

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.6
(263 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
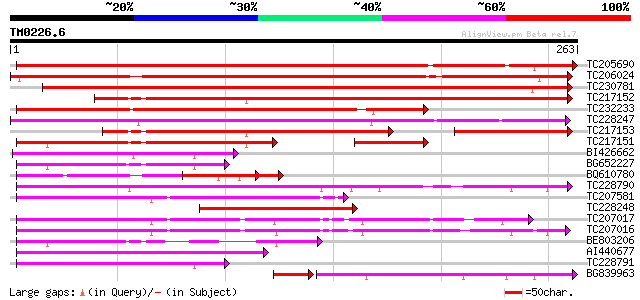
Score E
Sequences producing significant alignments: (bits) Value
TC205690 weakly similar to UP|Q6K3E9 (Q6K3E9) F-box family prote... 312 9e-86
TC206024 weakly similar to UP|Q9FLU7 (Q9FLU7) Phloem-specific le... 259 7e-70
TC230781 weakly similar to UP|Q6K3E9 (Q6K3E9) F-box family prote... 252 1e-67
TC217152 similar to UP|Q9FLU7 (Q9FLU7) Phloem-specific lectin-li... 217 5e-57
TC232233 weakly similar to GB|AAT41865.1|48310676|BT014882 At1g8... 192 2e-49
TC228247 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific le... 191 2e-49
TC217153 similar to GB|AAS76702.1|45773796|BT012215 At1g56240 {A... 126 6e-44
TC217151 similar to UP|Q6NPT8 (Q6NPT8) At2g02230, partial (6%) 126 2e-33
BI426662 102 2e-22
BG652227 97 7e-21
BQ610780 92 2e-19
TC228790 92 3e-19
TC207581 78 4e-15
TC228248 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific le... 77 8e-15
TC207017 76 1e-14
TC207016 75 2e-14
BE803206 72 3e-13
AI440677 61 5e-10
TC228791 57 7e-09
BG839963 weakly similar to GP|21592660|gb| phloem-specific lecti... 47 9e-07
>TC205690 weakly similar to UP|Q6K3E9 (Q6K3E9) F-box family protein-like,
partial (33%)
Length = 1360
Score = 312 bits (800), Expect = 9e-86
Identities = 165/261 (63%), Positives = 194/261 (74%), Gaps = 1/261 (0%)
Frame = +2
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPEGCIA+ILSRTTP D R S+VSKIFRSAA+SDAVW RFLPSD HSI+SQS S N
Sbjct: 215 LPEGCIASILSRTTPADVCRFSVVSKIFRSAAESDAVWKRFLPSDYHSIISQSPSPLNYP 394
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPES 123
SKK YL LSD P++ID GKKSFQL+KKSGKKC+ML ARAL I W + + Y WTT S
Sbjct: 395 SKKELYLALSDRPIIIDQGKKSFQLEKKSGKKCYMLAARALSIIWGDTEQYWNWTTDTNS 574
Query: 124 RFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIGGH 183
RFP+V LR + WLEI G++NT LSPNTQY A+LVFK+IDA FH V LSVN GGH
Sbjct: 575 RFPEVAELRDVCWLEIRGVLNTLVLSPNTQYAAYLVFKMIDARGFHNRPVELSVNVFGGH 754
Query: 184 NITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISGLEDEEVEMSVLQI- 242
TKI L+P+ E H R + +Q P+ RSDGWLEIEMGEFF +GL D+EV+MSV++
Sbjct: 755 GSTKIVCLDPN--EELPHRRVEGLQRPNARSDGWLEIEMGEFFNTGL-DDEVQMSVVETK 925
Query: 243 DGCFNVGLIIEGIEVRPKNDN 263
G + GL IEGIEV+PK +N
Sbjct: 926 GGNWKSGLFIEGIEVKPKEEN 988
>TC206024 weakly similar to UP|Q9FLU7 (Q9FLU7) Phloem-specific lectin-like
protein, partial (47%)
Length = 954
Score = 259 bits (663), Expect = 7e-70
Identities = 145/264 (54%), Positives = 182/264 (68%), Gaps = 3/264 (1%)
Frame = +3
Query: 1 MEL--LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLS 58
MEL LPEGCIA ILS TTPVD RLSLVSK FRSAA+SD VWD FL SD SI+ S
Sbjct: 21 MELQDLPEGCIAKILSYTTPVDVCRLSLVSKAFRSAAESDTVWDCFLLSDFTSIIPIS-- 194
Query: 59 RANASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWT 118
++SKK Y LSDHP +I G+KS QLDK++GKKC ML AR L I W + + WT
Sbjct: 195 ---STSKKDLYFTLSDHPTIIHQGRKSVQLDKRTGKKCCMLSARNLTIIWGDTVQHWEWT 365
Query: 119 TLPESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVN 178
+LPESRF +V +L+ + W +I G INT +LS NT Y FLVFK+I+A FH VLSV
Sbjct: 366 SLPESRFQEVAMLQAVCWFDISGSINTLTLSSNTHYATFLVFKMINASGFHYHPTVLSVG 545
Query: 179 NIGGHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISGLEDEEVEMS 238
+GG++ TK L+P +L+G + R +Q P VRSDGWLEIEMGEFF SG E+++V+M
Sbjct: 546 VLGGNSNTKYVCLDP-NLKG--NHRLQELQFPKVRSDGWLEIEMGEFFNSGQEEKQVQMK 716
Query: 239 VLQIDG-CFNVGLIIEGIEVRPKN 261
V++ + G I+EGIE+RPK+
Sbjct: 717 VMETTSHIWKCGFILEGIEIRPKH 788
>TC230781 weakly similar to UP|Q6K3E9 (Q6K3E9) F-box family protein-like,
partial (29%)
Length = 971
Score = 252 bits (644), Expect = 1e-67
Identities = 134/247 (54%), Positives = 169/247 (68%), Gaps = 1/247 (0%)
Frame = +2
Query: 16 TTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANASSKKRFYLDLSDH 75
TT D RLSLVSK F SAA+++ VWD FLPSD SI+S S S S K YL LSD
Sbjct: 2 TTRADVCRLSLVSKAFHSAAEANTVWDCFLPSDLSSIISPSSSVPPFRSIKDLYLYLSDR 181
Query: 76 PVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPESRFPDVVVLRHLL 135
P +ID G KSFQL+K++G KC+ML AR L I W + HY WTTLPESRF +V LR +
Sbjct: 182 PTIIDQGIKSFQLEKRTGNKCYMLSARDLSIIWGDTTHYWEWTTLPESRFEEVARLRAVC 361
Query: 136 WLEIHGMINTFSLSPNTQYGAFLVFKIIDAPNFHEVLVVLSVNNIGGHNITKIAYLNPDS 195
W +I G +NT LSPNT Y AFLVFK+IDA FH VLSV +GG++ TK L+P+
Sbjct: 362 WFDITGRMNTRVLSPNTNYAAFLVFKMIDAGGFHYDPAVLSVGILGGNSSTKNVCLDPNL 541
Query: 196 LEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISGLEDEEVEMSVLQ-IDGCFNVGLIIEG 254
++ + DR+ +Q P+VRSDGWLEIEMGEFF GLE++E+++ V + + G I+EG
Sbjct: 542 VDNRLDDRFHGLQRPTVRSDGWLEIEMGEFFNPGLEEDELQIKVSETTSNWWKRGFILEG 721
Query: 255 IEVRPKN 261
IEVRPK+
Sbjct: 722 IEVRPKH 742
>TC217152 similar to UP|Q9FLU7 (Q9FLU7) Phloem-specific lectin-like protein,
partial (8%)
Length = 909
Score = 217 bits (552), Expect = 5e-57
Identities = 114/226 (50%), Positives = 149/226 (65%), Gaps = 4/226 (1%)
Frame = +3
Query: 40 VWDRFLPSDSHSIVSQSLSRANASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFML 99
VWDRF+PSD S +S LS +N SKK Y LSD P +ID G+KSFQL+K++ KKC+ML
Sbjct: 3 VWDRFIPSDFSSTISP-LSSSN--SKKDLYFTLSDRPTIIDQGRKSFQLEKRTAKKCYML 173
Query: 100 GARALFIAW----DNGDHYRCWTTLPESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYG 155
AR + I W Y W +LPESRF +V L + W I G I T LSPNTQY
Sbjct: 174 SARDISITWAPTQGEASQYWEWKSLPESRFQEVARLYAVCWFNITGQIKTRVLSPNTQYA 353
Query: 156 AFLVFKIIDAPNFHEVLVVLSVNNIGGHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSD 215
AFLVF++IDA FH +LSV+N+GG +K L+P+ + ++ DR+ +Q P+VR D
Sbjct: 354 AFLVFQMIDASGFHHHPAMLSVSNVGGSRTSKYVCLDPNLEDNDLDDRFRGLQRPNVRKD 533
Query: 216 GWLEIEMGEFFISGLEDEEVEMSVLQIDGCFNVGLIIEGIEVRPKN 261
WLEIEMGEFF SGLE++E+ M+V + + G I+EGIEVRPK+
Sbjct: 534 KWLEIEMGEFFNSGLEEDEIYMNVRETSDMWKHGFILEGIEVRPKH 671
>TC232233 weakly similar to GB|AAT41865.1|48310676|BT014882 At1g80110
{Arabidopsis thaliana;} , partial (47%)
Length = 687
Score = 192 bits (487), Expect = 2e-49
Identities = 106/197 (53%), Positives = 132/197 (66%), Gaps = 6/197 (3%)
Frame = +1
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPEGCIA ILS T+P D RLSL+S FRSAA SDAVW++FLPSD H+I+SQS S +
Sbjct: 25 LPEGCIANILSFTSPRDVCRLSLLSSTFRSAAQSDAVWNKFLPSDFHTILSQS-SSLSLP 201
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPES 123
SKK +L L P++ID+GKKS+QLDK GKKC+ML AR LFI W + Y WT+LP++
Sbjct: 202 SKKDLFLYLCQKPLLIDDGKKSYQLDKVYGKKCYMLSARNLFIVWGDTPRYWRWTSLPDA 381
Query: 124 RFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAPN------FHEVLVVLSV 177
RF +V LR + WLEI G INT LSP T YGA+LVFK PN F LV +S+
Sbjct: 382 RFSEVAELRSVCWLEIRGWINTGMLSPETLYGAYLVFK----PNPSGFYGFDYQLVEVSI 549
Query: 178 NNIGGHNITKIAYLNPD 194
GG N + +L+ +
Sbjct: 550 GIAGGENRKRNVFLDAE 600
>TC228247 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific lectin
PP2-like protein, partial (46%)
Length = 1124
Score = 191 bits (486), Expect = 2e-49
Identities = 112/263 (42%), Positives = 153/263 (57%), Gaps = 3/263 (1%)
Frame = +3
Query: 1 MELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLS-- 58
M+ LPE C++ ILS T+P DA R S+VS RS+ADSD +W F PSD IVS++L+
Sbjct: 72 MDTLPEDCVSKILSYTSPPDACRFSMVSSTLRSSADSDLLWRTFFPSDYSDIVSRALNPL 251
Query: 59 RANASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWT 118
N+SS + HP+++D G SF+LDK SGKK ++L AR L I W N Y W
Sbjct: 252 SLNSSSSYKHLFYALCHPLLLDGGNMSFKLDKSSGKKSYILSARQLSITWSNDPLYWSWR 431
Query: 119 TLPESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKIIDAP-NFHEVLVVLSV 177
+PESRF +V LR + WLEI G I T L+PNT Y +L+ K V +S+
Sbjct: 432 PVPESRFKEVAELRTVSWLEIQGKIGTRILTPNTSYVVYLIMKTSHREYGLDSVACEVSI 611
Query: 178 NNIGGHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISGLEDEEVEM 237
+ YL + + E + + + + P R DGW+EIEMGEFF G DEEV M
Sbjct: 612 AVDNKVKQSGRVYLCQNEKD-ENNLKKESIGIPMRREDGWMEIEMGEFFC-GEADEEVLM 785
Query: 238 SVLQIDGCFNVGLIIEGIEVRPK 260
S++++ GLI+EG+E+RPK
Sbjct: 786 SLMEVGYQLKGGLIVEGVEIRPK 854
>TC217153 similar to GB|AAS76702.1|45773796|BT012215 At1g56240 {Arabidopsis
thaliana;} , partial (13%)
Length = 635
Score = 126 bits (316), Expect(2) = 6e-44
Identities = 69/139 (49%), Positives = 87/139 (61%), Gaps = 4/139 (2%)
Frame = +3
Query: 44 FLPSDSHSIVSQSLSRANASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARA 103
F+PSD S +S LS +N SKK Y LSD P +ID G+KSFQL+K++ KKC+ML AR
Sbjct: 3 FIPSDFSSTISP-LSSSN--SKKDLYFTLSDRPTIIDQGRKSFQLEKRTAKKCYMLSARD 173
Query: 104 LFIAW----DNGDHYRCWTTLPESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLV 159
+ I W Y W +LPESRF +V L + W I G I T LSPNTQY AFLV
Sbjct: 174 ISITWAPTQGEASQYWEWKSLPESRFQEVARLYAVCWFNITGQIKTRVLSPNTQYAAFLV 353
Query: 160 FKIIDAPNFHEVLVVLSVN 178
F++IDA FH +LS++
Sbjct: 354 FQMIDASGFHHHPAMLSIS 410
Score = 68.9 bits (167), Expect(2) = 6e-44
Identities = 32/55 (58%), Positives = 42/55 (76%)
Frame = +1
Query: 207 VQGPSVRSDGWLEIEMGEFFISGLEDEEVEMSVLQIDGCFNVGLIIEGIEVRPKN 261
+Q P+VR D WLEIEMGEFF SGLE++E+ M+V + + G I+EGIEVRPK+
Sbjct: 415 LQRPNVRKDKWLEIEMGEFFNSGLEEDEIYMNVRETSDMWKHGFILEGIEVRPKH 579
>TC217151 similar to UP|Q6NPT8 (Q6NPT8) At2g02230, partial (6%)
Length = 514
Score = 126 bits (316), Expect(2) = 2e-33
Identities = 71/126 (56%), Positives = 83/126 (65%), Gaps = 5/126 (3%)
Frame = +3
Query: 4 LPEGCIAAILSRT-TPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANA 62
LPEGC+A ILS TP D RLSLVSK F SAAD D VWDRF+PSD S +S LS +N
Sbjct: 36 LPEGCVAHILSYICTPEDIVRLSLVSKAFYSAADYDTVWDRFIPSDFSSTISP-LSSSN- 209
Query: 63 SSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAW----DNGDHYRCWT 118
SKK Y LSD P +ID G+KSFQL+K++ KKC+ML AR + I W Y W
Sbjct: 210 -SKKDLYFTLSDRPTIIDQGRKSFQLEKRTAKKCYMLSARDISITWAPTQGEASQYWEWK 386
Query: 119 TLPESR 124
+LPESR
Sbjct: 387 SLPESR 404
Score = 33.9 bits (76), Expect(2) = 2e-33
Identities = 15/34 (44%), Positives = 22/34 (64%)
Frame = +2
Query: 161 KIIDAPNFHEVLVVLSVNNIGGHNITKIAYLNPD 194
K+IDA FH +LSV+N+GG +K L+P+
Sbjct: 401 KMIDASGFHHHPAMLSVSNVGGSRTSKYVCLDPN 502
>BI426662
Length = 421
Score = 102 bits (254), Expect = 2e-22
Identities = 65/135 (48%), Positives = 78/135 (57%), Gaps = 30/135 (22%)
Frame = +3
Query: 2 ELLPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQS---LS 58
E LPEGCIA I+S TTP DA LSLVS FRSA+ +D VW+RFLPSD +I+SQS +
Sbjct: 15 EHLPEGCIANIVSFTTPPDACVLSLVSSSFRSASVTDFVWERFLPSDYQAIISQSSKPST 194
Query: 59 RANASSKKRFYLDLSDHPVVIDNGKK---------------------------SFQLDKK 91
N SSKK YL L +P++ID GKK SF LDK
Sbjct: 195 LTNYSSKKDLYLHLCHNPLLIDAGKKVLQIKSFFLSIIIHVYLSLIGKLINHQSFALDKL 374
Query: 92 SGKKCFMLGARALFI 106
+GK C+ML AR+L I
Sbjct: 375 NGKICYMLSARSLSI 419
>BG652227
Length = 416
Score = 97.1 bits (240), Expect = 7e-21
Identities = 62/132 (46%), Positives = 72/132 (53%), Gaps = 33/132 (25%)
Frame = +3
Query: 4 LPEGCIAAILSRT-TPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANA 62
LPEGC+A ILS TP D RLSLVSK F SAAD D VWDRF+PSD S +S LS +N
Sbjct: 30 LPEGCVAHILSYICTPEDIVRLSLVSKAFYSAADYDTVWDRFIPSDFSSTIS-PLSSSN- 203
Query: 63 SSKKRFYLDLSDHPVVIDNGKK--------------------------------SFQLDK 90
SKK Y LSD P +ID G+K SFQL+K
Sbjct: 204 -SKKDLYFTLSDRPTIIDQGRKVRTLFLLACSDVFYGIPEMKCVVYVASLKFLQSFQLEK 380
Query: 91 KSGKKCFMLGAR 102
++ KKC+ML AR
Sbjct: 381 RTAKKCYMLSAR 416
>BQ610780
Length = 447
Score = 92.4 bits (228), Expect = 2e-19
Identities = 60/117 (51%), Positives = 70/117 (59%), Gaps = 4/117 (3%)
Frame = +3
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
LPEGCIA ILS TTPVDA RLS VS FRSAA+SD VWD FL SD S + S ++
Sbjct: 15 LPEGCIAKILSYTTPVDACRLS-VSIAFRSAAESDTVWDCFLLSDFTSFIPPS-----ST 176
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKK--CFMLGARALF--IAWDNGDHYRC 116
SK Y LSD P ++D G+KS QL K G+ CF L L+ I +D G C
Sbjct: 177 SKNDLYFTLSDLPTIMDQGRKSVQLAKGPGRSVTCFPLEIWPLYGVILFDIGSGQAC 347
Score = 46.2 bits (108), Expect = 2e-05
Identities = 21/47 (44%), Positives = 30/47 (63%)
Frame = +2
Query: 81 NGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPESRFPD 127
+G K + K++GKKC+ML AR L I W + + WT+LPES+ D
Sbjct: 227 SG*KECSIGKRTGKKCYMLSARNLAIIWGDTVRHWEWTSLPESKLVD 367
>TC228790
Length = 1215
Score = 91.7 bits (226), Expect = 3e-19
Identities = 71/274 (25%), Positives = 126/274 (45%), Gaps = 16/274 (5%)
Frame = +2
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVS--QSLSRAN 61
+PE C+A + TP + L+ +++ FR AA SD+VW+ LP + ++ R +
Sbjct: 185 IPESCVACVFLHLTPPEICNLARLNRAFRGAASSDSVWEAKLPRNYQDLLDLVPPPERHH 364
Query: 62 ASSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP 121
S K+ L P+ D+G K LD+ +GK C + A+A+ I + Y W
Sbjct: 365 RSLSKKDIFALLSRPLPFDHGHKEVWLDRVTGKVCMSISAKAMVITGIDDRRYWNWIPTE 544
Query: 122 ESRFPDVVVLRHLLWLEIHGMI------NTFSLSPNTQYGAF---LVFKIIDAPNFHE-- 170
ESRF V L+ + W E+ G + + ++LS G F L ++ + + H
Sbjct: 545 ESRFHTVAYLQQIWWFEVDGEVTFPFPADIYTLSFRLHLGRFSKRLGRRVCNYEHTHGWD 724
Query: 171 -VLVVLSVNNIGGHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISG 229
V + G + L+ E E++D + + + W++ ++GEF +SG
Sbjct: 725 IKPVKFEFSTSDGQQASCECCLD----ETEINDTYG-----NHKRGYWVDYKVGEFIVSG 877
Query: 230 LE-DEEVEMSVLQIDGCFNV-GLIIEGIEVRPKN 261
E +V S+ QID + GL ++ + + P +
Sbjct: 878 SEPTTQVRFSMKQIDCTHSKGGLCVDAVFIVPSD 979
>TC207581
Length = 1562
Score = 78.2 bits (191), Expect = 4e-15
Identities = 48/156 (30%), Positives = 78/156 (49%), Gaps = 2/156 (1%)
Frame = +2
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
+PE C+A +L P D +L+ +++ FR A+ +D +W+ LP + IV ++L +
Sbjct: 464 IPESCVALVLMYLDPPDICKLARLNRAFRDASSADFIWESKLPLNYKFIVEKALKDVSVE 643
Query: 64 S--KKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP 121
K+ Y L P DNG K LDK++G C + ++AL I + Y +
Sbjct: 644 QLGKRDIYARLC-RPNSFDNGTKEIWLDKRTGGVCLAISSQALRITGIDDRRYWSRISTE 820
Query: 122 ESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAF 157
ESRF V L+ + WLE+ ++ F P +Y F
Sbjct: 821 ESRFHTVAYLQQIWWLEVEDDVD-FQFPPG-KYSVF 922
>TC228248 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific lectin
PP2-like protein, partial (28%)
Length = 519
Score = 77.0 bits (188), Expect = 8e-15
Identities = 37/73 (50%), Positives = 45/73 (60%)
Frame = +3
Query: 89 DKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPESRFPDVVVLRHLLWLEIHGMINTFSL 148
DK SGKK ++L AR L I W N Y W +PESRF +V LR + WLEI G I T L
Sbjct: 15 DKSSGKKSYILSARQLSITWSNDPLYWSWRPVPESRFKEVAELRTVSWLEIQGKIGTRIL 194
Query: 149 SPNTQYGAFLVFK 161
+PNT Y +L+ K
Sbjct: 195 TPNTSYVVYLIMK 233
>TC207017
Length = 905
Score = 76.3 bits (186), Expect = 1e-14
Identities = 68/246 (27%), Positives = 116/246 (46%), Gaps = 6/246 (2%)
Frame = +3
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
+PE CI+++ P D +L+ V++ F A+ +D VW+ LP + ++ L N +
Sbjct: 189 IPESCISSLFMNLDPPDICKLARVNRAFHRASSADFVWESKLPPSYKFLANKVLGEENIA 368
Query: 64 S--KKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP 121
+ KK Y L P D G K LDK SG+ C + +++L I D R W +P
Sbjct: 369 TMTKKEIYAKLC-LPNRFDGGTKEVWLDKCSGQVCLFMSSKSLKIT--GIDDRRYWNYIP 539
Query: 122 --ESRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKI-IDAPNFHEVLVVLSVN 178
ESRF V L+ + W+E+ G + F P Y L+F++ + + V +V+
Sbjct: 540 TEESRFQSVAYLQQMWWVEVVGELE-FEF-PVGSYS--LIFRLQLGKASKRLGRRVCNVD 707
Query: 179 NIGGHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFI-SGLEDEEVEM 237
+ G +I I + S +G+ ++GP W+ +G+F + E ++
Sbjct: 708 QVHGWDIKPIRFQLSTS-DGQSSLSECYLRGPG----EWVYYNVGDFVVEKPKEPINIKF 872
Query: 238 SVLQID 243
S+ QID
Sbjct: 873 SLAQID 890
>TC207016
Length = 1310
Score = 75.5 bits (184), Expect = 2e-14
Identities = 71/268 (26%), Positives = 122/268 (45%), Gaps = 11/268 (4%)
Frame = +1
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
+PE CI++++ P + L+ V+K F A+ ++ VW+ LP + ++++ L N
Sbjct: 316 IPENCISSMMMNFDPQEICSLARVNKTFHRASSANFVWESKLPQNYKFLLNKVLGEQNLG 495
Query: 64 S--KKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLP 121
S KK Y L P D G K LD+ SG+ C + +++ I D R W +P
Sbjct: 496 SMTKKEIYAKLC-QPNFFDGGTKEVWLDRSSGQVCMFISSKSFKIT--GIDDRRYWNYIP 666
Query: 122 E-----SRFPDVVVLRHLLWLEIHGMINTFSLSPNTQYGAFLVFKI-IDAPNFHEVLVVL 175
RF V L+ + W+E+ G + F P Y F FK+ + P+ V
Sbjct: 667 TEESR*ERFKSVAYLQQMWWVEVIGELE-FEF-PKGNYSVF--FKLQLGKPSKRLGRRVC 834
Query: 176 SVNNIGGHNITKIAYLNPDSLEGEVHDRWDRVQGPSVRSDGWLEIEMGEFFISGLE-DEE 234
+++ + G +I + + S +G+ ++G S W +G+F I
Sbjct: 835 NLDQVHGWDIKPVRFQLSTS-DGQRSLSQCYLRG----SREWAYYHVGDFAIDKPNGPTN 999
Query: 235 VEMSVLQIDGCFNV--GLIIEGIEVRPK 260
+ S+ QID C + GL I+G+ + PK
Sbjct: 1000INFSLAQID-CTHTKGGLCIDGVVICPK 1080
>BE803206
Length = 404
Score = 71.6 bits (174), Expect = 3e-13
Identities = 60/156 (38%), Positives = 70/156 (44%), Gaps = 14/156 (8%)
Frame = +2
Query: 4 LPEGCIAAILSRT-TPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANA 62
LPEGC+A ILS TP D RLSLVSK F SAAD D VWDRF+PSD S +S LS +N
Sbjct: 29 LPEGCVAHILSYICTPEDIVRLSLVSKAFYSAADYDTVWDRFIPSDFSSTIS-PLSSSN- 202
Query: 63 SSKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTLPE 122
SKK Y L F +D+ KCF C +P+
Sbjct: 203 -SKKDLYFTL-----------PFFFIDELIFSKCF-----------------ACL*FVPQ 295
Query: 123 -------------SRFPDVVVLRHLLWLEIHGMINT 145
SRF +V L + W I G I T
Sbjct: 296 IIK*AIFELSYCTSRFQEVARLYAVCWFNITGQIKT 403
>AI440677
Length = 421
Score = 61.2 bits (147), Expect = 5e-10
Identities = 31/117 (26%), Positives = 55/117 (46%)
Frame = +1
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
+PE C+A + TP + L+ +++ FR AA +D+VW LP + ++ + +
Sbjct: 55 IPENCVARVFLHLTPPEICNLARLNRAFRGAASADSVWQTKLPRNYQDLLDLMPPERHRN 234
Query: 64 SKKRFYLDLSDHPVVIDNGKKSFQLDKKSGKKCFMLGARALFIAWDNGDHYRCWTTL 120
K+ L V D+G K LD+ + + C + A+A+ I + Y W L
Sbjct: 235 LSKKDIFALLSRAVPFDDGNKGVWLDRVTRRVCMSISAKAMSITGIDDRRYWTWFLL 405
>TC228791
Length = 584
Score = 57.4 bits (137), Expect = 7e-09
Identities = 30/105 (28%), Positives = 51/105 (48%), Gaps = 6/105 (5%)
Frame = +1
Query: 4 LPEGCIAAILSRTTPVDAARLSLVSKIFRSAADSDAVWDRFLPSDSHSIVSQSLSRANAS 63
+PE C+A + TP + L+ +++ FR AA SD+VW+ LP + ++ + S
Sbjct: 268 IPESCVACVFLHLTPPEICNLARLNRAFRGAASSDSVWEAKLPRNYQDLLDLVPPERHRS 447
Query: 64 SKKRFYLDLSDHPVVIDNGKK------SFQLDKKSGKKCFMLGAR 102
K+ L P+ D+G K LD+ +GK C + A+
Sbjct: 448 LSKKDIFALLSRPLPFDHGHKVGAALQEVWLDRVTGKVCMSISAK 582
>BG839963 weakly similar to GP|21592660|gb| phloem-specific lectin PP2-like
protein {Arabidopsis thaliana}, partial (11%)
Length = 707
Score = 47.0 bits (110), Expect(2) = 9e-07
Identities = 43/150 (28%), Positives = 66/150 (43%), Gaps = 29/150 (19%)
Frame = +3
Query: 143 INTFSLSPNTQYGAFLVFKIID--APNFHEVLVVLSVNNIGGHNITKIAYLNPDSLEGEV 200
I + +LS T YGA+LVFK A F+ V +SV + A E V
Sbjct: 75 IKSGTLSEKTLYGAYLVFKQRSGGAYGFYNQPVXVSVEGRRRTVYLEXAETPRRPREQIV 254
Query: 201 HDRWDRVQG-----------------------PSVRSDGWLEIEMGEFFISG---LEDEE 234
+ RV+ P RSDGW+E+E+G+ F G +++E
Sbjct: 255 PGIFXRVRSRFLDSFDAAPPPPPPNAKGGGEYPKERSDGWMEVELGDXFNVGGXKEKEKE 434
Query: 235 VEMSVLQI-DGCFNVGLIIEGIEVRPKNDN 263
VE+ V ++ G + G+ ++G E+RP N
Sbjct: 435 VEIGVYEVXSGGWKXGIFVQGXEIRPXPKN 524
Score = 22.7 bits (47), Expect(2) = 9e-07
Identities = 10/19 (52%), Positives = 12/19 (62%)
Frame = +1
Query: 123 SRFPDVVVLRHLLWLEIHG 141
SRF +V L + WLEI G
Sbjct: 16 SRFSEVAELVSVCWLEIKG 72
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.138 0.421
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,191,496
Number of Sequences: 63676
Number of extensions: 169190
Number of successful extensions: 771
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 740
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 745
length of query: 263
length of database: 12,639,632
effective HSP length: 95
effective length of query: 168
effective length of database: 6,590,412
effective search space: 1107189216
effective search space used: 1107189216
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0226.6


