

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.9
(223 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
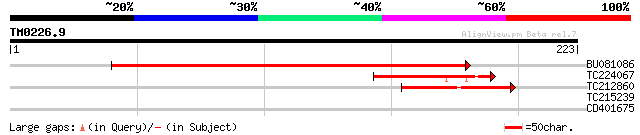
BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza s... 125 1e-29
TC224067 44 6e-05
TC212860 weakly similar to UP|PIF1_YEAST (P07271) DNA repair and... 42 3e-04
TC215239 UP|H3_PEA (P02300) Histone H3, partial (39%) 35 0.035
CD401675 similar to GP|25073695|gb NMDA receptor subunit NR2B {A... 27 5.6
>BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza sativa
(japonica cultivar-group)}, partial (7%)
Length = 426
Score = 125 bits (315), Expect = 1e-29
Identities = 66/141 (46%), Positives = 88/141 (61%)
Frame = +2
Query: 41 SDSALRSDEDSEIKVEWFTTEFSNDTKSSGMPNHQLLLKVGTPIMLLRNIDQAASLCNGT 100
SDS +SD TTEF N +S +PNH++ LKV + IMLLRN+DQ LCN T
Sbjct: 2 SDSIEKSDTIESEGFSTITTEFINSLSTSRLPNHKIKLKVDSRIMLLRNLDQNEGLCNST 181
Query: 101 RLIVDELGERFIGATVITGTNVNDKVHILRMDLVLSDSKFPIKFRIRQFPIAICFAMTIN 160
RLI+ + I A +++G + V+I R+ S S +P K R+FPI + +AMTIN
Sbjct: 182 RLIITMFADHIIEAKIMSGKGQGNTVYIPRLATSPSQSPWPFKLIRRKFPIIVSYAMTIN 361
Query: 161 KSQGQTLSQVGLFLPRPVFTH 181
KSQGQ L+ VGL+LP PVF+H
Sbjct: 362 KSQGQLLASVGLYLPTPVFSH 424
>TC224067
Length = 415
Score = 43.9 bits (102), Expect = 6e-05
Identities = 26/50 (52%), Positives = 34/50 (68%), Gaps = 2/50 (4%)
Frame = -2
Query: 144 FRIRQFPIAICFAMTINKSQGQTLSQV-GLFLPRPV-FTHGQLYVALSRV 191
F+ + +C+ M INKS GQTLSQV G+ LPRP +HGQ YV L++V
Sbjct: 387 FKFQDVNSLVCYTMKINKSHGQTLSQVGGVLLPRPN**SHGQ-YVTLNQV 241
>TC212860 weakly similar to UP|PIF1_YEAST (P07271) DNA repair and
recombination protein PIF1, mitochondrial precursor,
partial (8%)
Length = 596
Score = 41.6 bits (96), Expect = 3e-04
Identities = 25/45 (55%), Positives = 31/45 (68%)
Frame = +2
Query: 155 FAMTINKSQGQTLSQVGLFLPRPVFTHGQLYVALSRVRSRKGLKL 199
+AM+I+K QG TL +V L R F G +YVALSRVRS +GL L
Sbjct: 5 WAMSIHKCQGMTLERVHTDLSR-AFGCGMVYVALSRVRSLEGLHL 136
>TC215239 UP|H3_PEA (P02300) Histone H3, partial (39%)
Length = 340
Score = 34.7 bits (78), Expect = 0.035
Identities = 13/18 (72%), Positives = 16/18 (88%)
Frame = -3
Query: 149 FPIAICFAMTINKSQGQT 166
FP+ +CFAMT NKS+GQT
Sbjct: 326 FPLIVCFAMTTNKSEGQT 273
>CD401675 similar to GP|25073695|gb NMDA receptor subunit NR2B {Apteronotus
leptorhynchus}, partial (1%)
Length = 338
Score = 27.3 bits (59), Expect = 5.6
Identities = 10/36 (27%), Positives = 18/36 (49%)
Frame = +2
Query: 142 IKFRIRQFPIAICFAMTINKSQGQTLSQVGLFLPRP 177
+ F FP ++CF+ + S +++ L L RP
Sbjct: 203 LSFNFHSFPKSLCFSYASSSSSSSSVAAAALLLHRP 310
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.137 0.389
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,276,660
Number of Sequences: 63676
Number of extensions: 97700
Number of successful extensions: 372
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 371
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 372
length of query: 223
length of database: 12,639,632
effective HSP length: 94
effective length of query: 129
effective length of database: 6,654,088
effective search space: 858377352
effective search space used: 858377352
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0226.9


