

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0227.11
(425 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
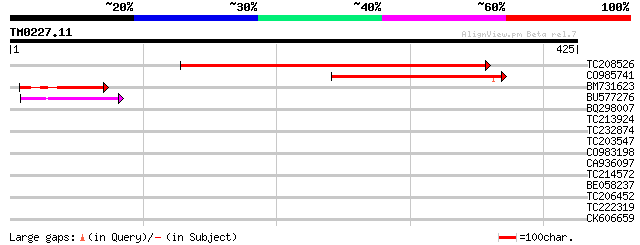
Score E
Sequences producing significant alignments: (bits) Value
TC208526 similar to UP|Q6YW97 (Q6YW97) MAP3K protein kinase-like... 328 3e-90
CO985741 193 1e-49
BM731623 weakly similar to PIR|D85436|D854 MAP3K-like protein ki... 61 1e-09
BU577276 50 2e-06
BQ298007 weakly similar to PIR|T47442|T47 disease resistance pro... 39 0.006
TC213924 33 0.32
TC232874 weakly similar to UP|O04020 (O04020) F7G19.3 protein, p... 32 0.42
TC203547 weakly similar to UP|Q75IX4 (Q75IX4) Expressed protein,... 32 0.42
CO983198 32 0.42
CA936097 31 1.2
TC214572 homologue to UP|TBA3_ARATH (P20363) Tubulin alpha-3/alp... 30 2.7
BE058237 28 6.0
TC206452 similar to UP|CBPY_YEAST (P00729) Carboxypeptidase Y pr... 28 6.0
TC222319 UP|Q39833 (Q39833) Alfa-carboxyltransferase precursor (... 28 7.8
CK606659 28 7.8
>TC208526 similar to UP|Q6YW97 (Q6YW97) MAP3K protein kinase-like protein,
partial (59%)
Length = 1214
Score = 328 bits (841), Expect = 3e-90
Identities = 149/232 (64%), Positives = 184/232 (79%)
Frame = +2
Query: 129 LDLYDFENDVWLCHSFGGQCYNYTAFQPAINVLKEIQVFLEANPSEIVTIIIEDYVTSPK 188
LD+YDF+ND+WLCHSF GQCYN+TAFQPA+N LKE++ FL NP+EIVTIIIEDYV +PK
Sbjct: 23 LDMYDFQNDIWLCHSFRGQCYNFTAFQPAVNTLKEVEAFLTENPTEIVTIIIEDYVHTPK 202
Query: 189 GLTKVFDAAGLRKYWFPVSRMPKNGGDWPKVDDMVQKNQRLVVFTSKASKEASEGIAYEW 248
GLT VF +AGL KYWFPVS+MPK G DWP V +MVQ N RLVVFTS ASKEA EGIAY+W
Sbjct: 203 GLTNVFTSAGLDKYWFPVSKMPKKGDDWPTVTEMVQANHRLVVFTSDASKEAGEGIAYQW 382
Query: 249 RYLVENQYGNGGMKAGSCPNRAESPSMNTTSRSLVLVNFFRDLPDVAQSCKDNSAPLLSM 308
+++VEN+ G+ G++ GSCP+R ES ++N+ S SL L+N+F P A SCK++SAPL M
Sbjct: 383 KHMVENESGDPGVQQGSCPHRKESKALNSKSHSLFLMNYFPTYPVEADSCKEHSAPLAEM 562
Query: 309 VSTCNQAAGKRWPNFIAVDFYKRSDGGGAPEAVDVANGHLVCGCGNIATCKV 360
V+TC +AAG PNFIAV+FY RSDGGG + VD NGH +CGC + C+V
Sbjct: 563 VNTCYKAAGNLMPNFIAVNFYMRSDGGGVFDIVDKMNGHTLCGCSTVTACQV 718
>CO985741
Length = 761
Score = 193 bits (490), Expect = 1e-49
Identities = 92/138 (66%), Positives = 108/138 (77%), Gaps = 7/138 (5%)
Frame = -1
Query: 242 EGIAYEWRYLVENQYGNGGMKAGSCPNRAESPSMNTTSRSLVLVNFFRDLPDVAQSCKDN 301
EGIAY+W Y+VENQYG+ GMKAGSCP+RAESP+MNT SRSLVLVN+F P+ +Q+C DN
Sbjct: 761 EGIAYQWTYVVENQYGDDGMKAGSCPSRAESPAMNTKSRSLVLVNYFHSAPNRSQACADN 582
Query: 302 SAPLLSMVSTCNQAAGKRWPNFIAVDFYKRSDGGGAPEAVDVANGHLVCGCGNIATCKVR 361
SAPLL M+ TC++AAG RW NFIAVD+Y+RSDGGGAP AVD ANGHL CGC NIA CK
Sbjct: 581 SAPLLDMMKTCHEAAGNRWANFIAVDYYQRSDGGGAPLAVDEANGHLTCGCDNIAYCKEN 402
Query: 362 -------LTPITFIPPHT 372
+ PI+ PP T
Sbjct: 401 AKSGTCDVPPISPPPPAT 348
>BM731623 weakly similar to PIR|D85436|D854 MAP3K-like protein kinase
[imported] - Arabidopsis thaliana, partial (4%)
Length = 422
Score = 60.8 bits (146), Expect = 1e-09
Identities = 32/67 (47%), Positives = 41/67 (60%)
Frame = +2
Query: 8 LAIATTLFTILVLLDSSLALKFKPLQQEGQICVANKNCNSGLHCETCVTNGNVRPRCTRT 67
L IA LFT SS A K G+ C ++ C++GL C+TC NGN RPRC+RT
Sbjct: 257 LLIAVCLFT------SSSASKI------GENCGSDNKCDAGLSCQTCPANGNTRPRCSRT 400
Query: 68 QPINPTS 74
QP++PTS
Sbjct: 401 QPLSPTS 421
>BU577276
Length = 428
Score = 50.1 bits (118), Expect = 2e-06
Identities = 31/77 (40%), Positives = 40/77 (51%)
Frame = +2
Query: 9 AIATTLFTILVLLDSSLALKFKPLQQEGQICVANKNCNSGLHCETCVTNGNVRPRCTRTQ 68
A A + +LV L SL+ Q + C A +C GL C C + G +P CTR Q
Sbjct: 200 APANAIIFLLVPLLCSLSF-INVNSQILEACSAATDCGPGLFCGNCPSLGLKQPICTRGQ 376
Query: 69 PINPTSKVKGLPFNRYS 85
PTS V GLPFN+Y+
Sbjct: 377 VTLPTSIVNGLPFNKYT 427
>BQ298007 weakly similar to PIR|T47442|T47 disease resistance protein homlog
- Arabidopsis thaliana, partial (2%)
Length = 367
Score = 38.5 bits (88), Expect = 0.006
Identities = 19/43 (44%), Positives = 22/43 (50%)
Frame = +1
Query: 37 QICVANKNCNSGLHCETCVTNGNVRPRCTRTQPINPTSKVKGL 79
+ C A +C GL C C G +P CTR Q PTS V GL
Sbjct: 238 EACSAATDCGPGLFCGNCPALGLKQPICTRGQATLPTSIVNGL 366
>TC213924
Length = 380
Score = 32.7 bits (73), Expect = 0.32
Identities = 12/23 (52%), Positives = 17/23 (73%)
Frame = +1
Query: 336 GAPEAVDVANGHLVCGCGNIATC 358
G+ +AVD NG L+CGC ++ TC
Sbjct: 7 GSFQAVDTLNGKLLCGCDDVHTC 75
>TC232874 weakly similar to UP|O04020 (O04020) F7G19.3 protein, partial (32%)
Length = 715
Score = 32.3 bits (72), Expect = 0.42
Identities = 17/47 (36%), Positives = 22/47 (46%)
Frame = +1
Query: 333 DGGGAPEAVDVANGHLVCGCGNIATCKVRLTPITFIPPHTHLNLHVL 379
DGGG + NGH +C +AT V T +PP H H+L
Sbjct: 370 DGGGQWDRHGGVNGHCICTGSQVATASV--TGAAGVPPSCHWLQHIL 504
>TC203547 weakly similar to UP|Q75IX4 (Q75IX4) Expressed protein, partial
(6%)
Length = 302
Score = 32.3 bits (72), Expect = 0.42
Identities = 14/26 (53%), Positives = 18/26 (68%)
Frame = +3
Query: 292 PDVAQSCKDNSAPLLSMVSTCNQAAG 317
P A SCK++ PL MV+TC +AAG
Sbjct: 27 PVEADSCKEHLVPLADMVNTCYKAAG 104
>CO983198
Length = 736
Score = 32.3 bits (72), Expect = 0.42
Identities = 14/26 (53%), Positives = 18/26 (68%)
Frame = -1
Query: 292 PDVAQSCKDNSAPLLSMVSTCNQAAG 317
P A SCK++ PL MV+TC +AAG
Sbjct: 178 PVEADSCKEHLVPLADMVNTCYKAAG 101
>CA936097
Length = 426
Score = 30.8 bits (68), Expect = 1.2
Identities = 14/43 (32%), Positives = 24/43 (55%)
Frame = +2
Query: 120 LNNGVRGLMLDLYDFENDVWLCHSFGGQCYNYTAFQPAINVLK 162
L NGVR + +DL + CHSF C+ ++ ++ +N L+
Sbjct: 104 LRNGVRRVTVDLLQVMANY*TCHSFFFLCWTFSLYR*PLNTLQ 232
>TC214572 homologue to UP|TBA3_ARATH (P20363) Tubulin alpha-3/alpha-5 chain,
partial (57%)
Length = 961
Score = 29.6 bits (65), Expect = 2.7
Identities = 14/41 (34%), Positives = 24/41 (58%), Gaps = 5/41 (12%)
Frame = +1
Query: 368 IPPHTHLNLHVLN-----FDLCRKTWDLEHVNYLKLKQLLN 403
IP H H ++ VL +D+CR++ D+E Y L +L++
Sbjct: 1 IPLHEHTDVVVLLDNEAIYDICRRSLDIERPTYTNLNRLIS 123
>BE058237
Length = 413
Score = 28.5 bits (62), Expect = 6.0
Identities = 13/31 (41%), Positives = 20/31 (63%)
Frame = -2
Query: 300 DNSAPLLSMVSTCNQAAGKRWPNFIAVDFYK 330
+NS L+ M+ TC+ A N++AVD+YK
Sbjct: 376 NNSGGLIDMLQTCHGA------NYVAVDYYK 302
>TC206452 similar to UP|CBPY_YEAST (P00729) Carboxypeptidase Y precursor
(Carboxypeptidase YSCY) , partial (5%)
Length = 1330
Score = 28.5 bits (62), Expect = 6.0
Identities = 15/44 (34%), Positives = 21/44 (47%)
Frame = +1
Query: 45 CNSGLHCETCVTNGNVRPRCTRTQPINPTSKVKGLPFNRYSWLT 88
C L T V GN+ P TQP N T+ ++ RY W++
Sbjct: 412 CMQALRVAT-VEQGNIDPYSVYTQPCNNTASLRRGLKGRYPWMS 540
>TC222319 UP|Q39833 (Q39833) Alfa-carboxyltransferase precursor (Carboxyl
transferase alpha subunit) , complete
Length = 2588
Score = 28.1 bits (61), Expect = 7.8
Identities = 22/59 (37%), Positives = 28/59 (47%), Gaps = 8/59 (13%)
Frame = -3
Query: 263 AGSCPNRAESPSMNTTSRSLVL------VNFFRDLP--DVAQSCKDNSAPLLSMVSTCN 313
+ SC N S S+ + RSL L VN F LP A+SC NS S++S N
Sbjct: 2040 SNSCLNVTASLSLFASDRSLTLHNSLR*VNLFCSLP*LGAARSCCSNSTLSFSILSFSN 1864
>CK606659
Length = 443
Score = 28.1 bits (61), Expect = 7.8
Identities = 16/37 (43%), Positives = 19/37 (51%)
Frame = -1
Query: 65 TRTQPINPTSKVKGLPFNRYSWLTTHNSFALLGQKSM 101
TR INP S V +PF W TT S +G+K M
Sbjct: 311 TRIWRINPRSTVMYVPFVWIMWGTTAASMYAMGRKVM 201
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.136 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,054,815
Number of Sequences: 63676
Number of extensions: 318351
Number of successful extensions: 1572
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 1564
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1572
length of query: 425
length of database: 12,639,632
effective HSP length: 100
effective length of query: 325
effective length of database: 6,272,032
effective search space: 2038410400
effective search space used: 2038410400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0227.11


