

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.7
(399 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
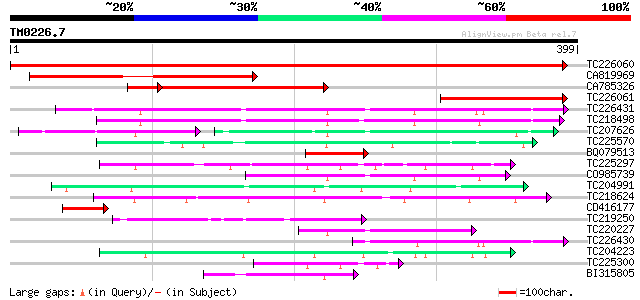
Score E
Sequences producing significant alignments: (bits) Value
TC226060 similar to GB|AAP68293.1|31711874|BT008854 At1g80360 {A... 669 0.0
CA819969 237 6e-63
CA785326 234 7e-62
TC226061 similar to GB|AAP68293.1|31711874|BT008854 At1g80360 {A... 157 1e-38
TC226431 similar to GB|AAM70520.1|21700793|AY124811 At2g22250/T2... 127 9e-30
TC218498 similar to GB|AAM70520.1|21700793|AY124811 At2g22250/T2... 124 6e-29
TC207626 49 2e-14
TC225570 similar to GB|AAR99909.1|41323503|AY518701 aminotransfe... 74 2e-13
BQ079513 73 2e-13
TC225297 similar to UP|Q6VEJ5 (Q6VEJ5) Alanine aminotransferase,... 73 3e-13
CO985739 63 2e-10
TC204991 similar to UP|Q9FN30 (Q9FN30) Tyrosine aminotransferase... 62 6e-10
TC218624 homologue to UP|1A1C_SOYBN (P31531) 1-aminocyclopropane... 58 9e-09
CD416177 similar to GP|29293694|gb| small GTP binding protein {O... 55 6e-08
TC219250 similar to UP|O82030 (O82030) Histidinol-phosphate amin... 55 7e-08
TC220227 weakly similar to UP|Q6ICW1 (Q6ICW1) LOC56267 protein (... 50 1e-06
TC226430 similar to GB|AAM70520.1|21700793|AY124811 At2g22250/T2... 46 3e-05
TC204223 similar to PIR|B86367|B86367 protein F26F24.16 [importe... 45 6e-05
TC225300 44 1e-04
BI315805 homologue to PIR|PN0477|PN0 1-aminocyclopropane-1-carbo... 42 4e-04
>TC226060 similar to GB|AAP68293.1|31711874|BT008854 At1g80360 {Arabidopsis
thaliana;} , partial (89%)
Length = 1770
Score = 669 bits (1727), Expect = 0.0
Identities = 327/393 (83%), Positives = 362/393 (91%), Gaps = 1/393 (0%)
Frame = +1
Query: 1 MGSYANLARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPS 60
MGS L+RRALETEMP+MVQMQE+ RGAKN +SLAQGVVYWQPPK AL+KVKELV EP
Sbjct: 175 MGSSVKLSRRALETEMPVMVQMQELQRGAKNALSLAQGVVYWQPPKQALEKVKELVSEPL 354
Query: 61 VSRYGADEGLPELRAALVKKLRQENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFA 120
+SRYG DEG+PELRAALVKKLR ENNLHKSSVMVT+GANQAFVNLVLTLCD GDSVVMFA
Sbjct: 355 ISRYGNDEGIPELRAALVKKLRDENNLHKSSVMVTSGANQAFVNLVLTLCDPGDSVVMFA 534
Query: 121 PYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGT 180
PYYFNAYMSFQMTGITNI VGPGS DTL+PD DWLERIL+E+KP PKLVTVVNPGNP+GT
Sbjct: 535 PYYFNAYMSFQMTGITNILVGPGSSDTLHPDADWLERILSENKPAPKLVTVVNPGNPSGT 714
Query: 181 YIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCVEGNHIVNIFSFSKAYGMMGW 240
YIPE LL+RI++LCKNAGSWL+VDNTYEYFM+DGLKHSCVEGNHIVN+FSFSKA+GMMGW
Sbjct: 715 YIPEPLLKRISDLCKNAGSWLVVDNTYEYFMYDGLKHSCVEGNHIVNVFSFSKAFGMMGW 894
Query: 241 RVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNRE 300
RVGYIAYPSEV+ + QLLKVQDNIPICASI+SQ+LAL+SLE+GP+WV ++V+TL KNRE
Sbjct: 895 RVGYIAYPSEVKDFAEQLLKVQDNIPICASILSQYLALYSLEVGPQWVVDQVKTLEKNRE 1074
Query: 301 IVSEALSPLGEGSVKGGEGAIYLFAKLPK-GGYDDFDVVRWLANRHGIAVIPGSACGAAG 359
IV EALSPLGEGSVKGGEGAIYL+AKLP +DDFDVVRWLAN+HG+AVIPG ACG
Sbjct: 1075IVLEALSPLGEGSVKGGEGAIYLWAKLPDLDAHDDFDVVRWLANKHGVAVIPGKACGCPS 1254
Query: 360 NLRISFGGLTESDCRAAAERLKKGLEELVANGL 392
NLRISFGGLTE+DCRAAAERLKKGLEELV +GL
Sbjct: 1255NLRISFGGLTENDCRAAAERLKKGLEELVRDGL 1353
>CA819969
Length = 422
Score = 237 bits (605), Expect = 6e-63
Identities = 121/160 (75%), Positives = 130/160 (80%)
Frame = +3
Query: 15 EMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELR 74
EMP+MVQMQE+LRGAKN VSLAQGVVYWQPPK AL+KVKELV EP +SRYG DEG+PELR
Sbjct: 3 EMPVMVQMQELLRGAKNAVSLAQGVVYWQPPKQALEKVKELVSEPLISRYGNDEGIPELR 182
Query: 75 AALVKKLRQENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTG 134
AALVKK AFVNLVLTLCD GDSVVMFAPYYFNAYMSFQMTG
Sbjct: 183 AALVKK--------------------AFVNLVLTLCDPGDSVVMFAPYYFNAYMSFQMTG 302
Query: 135 ITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNP 174
+TNI VGPGS DTL+PD DWLERIL+E+KP PKLVTVVNP
Sbjct: 303 VTNILVGPGSSDTLHPDADWLERILSETKPPPKLVTVVNP 422
>CA785326
Length = 425
Score = 234 bits (596), Expect = 7e-62
Identities = 110/141 (78%), Positives = 123/141 (87%)
Frame = +1
Query: 84 ENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHVGPG 143
EN +HK + FV+LVLTLCD GDSVVMFAPYYFNAYMSFQMTG+TNI VGPG
Sbjct: 1 EN*IHKCLIGT*CNHY*RFVHLVLTLCDPGDSVVMFAPYYFNAYMSFQMTGVTNILVGPG 180
Query: 144 SPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLV 203
S DTL+PD DWLERIL+E+KP PKLVTVVNPGNP+GTYIPE LL+RI++LCKNAGSWL+V
Sbjct: 181 SSDTLHPDADWLERILSETKPPPKLVTVVNPGNPSGTYIPEPLLKRISDLCKNAGSWLVV 360
Query: 204 DNTYEYFMFDGLKHSCVEGNH 224
DNTYEYFM+DGLKHSCVEGNH
Sbjct: 361 DNTYEYFMYDGLKHSCVEGNH 423
Score = 42.4 bits (98), Expect = 4e-04
Identities = 20/24 (83%), Positives = 23/24 (95%)
Frame = -3
Query: 84 ENNLHKSSVMVTAGANQAFVNLVL 107
+N +HKSSVMVT+GANQAFVNLVL
Sbjct: 72 KN*MHKSSVMVTSGANQAFVNLVL 1
>TC226061 similar to GB|AAP68293.1|31711874|BT008854 At1g80360 {Arabidopsis
thaliana;} , partial (7%)
Length = 478
Score = 157 bits (396), Expect = 1e-38
Identities = 77/90 (85%), Positives = 83/90 (91%), Gaps = 1/90 (1%)
Frame = +2
Query: 304 EALSPLGEGSVKGGEGAIYLFAKLPKGG-YDDFDVVRWLANRHGIAVIPGSACGAAGNLR 362
EALSPLGEGSVKGGEGAIYL+AKLP G +DDFDVVRWLAN+HG+AVIPG ACG GNLR
Sbjct: 8 EALSPLGEGSVKGGEGAIYLWAKLPHGNAHDDFDVVRWLANKHGVAVIPGKACGCPGNLR 187
Query: 363 ISFGGLTESDCRAAAERLKKGLEELVANGL 392
ISFGGLTE+DCRAAAERLKKGLEELV +GL
Sbjct: 188 ISFGGLTENDCRAAAERLKKGLEELVRDGL 277
>TC226431 similar to GB|AAM70520.1|21700793|AY124811 At2g22250/T26C19.9
{Arabidopsis thaliana;} , partial (85%)
Length = 1715
Score = 127 bits (319), Expect = 9e-30
Identities = 108/382 (28%), Positives = 180/382 (46%), Gaps = 21/382 (5%)
Frame = +2
Query: 33 VSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQENNLHKS-- 90
+ LA G + P P + + E +RY + G ELR A+ KL++EN + +
Sbjct: 365 IRLAAGEPDFDTPAPIAEAGINAIRE-GYTRYTPNAGTMELRQAICHKLKEENGITYTPD 541
Query: 91 SVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHVGPGSPDTLYP 150
V+V+ GA Q+ VL + GD V++ AP++ + ++ T + + D
Sbjct: 542 QVVVSNGAKQSIAQAVLAVSSPGDEVIIPAPFWVSYPEMARLADATPVILPTLISDNFLL 721
Query: 151 DVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANL-CKNAGSWLLVDNTYEY 209
D LE +TE +L+ + +P NPTG+ P+ LL+ IA + K+ +L D YE+
Sbjct: 722 DPKLLESKITERS---RLLILCSPSNPTGSVYPKELLEEIARIVAKHPRLLVLSDEIYEH 892
Query: 210 FMFDGLKHSCVEG-----NHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDN 264
++ H+ + + + FSKA+ M GWR+GYIA P A K+Q
Sbjct: 893 IIYAPATHTSFASLPGMWDRTLTVNGFSKAFAMTGWRLGYIAGPKH---FVAACGKIQSQ 1063
Query: 265 IPICASIISQHLALHSLEM---GPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAI 321
AS I+Q A+ +L + G E V V+ + R+ + ++ + + +GA
Sbjct: 1064FTSGASSIAQKAAVAALGLGHAGGEAVSTMVKAFRERRDFLVQSFREIDGIKISEPQGAF 1243
Query: 322 YLFAKL------PKGGY----DDFDVVRWLANRHGIAVIPGSACGAAGNLRISFGGLTES 371
YLF L G+ D + ++L +A++PGSA G +RIS+ + +
Sbjct: 1244YLFLDLSFYYGREAEGFGKIVDSESLCQYLLEVGQVALVPGSAFGDDTCIRISYAE-SLT 1420
Query: 372 DCRAAAERLKKGLEELVANGLV 393
+AA ER+KK L L + LV
Sbjct: 1421TLQAAVERIKKALVPLSSAALV 1486
>TC218498 similar to GB|AAM70520.1|21700793|AY124811 At2g22250/T26C19.9
{Arabidopsis thaliana;} , partial (78%)
Length = 1214
Score = 124 bits (312), Expect = 6e-29
Identities = 99/350 (28%), Positives = 171/350 (48%), Gaps = 21/350 (6%)
Frame = +2
Query: 62 SRYGADEGLPELRAALVKKLRQENNLHKS--SVMVTAGANQAFVNLVLTLCDAGDSVVMF 119
+RY + G ELR A+ KL++EN + + ++V+ GA Q+ V VL +C GD V++
Sbjct: 92 TRYTPNAGTLELRQAICHKLKEENEITYTPDEIVVSNGAKQSVVQAVLAVCSPGDEVIIP 271
Query: 120 APYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTG 179
AP+Y + ++ T + + + D LE LTE +L+ + +P NPTG
Sbjct: 272 APFYTSYPEMARLADATPVILPSHISNNFLLDPKLLEANLTERS---RLLILCSPCNPTG 442
Query: 180 TYIPESLLQRIANL-CKNAGSWLLVDNTYEYFMFDGLKHSCVEG-----NHIVNIFSFSK 233
+ + LL+ IA + K+ +L D YE+ ++ H+ + + + FSK
Sbjct: 443 SVYSKKLLEEIAQIVAKHPRLLVLSDEIYEHIIYAPATHTSFASLPGMWDRTLTVNGFSK 622
Query: 234 AYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEM---GPEWVRE 290
+ M GWR+GYIA + A K+Q AS ISQ + +L + G E V
Sbjct: 623 TFAMTGWRLGYIA---GTKHFVAACGKIQSQFTSGASSISQKAGVAALGLGYAGGEAVST 793
Query: 291 RVQTLAKNREIVSEALSPLGEGSVKGGEGAIYLFAKLPK---------GGYDDFD-VVRW 340
V+ + R+ + E+ + + +GA YLF G ++ D + R+
Sbjct: 794 MVKAFRERRDFLVESFREMDGVKISEPQGAFYLFIDFSSYYGREVEGFGIIENSDSLCRY 973
Query: 341 LANRHGIAVIPGSACGAAGNLRISFGGLTESDCRAAAERLKKGLEELVAN 390
L ++ +A++PGSA G +RIS+ + ++ + A ER+KK L L+++
Sbjct: 974 LLDKGLVALVPGSAFGDDSCIRISYAE-SLTNLKTAVERIKKALIPLISS 1120
>TC207626
Length = 1824
Score = 48.5 bits (114), Expect(2) = 2e-14
Identities = 30/131 (22%), Positives = 64/131 (47%), Gaps = 3/131 (2%)
Frame = +1
Query: 7 LARRALETEMPIMVQMQEVLRGAKNCVSLAQGVVYWQPPKPALDKVKELVWEPSVSRYGA 66
+A+R + + I QM +L ++L QG + P+ + + + + ++Y
Sbjct: 199 IAKRLEKFQTTIFTQMS-LLAIKHGAINLGQGFPNFDGPEFVKEAAIQAIRDGK-NQYAR 372
Query: 67 DEGLPELRAALVKKLRQENNL---HKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYY 123
G+P+L A+ ++ +++ L + + VT+G + ++ L + GD V+MFAP+Y
Sbjct: 373 GYGVPDLNIAIAERFKKDTGLVVDPEKEITVTSGCTETIAATMIGLINPGDEVIMFAPFY 552
Query: 124 FNAYMSFQMTG 134
+ + M G
Sbjct: 553 DSYEATLSMAG 585
Score = 48.1 bits (113), Expect(2) = 2e-14
Identities = 56/252 (22%), Positives = 91/252 (35%), Gaps = 10/252 (3%)
Frame = +2
Query: 145 PDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVD 204
PD P LE + + + + + P NPTG L IA+LC + D
Sbjct: 617 PDFAVP----LEELKSTISKNTRAILINTPHNPTGKMFTREELNCIASLCIENDVLVFTD 784
Query: 205 NTYEYFMFDGLKHSCVEG-----NHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLL 259
Y+ FD ++H + V + S K + + GW++G+ P LS +
Sbjct: 785 EVYDKLAFD-MEHISMASLPGMFERTVTLNSLGKTFSLTGWKIGWAIAPPH---LSWGVR 952
Query: 260 KVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEG 319
+ + + Q A +L + E + R I+ E L +G
Sbjct: 953 QAHAFLTFATAHPFQCAAAAALRAPDSYYVELKRDYMAKRAILIEGLKAVGFKVFPSSGT 1132
Query: 320 AIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSAC-----GAAGNLRISFGGLTESDCR 374
+ P G +D +L N G+ IP S +R +F E R
Sbjct: 1133YFVVVDHTPFGLENDVAFCEYLVNEVGVVAIPTSVFYLNPEEGKNLVRFTF-CKDEETIR 1309
Query: 375 AAAERLKKGLEE 386
+A ER+K L +
Sbjct: 1310SAVERMKAKLRK 1345
>TC225570 similar to GB|AAR99909.1|41323503|AY518701 aminotransferase AGD2
{Arabidopsis thaliana;} , partial (86%)
Length = 1414
Score = 73.6 bits (179), Expect = 2e-13
Identities = 83/336 (24%), Positives = 135/336 (39%), Gaps = 26/336 (7%)
Frame = +3
Query: 62 SRYGADEGLPELRAALVKKLRQENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFA- 120
S YGA++G LR AL + + + + V+ GA L + G +V M
Sbjct: 189 SGYGAEQGEKPLRRALASTFYSDLGIEEDDIFVSDGAKCDISRLQIVF---GSNVKMAVQ 359
Query: 121 ----PYYFNAYMSFQMTGI--------TNIHVGPGSPDT-LYPDVDWLERILTESKPVPK 167
P Y ++ + TG+ NI +P+ +PD+ + R P
Sbjct: 360 DPSYPAYVDSSVIMGQTGLFQKNVEKFANIEYMRCNPENGFFPDLSSISR--------PD 515
Query: 168 LVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCVE----GN 223
++ +P NPTG L ++ K+ GS ++ D+ Y ++ S E
Sbjct: 516 IIFFCSPNNPTGAVATREQLTQLVQFAKDNGSIVIHDSAYAMYISGDNPRSIFEIPGAKE 695
Query: 224 HIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPIC-----ASIISQHLAL 278
+ SFSK G G R+G+ P ++ + N +C AS ISQ L
Sbjct: 696 VAIETSSFSKYAGFTGVRLGWTVVPKQLLFSDGFPVAKDFNRIVCTCFNGASNISQAGGL 875
Query: 279 HSLE-MGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIYLFAKLPKGGYDDFDV 337
L G + +R+ + +N I+ E LG V GG+ A Y++ P G +DV
Sbjct: 876 ACLSPEGLKAMRDVIGFYKENTNIIMETFDSLG-FKVYGGKDAPYVWVHFP--GRSSWDV 1046
Query: 338 VRWLANRHGIAVIPGSACGAAGN--LRISFGGLTES 371
+ + + PGS G G +R+S G E+
Sbjct: 1047FAEILEKTHVVTTPGSGFGPGGEGFIRVSAFGHREN 1154
>BQ079513
Length = 415
Score = 73.2 bits (178), Expect = 2e-13
Identities = 33/44 (75%), Positives = 36/44 (81%)
Frame = -1
Query: 209 YFMFDGLKHSCVEGNHIVNIFSFSKAYGMMGWRVGYIAYPSEVE 252
YFM+DGLKHSCVEGNHIVN+FS SK YGMMG AYPSEV+
Sbjct: 133 YFMYDGLKHSCVEGNHIVNVFSLSKTYGMMGCLRSCSAYPSEVK 2
>TC225297 similar to UP|Q6VEJ5 (Q6VEJ5) Alanine aminotransferase, partial
(98%)
Length = 1833
Score = 72.8 bits (177), Expect = 3e-13
Identities = 94/347 (27%), Positives = 144/347 (41%), Gaps = 54/347 (15%)
Frame = +1
Query: 64 YGADEGLPELRAALVKKLRQENNL--HKSSVMVTAGANQAFVNLV-LTLCDAGDSVVMFA 120
Y +G+ LR + + + + + + +T GA+ A N++ L + D ++
Sbjct: 481 YSHSQGVKGLRDTIAAGIEERDGFPANPDDIFMTDGASPAVHNMMQLLIRSENDGILCPI 660
Query: 121 PYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDW----------LERILTESKPVPKLVT 170
P Y S ++H G P L W LE ++ V LV
Sbjct: 661 PQYPLYSASI------DLHGGCLVPYYLDEATGWGLEIPELKKQLEAAKSKGINVRALV- 819
Query: 171 VVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYE---------YFMFDGLKHSCVE 221
V+NPGNPTG + E+ + I CK G LL D Y+ + F + S
Sbjct: 820 VINPGNPTGQVLGEANQRDIVEFCKQEGLVLLADEVYQENVYVPEKKFHSFKKVSRSMGY 999
Query: 222 GNHIVNIFSF---SKAY-GMMGWRVGYIAYPSEVEGLSA----QLLKVQDNIPICASIIS 273
G + + + SF SK Y G G R GY+ EV G SA Q+ KV ++ +C++I
Sbjct: 1000GENDITLVSFQSVSKGYHGECGKRGGYM----EVTGFSAEVREQIYKVA-SVNLCSNISG 1164
Query: 274 QHLALHSLEMGPEWVRER------------VQTLAKNREIVSEALSPLGEGSVKGGEGAI 321
Q LA SL M P V + + +LA+ + + +A + L + EGA+
Sbjct: 1165QILA--SLVMSPPKVGDESYDSFMAEKENILASLARRAKTLEDAFNKLEGVTCNKAEGAM 1338
Query: 322 YLF------------AKLPKGGYDDFDVVRWLANRHGIAVIPGSACG 356
YLF A+ D+F R L N G+ V+PGS G
Sbjct: 1339YLFPQIRLSEKAIKAAEAANATPDNFYCKR-LLNATGVVVVPGSGFG 1476
>CO985739
Length = 806
Score = 63.2 bits (152), Expect = 2e-10
Identities = 52/205 (25%), Positives = 93/205 (45%), Gaps = 19/205 (9%)
Frame = -3
Query: 167 KLVTVVNPGNPTGTYIPESLLQRIANL-CKNAGSWLLVDNTYEYFMFDGLKHSCVEG--- 222
+L+ + +P NPTG+ + LL+ IA + K+ +L D YE+ ++ H+
Sbjct: 795 RLLILCSPCNPTGSVYSKKLLEEIAQIVAKHPRLLVLSDENYEHIIYAPATHTSFASLPG 616
Query: 223 --NHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHS 280
+ + + SK + M GWR+GYIA P A K+Q AS ISQ + +
Sbjct: 615 MWDRTLIVNGLSKTFAMTGWRLGYIAGPKH---FVAACEKIQSQFTSGASSISQKAGVAA 445
Query: 281 LEM---GPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIYLFAKLPK-------- 329
L + G E V V+ + R+ + E+ + + +G Y+F
Sbjct: 444 LGLGYAGGEAVSTMVKAFRERRDFLVESFREMDGVKICEPQGGFYVFLDFSSYYGREAEG 265
Query: 330 -GGYDDFD-VVRWLANRHGIAVIPG 352
G ++ D + R+L ++ +A++PG
Sbjct: 264 FGVIENSDSLCRYLLDKGLVALVPG 190
>TC204991 similar to UP|Q9FN30 (Q9FN30) Tyrosine aminotransferase, partial
(89%)
Length = 1743
Score = 61.6 bits (148), Expect = 6e-10
Identities = 80/362 (22%), Positives = 137/362 (37%), Gaps = 26/362 (7%)
Frame = +2
Query: 30 KNCVSLAQG----VVYWQPPKPALDKVKELVWEPSVSRYGADEGLPELRAALVKKLRQE- 84
K +SL G + PK + V + + Y GL + R A+ + L ++
Sbjct: 401 KRVISLGMGDPTLTTLFHTPKVVEEAVADALQSRKFHGYAPTAGLLQARIAIAEYLSRDL 580
Query: 85 -NNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQMTGITNIHVGPG 143
L + V +T G QA V L G ++++ P + + G+ H
Sbjct: 581 PYQLSRDDVFITCGCTQAIDVSVAMLARPGANILLPRPGFPIYELCAAFRGVEVRHYDLL 760
Query: 144 SPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKNAGSWLLV 203
D+D +E + ++ + ++NPGNP G L++IA K G+ ++
Sbjct: 761 PEKGWEVDLDAVEALADQNTVA---LAIINPGNPCGNVYSYHHLEKIAETAKRVGTIVIS 931
Query: 204 DNTYEYFMFD-------GLKHSCVEGNHIVNIFSFSKAYGMMGWRVGYIA---------Y 247
D Y + F G+ S V ++ + S SK + + GWR+G+
Sbjct: 932 DEVYGHLAFGSKPFVPMGVFGSTVP---VLTLGSLSKRWIVPGWRLGWFVTNDPSGTFRE 1102
Query: 248 PSEVEGLSAQLLKVQDNIPICASIISQHLALHS---LEMGPEWVRERVQTLAKNREIVSE 304
P VE + + + + Q +A E + +R K E +
Sbjct: 1103PKVVERIKKYFDLLGGPATFLQAAVPQIIANTEEIFFEKTIDNLRHTADICCKEIEDIPC 1282
Query: 305 ALSPL-GEGSVKGGEGAIYLFAKLPKGGYDDFDVVRWLANRHGIAVIPGSACGAAGNLRI 363
P EGS+ + L L + DD D LA + ++PG+A G LRI
Sbjct: 1283IFCPYKPEGSM---AMMVKLNLSLLEDISDDIDFCFKLAKEESVIILPGTAVGLKDWLRI 1453
Query: 364 SF 365
+F
Sbjct: 1454TF 1459
>TC218624 homologue to UP|1A1C_SOYBN (P31531)
1-aminocyclopropane-1-carboxylate synthase (ACC
synthase) (S-adenosyl-L-methionine
methylthioadenosine-lyase) , partial (90%)
Length = 1713
Score = 57.8 bits (138), Expect = 9e-09
Identities = 79/353 (22%), Positives = 145/353 (40%), Gaps = 31/353 (8%)
Frame = +3
Query: 60 SVSRYGADEGLPELRAALVKKL-RQENN---LHKSSVMVTAGANQAFVNLVLTLCDAGDS 115
+++ + GLPE R A+ K + R N ++++ GA A L D GD+
Sbjct: 108 AIANFQDYHGLPEFRNAVAKFMGRTRGNRVTFDPDRIVMSGGATGAHEVTTFCLADPGDA 287
Query: 116 VVMFAPYY--FNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESKPVP---KLVT 170
++ PYY F+ + ++ TGI + V S + LE ++K K +
Sbjct: 288 FLVPIPYYPGFDRDLRWR-TGIKLVPVMCDSSNNFKLTKQALEDAYEKAKEDNIRVKGLL 464
Query: 171 VVNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFDGLKHSCV---------- 220
+ NP NP GT + + L+ + + L+ D Y +F +
Sbjct: 465 ITNPSNPLGTVMDRNTLRTVMSFINEKRIHLVSDEIYSATVFSHPSFISIAEILEEDTDI 644
Query: 221 --EGNHIVNIFSFSKAYGMMGWRVGYI-AYPSEVEGLSAQLLKVQDNIPICASIISQHL- 276
+ N + ++S SK G G+RVG I +Y V + ++ S +QHL
Sbjct: 645 ECDRNLVHIVYSLSKDMGFPGFRVGIIYSYNDAVVHCARKMSSFG-----LVSTQTQHLL 809
Query: 277 --ALHSLEMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGEGAIY---LFAKLPKGG 331
L+ E ++ E + LA+ + + L+ +G ++ G L L K
Sbjct: 810 ASMLNDDEFVERFLEESAKRLAQRHRVFTSGLAKVGIKCLQSNAGLFVWMDLRQLLKKPT 989
Query: 332 YD-DFDVVRWLANRHGIAVIPGSA--CGAAGNLRISFGGLTESDCRAAAERLK 381
D + ++ R + + I V PGS+ C G R+ + + + + A +R++
Sbjct: 990 LDSEMELWRVIIHEVKINVSPGSSFHCTEPGWFRVCYANMDDMAVQIALQRIR 1148
>CD416177 similar to GP|29293694|gb| small GTP binding protein {Oryza sativa
(indica cultivar-group)}, partial (30%)
Length = 625
Score = 55.1 bits (131), Expect = 6e-08
Identities = 25/32 (78%), Positives = 27/32 (84%)
Frame = -1
Query: 38 GVVYWQPPKPALDKVKELVWEPSVSRYGADEG 69
GVVYWQPPK AL+KVKELV E +SRYG DEG
Sbjct: 475 GVVYWQPPKEALEKVKELVSETLISRYGNDEG 380
>TC219250 similar to UP|O82030 (O82030) Histidinol-phosphate aminotransferase
precursor , partial (87%)
Length = 1564
Score = 54.7 bits (130), Expect = 7e-08
Identities = 47/180 (26%), Positives = 86/180 (47%), Gaps = 1/180 (0%)
Frame = +1
Query: 73 LRAALVKKLRQENNLHKSSVMVTAGANQAFVNLVLTLCDAGDSVVMFAPYYFNAYMSFQM 132
LRAAL ++ L ++ GA++ ++ + D GD +V P + +
Sbjct: 580 LRAALA----HDSGLEAEYILAGCGADELIDLIMRCVLDPGDKIVDCPPTFTMYEFDAAV 747
Query: 133 TGITNIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIAN 192
G I V P PD +V+ + ++ + KP K + + +P NP G+ I + +L +I
Sbjct: 748 NGALVIKV-PRRPDFSL-NVEQIAEVVKQEKP--KCIFLTSPNNPDGSIIDDEVLLKILE 915
Query: 193 LCKNAGSWLLVDNTYEYFMFDGLKHSCVEGN-HIVNIFSFSKAYGMMGWRVGYIAYPSEV 251
L +++D Y F + S V+ + +++ + +FSK G+ G RVGY A+P +
Sbjct: 916 L----PILVILDEAYIEFSAIESRMSWVKKHDNLIVLRTFSKRAGLAGLRVGYGAFPLSI 1083
>TC220227 weakly similar to UP|Q6ICW1 (Q6ICW1) LOC56267 protein (Fragment),
partial (27%)
Length = 1196
Score = 50.4 bits (119), Expect = 1e-06
Identities = 34/130 (26%), Positives = 59/130 (45%), Gaps = 5/130 (3%)
Frame = +2
Query: 204 DNTYEYFMFDGLKHSCVEG-----NHIVNIFSFSKAYGMMGWRVGYIAYPSEVEGLSAQL 258
+ YE+ +D LKH + V S SK++ + GWRVG+ P+ L++ +
Sbjct: 89 EQVYEHITYDNLKHISLASFPGMLERTVITSSLSKSFSVTGWRVGWAIAPAF---LASAI 259
Query: 259 LKVQDNIPICASIISQHLALHSLEMGPEWVRERVQTLAKNREIVSEALSPLGEGSVKGGE 318
+ + A Q AL +L PE+ + R+ + + L +G V +
Sbjct: 260 RNIHGRVTDSAPAPFQEAALTALRSPPEYFESLRRDYQSKRDYIIKLLDGVGFKIVFIPQ 439
Query: 319 GAIYLFAKLP 328
G+ +LFA+LP
Sbjct: 440 GSFFLFAELP 469
>TC226430 similar to GB|AAM70520.1|21700793|AY124811 At2g22250/T26C19.9
{Arabidopsis thaliana;} , partial (33%)
Length = 686
Score = 45.8 bits (107), Expect = 3e-05
Identities = 46/165 (27%), Positives = 76/165 (45%), Gaps = 13/165 (7%)
Frame = +1
Query: 242 VGYIAYPSEVEGLSAQLLKVQDNIPICASIISQHLALHSLEMGP---EWVRERVQTLAKN 298
+GYIA P A K+Q AS I+Q A+ +L +G E V V+ +
Sbjct: 1 LGYIAGPKH---FVAACGKIQSQFTSGASSIAQKAAVAALGLGHAGGEAVSTMVKAFRER 171
Query: 299 REIVSEALSPLGEGSVKGGEGAIYLFAKLP------KGGY----DDFDVVRWLANRHGIA 348
R+ + ++ + + +GA YLF G+ D + R+L + +A
Sbjct: 172 RDFLVKSFREIDGVKISEPQGAFYLFLDFSFYYGREAEGFGKIEDSESLCRYLLDVGQVA 351
Query: 349 VIPGSACGAAGNLRISFGGLTESDCRAAAERLKKGLEELVANGLV 393
++PGSA G +RIS+ + + +AA ER+K+ L L + LV
Sbjct: 352 LVPGSAFGDDTCIRISYAE-SLTTLQAAVERVKRALIPLSSAALV 483
>TC204223 similar to PIR|B86367|B86367 protein F26F24.16 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , complete
Length = 2050
Score = 45.1 bits (105), Expect = 6e-05
Identities = 69/336 (20%), Positives = 133/336 (39%), Gaps = 43/336 (12%)
Frame = +1
Query: 64 YGADEGLPELRAALVKKLRQENNLHKSSVMV--TAGANQAFVNLVLTLCDA-GDSVVMFA 120
Y GLP +R + + + + + ++ T GA++ + ++ T+ D +++
Sbjct: 760 YSDSRGLPGVRKEVAEFILRRDGYPTDPELIYLTDGASKGVMQILNTIIRGQDDGILVPV 939
Query: 121 PYYFNAYMSFQMTGITNIHVGPGSPDTLYPDVDWLERILTESK---PVPKLVTVVNPGNP 177
P Y + + G T + DV+ L + + +++ K + ++NPGNP
Sbjct: 940 PQYPLYSATIALLGGTLVPYYLEETANWGLDVNELRQSVEQARFKGITVKAMVIINPGNP 1119
Query: 178 TGTYIPESLLQRIANLCKNAGSWLLVDNTYEYFMFD-------------GLKHSCVEGNH 224
TG + E+ L+ + C LL D Y+ ++ L +
Sbjct: 1120 TGQCLSEANLREVLQFCYQENLALLGDEVYQTNIYQDERPFISSRKVLMDLGPPISKEVQ 1299
Query: 225 IVNIFSFSKA-YGMMGWRVGYIAY----PSEVEGL-SAQLLKVQDNIPICASIISQHLAL 278
+++ S SK YG G R GY P V+ + + + N+P + I + L
Sbjct: 1300 LISFHSVSKGYYGECGQRGGYFEMTNIPPETVDEIYKVASISLSPNVP---AQIFMGVML 1470
Query: 279 HSLEMG----PEWVRER---VQTLAKNREIVSEALSPLGEGSVKGGEGAIYLF--AKLPK 329
H + G ++VRE +++L + ++++ + EGA+Y F +LP
Sbjct: 1471 HPPQPGDISYDKFVRESTGILESLRRRARLMTDGFNSCRNVVCNFTEGAMYSFPQIRLPP 1650
Query: 330 GGYD---------DFDVVRWLANRHGIAVIPGSACG 356
+ D L GI+ +PGS G
Sbjct: 1651 RALEAAKQAAKVPDVFYCLKLLEATGISTVPGSGFG 1758
>TC225300
Length = 1269
Score = 43.9 bits (102), Expect = 1e-04
Identities = 39/123 (31%), Positives = 56/123 (44%), Gaps = 17/123 (13%)
Frame = +1
Query: 172 VNPGNPTGTYIPESLLQRIANLCKNAGSWLLVDNTYE---------YFMFDGLKHSCVEG 222
+NPGNPTG + E + I CK G L D Y+ + F + S G
Sbjct: 802 INPGNPTGQVLGEENQRDIVEFCKQEGLVRLADEVYQENVYVPEKKFHSFKKVSRSMGYG 981
Query: 223 NHIVNIFSF---SKAY-GMMGWRVGYIAYPSEVEGLSAQ----LLKVQDNIPICASIISQ 274
+ + + SF SK Y G G R GY+ EV G SA+ + KV+ +I +C++
Sbjct: 982 ENDITLVSFQSVSKGYHGECGKRGGYM----EVTGFSAKVRELMYKVRSDI-LCSNFFG* 1146
Query: 275 HLA 277
LA
Sbjct: 1147ILA 1155
>BI315805 homologue to PIR|PN0477|PN0 1-aminocyclopropane-1-carboxylate
synthase (EC 4.4.1.14) 5 - mung bean (fragment), partial
(34%)
Length = 408
Score = 42.4 bits (98), Expect = 4e-04
Identities = 29/119 (24%), Positives = 50/119 (41%), Gaps = 10/119 (8%)
Frame = +1
Query: 137 NIHVGPGSPDTLYPDVDWLERILTESKPVPKLVTVVNPGNPTGTYIPESLLQRIANLCKN 196
N + P + + Y D + + + + V + NP NP G IP S+L+ I +
Sbjct: 40 NFQITPEALEAAYKDAEAMNSKV-------RGVLITNPSNPLGVTIPRSVLEEILDFVTR 198
Query: 197 AGSWLLVDNTYEYFMFDGLKHSCV----------EGNHIVNIFSFSKAYGMMGWRVGYI 245
L+ D Y +F + + V + + ++S SK G+ G+RVG I
Sbjct: 199 KNIHLVSDEIYSGSVFSSSEFTSVAEILEARQYKDAERVHIVYSLSKDLGLPGFRVGTI 375
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.136 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,724,652
Number of Sequences: 63676
Number of extensions: 250090
Number of successful extensions: 1080
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 1059
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1066
length of query: 399
length of database: 12,639,632
effective HSP length: 99
effective length of query: 300
effective length of database: 6,335,708
effective search space: 1900712400
effective search space used: 1900712400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0226.7


