

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0228a.2
(510 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
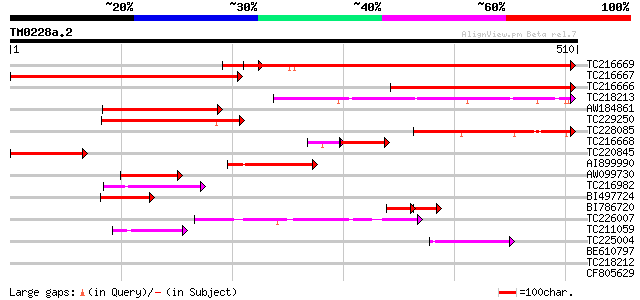
Score E
Sequences producing significant alignments: (bits) Value
TC216669 similar to GB|BAC06968.1|22093674|AP003815 kinase-like ... 401 e-112
TC216667 weakly similar to GB|BAC06968.1|22093674|AP003815 kinas... 310 1e-84
TC216666 similar to GB|BAC06968.1|22093674|AP003815 kinase-like ... 268 6e-72
TC218213 similar to UP|Q9LK27 (Q9LK27) Gb|AAF01563.1, partial (21%) 134 1e-31
AW184861 114 1e-25
TC229250 similar to UP|Q9AR00 (Q9AR00) PSTVd RNA-biding protein,... 103 2e-22
TC228085 similar to UP|Q9LK27 (Q9LK27) Gb|AAF01563.1, partial (17%) 100 1e-21
TC216668 79 8e-19
TC220845 82 7e-16
AI899990 78 8e-15
AW099730 weakly similar to GP|29569106|gb| global transcription ... 68 9e-12
TC216982 similar to UP|Q9AR19 (Q9AR19) Histone acetyltransferase... 67 2e-11
BI497724 similar to GP|16323117|gb| AT3g01770/F28J7_10 {Arabidop... 64 2e-10
BI786720 36 4e-05
TC226007 similar to UP|Q8LC13 (Q8LC13) Remorin, partial (65%) 44 1e-04
TC211059 similar to UP|Q8LRK9 (Q8LRK9) HAF1, partial (7%) 43 4e-04
TC225004 42 7e-04
BE610797 39 0.004
TC218212 similar to UP|Q9LK27 (Q9LK27) Gb|AAF01563.1, partial (11%) 38 0.009
CF805629 31 0.010
>TC216669 similar to GB|BAC06968.1|22093674|AP003815 kinase-like protein
{Oryza sativa (japonica cultivar-group);} , partial
(16%)
Length = 1336
Score = 401 bits (1031), Expect = e-112
Identities = 226/354 (63%), Positives = 257/354 (71%), Gaps = 55/354 (15%)
Frame = +1
Query: 211 PRQKDTFPKKSQVSEHKRVLKISSLAARDAKVEVPKLSQV-------------------- 250
P K PKK+Q+ EHK + KI SLA +DA+VE PKLSQ+
Sbjct: 67 PGIKIPCPKKTQLFEHKGMHKIRSLATKDARVETPKLSQIPCKLIEKDLHKGSKGNHDVE 246
Query: 251 ---------------------------HCKS------IEKDLI-KGSEERDRIAPGAVAF 276
HC++ + D+ +GSE RD IA GA +
Sbjct: 247 QPAGSLKSYFSVRFVTSKCRICGNIICHCENPSNSTQVSSDISSEGSEGRDLIARGADSL 426
Query: 277 KQKRQIKSTSPLERKSDPDSDGAVSSLDSEHVYPGSQHATPATDASSGEVWSTPDFPVQL 336
+Q + K TSP+E+KSD DSDGAVSSLDSEHV P SQH TP TDASSGEVWS P PVQL
Sbjct: 427 RQDCRTKCTSPMEKKSDLDSDGAVSSLDSEHVCPSSQHVTPTTDASSGEVWSMPVLPVQL 606
Query: 337 SPKKALRAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQ 396
SPKKALRAA+LKSRFADTILKAQQKTLL+HGDKG+P KMQ EKERLERIQREERARIEAQ
Sbjct: 607 SPKKALRAAILKSRFADTILKAQQKTLLDHGDKGNPQKMQQEKERLERIQREERARIEAQ 786
Query: 397 IKTAEAAARARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELETLSGCTLSY 456
IKTAEAAAR RAEEE RQ+REKEREA+RAAIE+MKRTV+IEHN+EI+KELE+LSGCTLSY
Sbjct: 787 IKTAEAAARMRAEEESRQRREKEREASRAAIEKMKRTVDIEHNMEIIKELESLSGCTLSY 966
Query: 457 KALGGRNDYKVALETLDKLQFENPLEQLGLFIKDEL-ADEDEEVINGTVEEGEI 509
KA+GGRN YKVALETLDK QFENPLE+LGLFIK++ D DEEV+NG EEGEI
Sbjct: 967 KAVGGRNGYKVALETLDKHQFENPLERLGLFIKEDFTTDLDEEVLNGGFEEGEI 1128
Score = 41.6 bits (96), Expect = 9e-04
Identities = 20/37 (54%), Positives = 25/37 (67%)
Frame = +3
Query: 192 KSVTGTIKETARKSCNVMHPRQKDTFPKKSQVSEHKR 228
KS T TIKET RKS + M PR KD+ PKK+ + +R
Sbjct: 9 KSETETIKETGRKSLDAMLPRHKDSLPKKNTIIRTQR 119
>TC216667 weakly similar to GB|BAC06968.1|22093674|AP003815 kinase-like
protein {Oryza sativa (japonica cultivar-group);} ,
partial (21%)
Length = 892
Score = 310 bits (793), Expect = 1e-84
Identities = 151/209 (72%), Positives = 172/209 (82%)
Frame = +3
Query: 1 MIATETIVPTRKLKIKFSTKRIEVDPGQKCEFGQRGLQTDENGRCNSAEKLFKPDSIKRG 60
MIATETIVP+ KLKIKFSTKRIE+D G KCEFGQ+ D+N RCNS E +S KRG
Sbjct: 252 MIATETIVPSTKLKIKFSTKRIEIDSGPKCEFGQKVYHNDKNRRCNSNENSSALNSNKRG 431
Query: 61 PPEGFEGQKEKRQKIDRKGSTQCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKP 120
PP EGQKEKRQKIDRKGS QCA ILK LMS+ YSWVF+KPVDP+AL+IPDYF II+ P
Sbjct: 432 PPVSVEGQKEKRQKIDRKGSMQCATILKSLMSHTYSWVFSKPVDPIALSIPDYFTIISHP 611
Query: 121 MDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDM 180
MDLGTI SKL+KN+YS EEFAADVRLTFSNAM YNPP NDVH+MAKEL++IFD KWKD+
Sbjct: 612 MDLGTIKSKLEKNIYSGTEEFAADVRLTFSNAMKYNPPSNDVHLMAKELSKIFDRKWKDL 791
Query: 181 DKKWKCEDELGKSVTGTIKETARKSCNVM 209
+KWKCEDE KS + TIKET RKS +++
Sbjct: 792 GRKWKCEDEHDKSESETIKETGRKSLDML 878
>TC216666 similar to GB|BAC06968.1|22093674|AP003815 kinase-like protein
{Oryza sativa (japonica cultivar-group);} , partial
(14%)
Length = 667
Score = 268 bits (684), Expect = 6e-72
Identities = 141/168 (83%), Positives = 155/168 (91%), Gaps = 1/168 (0%)
Frame = +2
Query: 343 RAAMLKSRFADTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEA 402
RAAMLKSRFADTILKAQ KTLL+HGDKG+P +MQ EKERLERIQREERARIEAQIK AEA
Sbjct: 2 RAAMLKSRFADTILKAQLKTLLDHGDKGNPPEMQQEKERLERIQREERARIEAQIKIAEA 181
Query: 403 AARARAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELETLSGCTLSYKALGGR 462
AAR RAEEE RQ+REKEREAARAAIE+MKRTV+IEHN+EI+KELE+LSGCTLSYKA+G R
Sbjct: 182 AARMRAEEESRQRREKEREAARAAIEKMKRTVDIEHNVEIIKELESLSGCTLSYKAVGCR 361
Query: 463 NDYKVALETLDKLQFENPLEQLGLFIKDEL-ADEDEEVINGTVEEGEI 509
N YKVALETLDK QFENPLE+LGLFIK++ ADEDEEV+NG EEGEI
Sbjct: 362 NGYKVALETLDKPQFENPLERLGLFIKEDFTADEDEEVLNGGFEEGEI 505
>TC218213 similar to UP|Q9LK27 (Q9LK27) Gb|AAF01563.1, partial (21%)
Length = 1177
Score = 134 bits (336), Expect = 1e-31
Identities = 109/292 (37%), Positives = 155/292 (52%), Gaps = 20/292 (6%)
Frame = +1
Query: 238 RDAKVEVPK-LSQVHCKSIEKDLIKGSEERDRIAPGAVAFKQKRQIKSTSPLERKSDP-- 294
+D V K LS + D SE D A A K + S + L+ +
Sbjct: 157 KDTTYRVNKSLSPGSSNDTDSDSSSDSEADDVKASPANVAKAPENLGSEAQLDEMTMAAA 336
Query: 295 --DSDGAVSSLDSEHVYPGSQHATPATDASSGEVWSTPDFPVQLSPKKALRAAMLKSRFA 352
+ + +VS LD + SQH + D+ + + Q SP K RAA+LK RF
Sbjct: 337 TLERNQSVSGLDQ--LEDNSQHKPSSFDSDCQQDGDSAATERQFSPDKLYRAAVLKKRFL 510
Query: 353 DTILKAQQKTLLEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAARARAEE-- 410
DTILKA++KTL + G+KGDP K++ E+E+LE Q++E+AR++A+ K AE A + EE
Sbjct: 511 DTILKAREKTLTQ-GEKGDPEKLRQEREKLEMEQKKEKARLQAEAKAAEDARKQAEEEAA 687
Query: 411 -ELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELETLSGCTLSYKALGGRNDYKVAL 469
E R++RE EREAAR A+ +M++TVEI N I+++LE L + + L D
Sbjct: 688 AEARRKRELEREAARQALLQMEKTVEINENSRILEDLEMLR--AVPAEQLPSSVDETSPA 861
Query: 470 ETLD-----KLQFENPLEQLGLFIKDELADEDEE-----VIN--GTVEEGEI 509
+ D K NPLEQLGL+IK + DE+EE + N VEEGEI
Sbjct: 862 HSQDGLGSFKFGSSNPLEQLGLYIKAD--DEEEEGEPPCIPNPVNDVEEGEI 1011
>AW184861
Length = 448
Score = 114 bits (284), Expect = 1e-25
Identities = 53/108 (49%), Positives = 73/108 (67%)
Frame = +2
Query: 84 AAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAA 143
+++L+ LM + + WVFN PVD L + DYF II PMDLGT+ ++L KN Y +EFA
Sbjct: 2 SSLLEKLMRHKHGWVFNSPVDVETLGLHDYFTIITHPMDLGTVKTRLNKNWYKSPKEFAE 181
Query: 144 DVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKWKCEDELG 191
DVRLTF NAMTYNP DVH+MA+ L++IF+ +W ++ + E G
Sbjct: 182 DVRLTFRNAMTYNPQGQDVHIMAELLSKIFEDRWAIIESDYNREMRYG 325
>TC229250 similar to UP|Q9AR00 (Q9AR00) PSTVd RNA-biding protein, Virp1
(PSTVd RNA-binding protein Virp1a) (PSTVd RNA-binding
protein Virp1b) (PSTVd RNA-binding protein Virp1c)
(PSTVd RNA-binding protein Virp1d), partial (34%)
Length = 1450
Score = 103 bits (257), Expect = 2e-22
Identities = 55/138 (39%), Positives = 85/138 (60%), Gaps = 9/138 (6%)
Frame = +2
Query: 83 CAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFA 142
C+ +L+ L+ + + WVF PVD V L + DY II +PMDLGT+ S L KNVY+ +FA
Sbjct: 668 CSQVLQKLIKHKHGWVFKAPVDVVGLKLHDYCDIIKQPMDLGTVKSNLSKNVYATPADFA 847
Query: 143 ADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKDMDKKW--------KCEDELGKSV 194
+DVRLTF+NA+ YNP +DV+ MA++L F+ ++ + +K+ + E+EL S
Sbjct: 848 SDVRLTFNNALAYNPKGHDVYTMAEQLLARFEELYRPVHEKFEGSIVHDRESEEELQASS 1027
Query: 195 TGTIK-ETARKSCNVMHP 211
++ E +K N + P
Sbjct: 1028WSQVEPERVKKKENPIPP 1081
>TC228085 similar to UP|Q9LK27 (Q9LK27) Gb|AAF01563.1, partial (17%)
Length = 947
Score = 100 bits (250), Expect = 1e-21
Identities = 68/161 (42%), Positives = 100/161 (61%), Gaps = 15/161 (9%)
Frame = +1
Query: 364 LEHGDKGDPLKMQLEKERLERIQREERARIEAQIKTAEAAAR---ARAEEELRQQREKER 420
LE +K DP K+++E+E LER Q+EE+AR++A+ K AE A R A A E +++RE ER
Sbjct: 1 LEKDEKRDPEKLRIEREDLERRQKEEKARLQAEAKAAEEAQRKAEAEAAAEAKRKRELER 180
Query: 421 EAARAAIEEMKRTVEIEHNLEIVKELETLSGC----TLSYKALGGRNDYKVALETLDKLQ 476
EAAR A+++M++TV+I N +++LE LS S+K + + L + KLQ
Sbjct: 181 EAARQALQKMEKTVDINENSHFLEDLEMLSAVHDEHLPSFKEETSADQPQDGLGGI-KLQ 357
Query: 477 FENPLEQLGLFIKDELADEDEEV--------INGTVEEGEI 509
NPLEQLGL++K+E +E+EE + VEEGEI
Sbjct: 358 -GNPLEQLGLYMKEEEEEEEEEEPPPSGAAGPSNDVEEGEI 477
>TC216668
Length = 435
Score = 79.3 bits (194), Expect(2) = 8e-19
Identities = 38/44 (86%), Positives = 38/44 (86%)
Frame = +1
Query: 298 GAVSSLDSEHVYPGSQHATPATDASSGEVWSTPDFPVQLSPKKA 341
GAVSSLDSEHV P SQH TP TDASSGEVWSTP PVQLSPKKA
Sbjct: 304 GAVSSLDSEHVCPSSQHVTPTTDASSGEVWSTPVLPVQLSPKKA 435
Score = 32.7 bits (73), Expect(2) = 8e-19
Identities = 19/41 (46%), Positives = 24/41 (58%), Gaps = 8/41 (19%)
Frame = +3
Query: 269 IAPGAVAFKQKR--------QIKSTSPLERKSDPDSDGAVS 301
+A AVA+K K+ K TSP+E+KSD DSDG S
Sbjct: 102 LAKLAVAYKSKKLYVYFQNCPTKCTSPIEKKSDSDSDGEKS 224
>TC220845
Length = 476
Score = 81.6 bits (200), Expect = 7e-16
Identities = 44/70 (62%), Positives = 49/70 (69%)
Frame = +3
Query: 1 MIATETIVPTRKLKIKFSTKRIEVDPGQKCEFGQRGLQTDENGRCNSAEKLFKPDSIKRG 60
MIATETIVP+ KLKIKFSTKR+EV+ G K EFGQ+ TDEN N K +S KRG
Sbjct: 267 MIATETIVPSTKLKIKFSTKRMEVESGPKYEFGQKVSHTDENRSFNLNGKSSALNSNKRG 446
Query: 61 PPEGFEGQKE 70
PP EGQKE
Sbjct: 447 PPTSIEGQKE 476
>AI899990
Length = 382
Score = 78.2 bits (191), Expect = 8e-15
Identities = 42/81 (51%), Positives = 57/81 (69%)
Frame = +2
Query: 197 TIKETARKSCNVMHPRQKDTFPKKSQVSEHKRVLKISSLAARDAKVEVPKLSQVHCKSIE 256
TIKET RKS +++ R +D+ PKK+QVSEHK + KI SLA +DA+VE KLSQ+ CK IE
Sbjct: 11 TIKETGRKSLDMLS-RHRDSLPKKTQVSEHKGMPKIRSLATKDARVEPSKLSQIPCKLIE 187
Query: 257 KDLIKGSEERDRIAPGAVAFK 277
KD+ KG++ + A + K
Sbjct: 188 KDMHKGNKGNHDVEQPAGSLK 250
>AW099730 weakly similar to GP|29569106|gb| global transcription factor group
E {Zea mays}, partial (8%)
Length = 177
Score = 68.2 bits (165), Expect = 9e-12
Identities = 30/56 (53%), Positives = 40/56 (70%)
Frame = +2
Query: 100 NKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNAMTY 155
N PVD V L + DY+ +I +PMDLGT+ S L N Y+ +FA+DVRLTF+NA+ Y
Sbjct: 8 NAPVDVVGLQLTDYYDVIKQPMDLGTVKSNLSMNKYTTPSDFASDVRLTFNNALAY 175
>TC216982 similar to UP|Q9AR19 (Q9AR19) Histone acetyltransferase GCN5
(At3g54610) (Expressed protein), partial (53%)
Length = 1203
Score = 66.6 bits (161), Expect = 2e-11
Identities = 36/93 (38%), Positives = 52/93 (55%), Gaps = 1/93 (1%)
Frame = +1
Query: 85 AILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKL-KKNVYSCVEEFAA 143
++LK + + +W F +PVD A ++PDY+ II PMDL T+S ++ + Y E F A
Sbjct: 595 SLLKSMFDHADAWPFKEPVD--ARDVPDYYDIIKDPMDLKTMSKRVDSEQYYVTFEMFVA 768
Query: 144 DVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSK 176
D R F+NA TYN P + + L F SK
Sbjct: 769 DARRMFANARTYNSPETIYYKCSTRLEAHFQSK 867
>BI497724 similar to GP|16323117|gb| AT3g01770/F28J7_10 {Arabidopsis
thaliana}, partial (10%)
Length = 408
Score = 63.9 bits (154), Expect = 2e-10
Identities = 30/49 (61%), Positives = 36/49 (73%)
Frame = +1
Query: 82 QCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKL 130
QC +LK +MS+ + VF+KPVD V NIPDYF II PMDLGT+ SKL
Sbjct: 232 QCETLLKRVMSHQFGKVFDKPVDIVKWNIPDYFTIIKHPMDLGTVKSKL 378
>BI786720
Length = 421
Score = 36.2 bits (82), Expect(2) = 4e-05
Identities = 17/26 (65%), Positives = 21/26 (80%)
Frame = +3
Query: 340 KALRAAMLKSRFADTILKAQQKTLLE 365
K RAA+LK RF DTILKA++KTL +
Sbjct: 3 KLYRAAVLKKRFLDTILKAREKTLTQ 80
Score = 29.3 bits (64), Expect(2) = 4e-05
Identities = 12/25 (48%), Positives = 19/25 (76%)
Frame = +2
Query: 364 LEHGDKGDPLKMQLEKERLERIQRE 388
L G+KGDP K++ E+E+LE Q++
Sbjct: 176 LGQGEKGDPEKLRQEREKLEMEQKK 250
>TC226007 similar to UP|Q8LC13 (Q8LC13) Remorin, partial (65%)
Length = 951
Score = 44.3 bits (103), Expect = 1e-04
Identities = 55/210 (26%), Positives = 88/210 (41%), Gaps = 5/210 (2%)
Frame = +1
Query: 167 KELNQIFD-SKWKDMDKKWKCEDELGKSVTGTIKETARKSCNVMHPRQKDTFPKKSQVSE 225
+ L+Q+F + WKD+ K K +E K V T + +P + K V+E
Sbjct: 163 RTLSQLFPITLWKDIHSKGKMTEEQSKKVAETESFPS-------NPAPEPVVVPKEDVAE 321
Query: 226 HKRVLKISSLAARD---AKVEVPKLSQVHCKSIEKDLIKGSEERDRIAPGAVAFKQKRQI 282
K V+ S + D A V V K S+V E+ I+GS RD + K+ I
Sbjct: 322 EKSVIPQPSPSPADESKALVIVEKTSEV----AEEKPIEGSVNRDAVLARVATEKRLSLI 489
Query: 283 KSTSPLER-KSDPDSDGAVSSLDSEHVYPGSQHATPATDASSGEVWSTPDFPVQLSPKKA 341
K+ E+ K+D S +S++ + + S+ A + E QL KKA
Sbjct: 490 KAWEESEKSKADNKSHKKLSAISA---WENSKKAAAEAELRKIEE--------QLEKKKA 636
Query: 342 LRAAMLKSRFADTILKAQQKTLLEHGDKGD 371
LK++ A +A++K KG+
Sbjct: 637 EYGEKLKNKIATIHREAEEKRAFIEAQKGE 726
>TC211059 similar to UP|Q8LRK9 (Q8LRK9) HAF1, partial (7%)
Length = 800
Score = 42.7 bits (99), Expect = 4e-04
Identities = 25/68 (36%), Positives = 33/68 (47%)
Frame = +2
Query: 93 YPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYSCVEEFAADVRLTFSNA 152
Y S++F KPV PDY II +PMDL I +++ Y E+F D+ NA
Sbjct: 272 YDLSYLFLKPVSKK--EAPDYLDIIERPMDLSRIRERVRNMEYKSREDFRHDMWQITFNA 445
Query: 153 MTYNPPHN 160
YN N
Sbjct: 446 HKYNDGRN 469
>TC225004
Length = 734
Score = 42.0 bits (97), Expect = 7e-04
Identities = 29/77 (37%), Positives = 37/77 (47%)
Frame = -3
Query: 378 EKERLERIQREERARIEAQIKTAEAAARARAEEELRQQREKEREAARAAIEEMKRTVEIE 437
E+ER ER + ER R + + E E E ++RE+ERE R E +R E E
Sbjct: 465 ERER-ERERERERERERERERERERERERERERERERERERERERERERERERERERERE 289
Query: 438 HNLEIVKELETLSGCTL 454
EIV E ET C L
Sbjct: 288 REREIVIERETDRKCVL 238
Score = 28.5 bits (62), Expect = 7.5
Identities = 17/41 (41%), Positives = 22/41 (53%)
Frame = -3
Query: 407 RAEEELRQQREKEREAARAAIEEMKRTVEIEHNLEIVKELE 447
RAE R++RE+ERE R E +R E E E +E E
Sbjct: 555 RAEFGTRRERERERERERERERERERERERERERERERERE 433
>BE610797
Length = 391
Score = 39.3 bits (90), Expect = 0.004
Identities = 25/101 (24%), Positives = 51/101 (49%), Gaps = 1/101 (0%)
Frame = +1
Query: 80 STQCAAILKCLMSYPYSWVFNKPVDPVALNIPDYFAIIAKPMDLGTISSKLKKNVYS-CV 138
S +L+ + + +S +F + +D + Y ++ +PMDL TI +L+K YS C
Sbjct: 49 SEPLVGVLELIKGHEHSSLFERRLDSQQ-DTDRYKDLVKQPMDLETIQLRLQKGHYSSCT 225
Query: 139 EEFAADVRLTFSNAMTYNPPHNDVHMMAKELNQIFDSKWKD 179
F D+ L F+NA + + ++L+++ ++ K+
Sbjct: 226 SAFFRDLLLLFTNATVFFSHDSLESQAGRQLHRLATAEMKN 348
>TC218212 similar to UP|Q9LK27 (Q9LK27) Gb|AAF01563.1, partial (11%)
Length = 712
Score = 38.1 bits (87), Expect = 0.009
Identities = 36/102 (35%), Positives = 49/102 (47%), Gaps = 12/102 (11%)
Frame = +2
Query: 420 REAARAAIEEMKRTVEIEHNLEIVKELETLSGCTLSYKALGGRNDYKVALETLD-----K 474
REAA A+ + +TVEI N I+++ + + L D + D K
Sbjct: 8 REAAXXALLQXXKTVEINENSRILEDXXLXRA--VPTEQLPSSVDETSPAHSQDGLGSFK 181
Query: 475 LQFENPLEQLGLFIKDELADEDEE-----VIN--GTVEEGEI 509
NPLEQLGL+IK + DE+EE + N VEEGEI
Sbjct: 182 FGSSNPLEQLGLYIKAD--DEEEEGEPPCIPNPVNDVEEGEI 301
>CF805629
Length = 556
Score = 31.2 bits (69), Expect(2) = 0.010
Identities = 14/22 (63%), Positives = 14/22 (63%)
Frame = +1
Query: 135 YSCVEEFAADVRLTFSNAMTYN 156
Y V E ADVRL F NAM YN
Sbjct: 52 YKHVREICADVRLVFKNAMKYN 117
Score = 25.8 bits (55), Expect(2) = 0.010
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = +2
Query: 160 NDVHMMAKELNQIFDSKW 177
+DVH+MAK L F+ KW
Sbjct: 128 SDVHVMAKTLLSKFEEKW 181
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.130 0.364
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,583,160
Number of Sequences: 63676
Number of extensions: 224955
Number of successful extensions: 1194
Number of sequences better than 10.0: 97
Number of HSP's better than 10.0 without gapping: 1151
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1180
length of query: 510
length of database: 12,639,632
effective HSP length: 101
effective length of query: 409
effective length of database: 6,208,356
effective search space: 2539217604
effective search space used: 2539217604
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0228a.2


