = 1/86 (1%)
Frame = -1
Query: 315 CSCINPEC-CNCSCINPECFNCSCINCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCC 373
C C C C C C + C C C +C C C+ CCC CC +C+
Sbjct: 508 CCCCGSVCWCWCWCCSN*CCGCDC-SCWC---CSNCCC------CCD----CSCW----- 386
Query: 374 TKCSRPSCCFKCCCNLRCARPSCCCM 399
CC CCCN +CCC+
Sbjct: 385 -------CCSNCCCNKYPGSCTCCCI 329
Score = 45.4 bits (106), Expect = 5e-05
Identities = 24/64 (37%), Positives = 26/64 (40%), Gaps = 6/64 (9%)
Frame = -1
Query: 356 IKCCHLPKFSTCFGSSC---CTKCSRPSCCFKCCCNLRCARPSCCCMSCFCW---RCCVG 409
IKC C GS C C CS C C C+ C CCC C CW CC
Sbjct: 535 IKCAP*INCCCCCGSVCWCWCWCCSN*CC--GCDCSCWCCSNCCCCCDCSCWCCSNCCCN 362
Query: 410 KHSG 413
K+ G
Sbjct: 361 KYPG 350
Score = 40.8 bits (94), Expect = 0.001
Identities = 26/76 (34%), Positives = 33/76 (43%), Gaps = 5/76 (6%)
Frame = -1
Query: 338 INCSCSLKCTQCCCLPSCIKCCHLPKFSTCFGSSCCTKCSRPSCCFKCCCNLRC-ARPSC 396
I C+ + C CCC S C + C G C C +CC CCC+ C +C
Sbjct: 535 IKCAP*INC--CCCCGSVCWCWCWCCSN*CCGCDCSCWCC-SNCC--CCCDCSCWCCSNC 371
Query: 397 CCM----SCFCWRCCV 408
CC SC C CC+
Sbjct: 370 CCNKYPGSCTC--CCI 329
Score = 34.7 bits (78), Expect = 0.083
Identities = 20/52 (38%), Positives = 24/52 (45%), Gaps = 7/52 (13%)
Frame = -1
Query: 294 CFSCS--CCRMKC---CSNNCTGCSNCSC--INPECCNCSCINPECFNCSCI 338
C+ CS CC C C +NC C +CSC + CCN P C CI
Sbjct: 475 CWCCSN*CCGCDCSCWCCSNCCCCCDCSCWCCSNCCCN---KYPGSCTCCCI 329
Score = 30.8 bits (68), Expect = 1.2
Identities = 15/37 (40%), Positives = 17/37 (45%)
Frame = -1
Query: 292 SSCFSCSCCRMKCCSNNCTGCSNCSCINPECCNCSCI 328
S+C C C CCSN C C+ P C C CI
Sbjct: 421 SNCCCCCDCSCWCCSNCC--CNK----YPGSCTCCCI 329
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.139 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,738,563
Number of Sequences: 63676
Number of extensions: 926478
Number of successful extensions: 18044
Number of sequences better than 10.0: 530
Number of HSP's better than 10.0 without gapping: 11307
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 15277
length of query: 420
length of database: 12,639,632
effective HSP length: 100
effective length of query: 320
effective length of database: 6,272,032
effective search space: 2007050240
effective search space used: 2007050240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0230.1


