

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
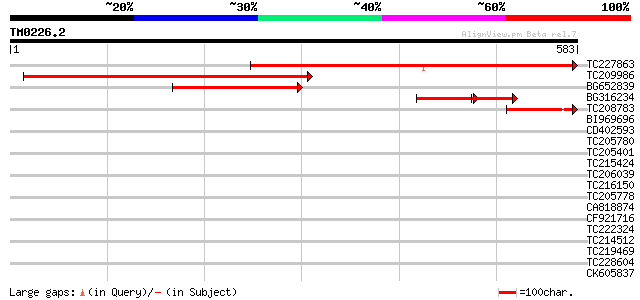
Query= TM0226.2
(583 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC227863 568 e-162
TC209986 weakly similar to UP|Q811Z9 (Q811Z9) N-terminal aceyltr... 549 e-156
BG652839 similar to GP|13365578|db P0686E09.16 {Oryza sativa (ja... 259 2e-69
BG316234 homologue to GP|1653666|dbj| chloride channel protein {... 99 1e-39
TC208783 87 2e-17
BI969696 38 0.014
CD402593 35 0.12
TC205780 weakly similar to UP|Q7TPA6 (Q7TPA6) Ab1-042, partial (... 34 0.16
TC205401 similar to GB|AAP37850.1|30725656|BT008491 At4g26630 {A... 33 0.35
TC215424 similar to UP|Q9MA04 (Q9MA04) F20B17.16, partial (26%) 33 0.35
TC206039 32 1.0
TC216150 weakly similar to UP|MNN4_YEAST (P36044) MNN4 protein, ... 31 1.3
TC205778 weakly similar to UP|Q7TPA6 (Q7TPA6) Ab1-042, partial (... 31 1.3
CA818874 31 1.8
CF921716 30 2.3
TC222324 30 2.3
TC214512 homologue to UP|O65334 (O65334) SAR DNA-binding protein... 30 2.3
TC219469 similar to UP|Q9ZS03 (Q9ZS03) RNA helicase (Fragment), ... 30 2.3
TC228604 weakly similar to UP|Q93X59 (Q93X59) Fructan 1-exohydro... 30 3.9
CK605837 30 3.9
>TC227863
Length = 1671
Score = 568 bits (1463), Expect = e-162
Identities = 292/367 (79%), Positives = 307/367 (83%), Gaps = 31/367 (8%)
Frame = +3
Query: 248 LRKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPA 307
LRKMT RTY+EMLKFQ +HSH+YFHKAAAGAIRCYIKLHDSPPKSTAEED+ MSKLLP+
Sbjct: 9 LRKMTXRTYVEMLKFQXGVHSHAYFHKAAAGAIRCYIKLHDSPPKSTAEEDDNMSKLLPS 188
Query: 308 QKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHGEKLLQVDDPLS 367
QKKKMRQKQRKAEARAKK AEEKNEE SASGVSKSGKRHVKPVDPDP+GEKLLQV+DPLS
Sbjct: 189 QKKKMRQKQRKAEARAKKEAEEKNEESSASGVSKSGKRHVKPVDPDPNGEKLLQVEDPLS 368
Query: 368 EAIKYLKLLQKNSPDSLETHLLSFELYTRKQKVLLTFQAVKQLLRLDAEHPDSHRCL--- 424
EA KYLKLLQKNSPDSLETHLLSFELYTRKQK+LL QAVKQLLRLDAEHPDSHRCL
Sbjct: 369 EATKYLKLLQKNSPDSLETHLLSFELYTRKQKILLALQAVKQLLRLDAEHPDSHRCLIKF 548
Query: 425 ----------------------------VCQLHEKTLFEANNSFLEKHKDSLMHRAAFAE 456
+ QLHEK+LFEANNSFLEKHKDSLMHRAAFAE
Sbjct: 549 FHKVGSMNAPVTDSEKLIWSVLEAERPTISQLHEKSLFEANNSFLEKHKDSLMHRAAFAE 728
Query: 457 TLYILDPNRKSEAVKLIEESTNNIVPRNGALGPIREWKLKDCIAVHKLLGTVLLDQDAAL 516
L+ILD NRKSEAVK +E+STNNIVPRNGALGPIREW L DCIAVHKLL TVL DQDA L
Sbjct: 729 ILHILDSNRKSEAVKFVEDSTNNIVPRNGALGPIREWNLTDCIAVHKLLETVLADQDAGL 908
Query: 517 RWKVRCAEYFPYSRYFEGSRSSASSNTALKQLSKNSENETLNHSVCTQNVGSITSNGKLE 576
RWKVRCAEYFPYS YFEG SSAS N+A QL KNSENE+LNHSV QNVGSITSNGKLE
Sbjct: 909 RWKVRCAEYFPYSTYFEGCHSSASPNSAFSQLRKNSENESLNHSVDGQNVGSITSNGKLE 1088
Query: 577 AFKQLAI 583
AFK L I
Sbjct: 1089AFKDLTI 1109
>TC209986 weakly similar to UP|Q811Z9 (Q811Z9) N-terminal aceyltransferase 1,
partial (18%)
Length = 1375
Score = 549 bits (1415), Expect = e-156
Identities = 265/297 (89%), Positives = 284/297 (95%)
Frame = +3
Query: 15 QRIPLDFLQGDKFREAADNYIRPLLTKGVPSLFSDLSSLYNHPGKADILEQLILDLEHSI 74
+RIPLDFLQGDKF+EAA+NYIRPLLTKG+PSLFSDLSSLYN PGKADILEQ+IL++E SI
Sbjct: 483 KRIPLDFLQGDKFQEAANNYIRPLLTKGIPSLFSDLSSLYNQPGKADILEQIILEIESSI 662
Query: 75 RTSGQYPGSMEKEPPSTLMWTLFLLAQHYDRRGQYELAIAKIDEAIEHTPTVIDLYSVKS 134
+T+ QYPG MEKEPPSTLMWTLFLLAQHYDRRGQYE+A+ KI+EAI+HTPTVIDLYSVKS
Sbjct: 663 KTTSQYPGGMEKEPPSTLMWTLFLLAQHYDRRGQYEIALFKINEAIDHTPTVIDLYSVKS 842
Query: 135 RILKHAGDFSAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTKEGDQH 194
RILKHAGD AAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTK+GDQH
Sbjct: 843 RILKHAGDLVAAAAFADEARCMDLADRYVNSECVKRMLQADQVALAEKTAVLFTKDGDQH 1022
Query: 195 NNLHDMQCMWYELASGESFFRQGDLGRALKKFLGVEKHYADINEDQFDFHSYCLRKMTLR 254
NNLHDMQCMWYELAS ES FRQG+LG ALKKFL VEKHYADI EDQFDFHSYCLRKMTLR
Sbjct: 1023NNLHDMQCMWYELASAESHFRQGNLGMALKKFLAVEKHYADITEDQFDFHSYCLRKMTLR 1202
Query: 255 TYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKK 311
TY+EMLKFQDQLHSH+YFHKAAAGAIRCYIKLHDSPPKSTAEED +MSKLLP+QKKK
Sbjct: 1203TYVEMLKFQDQLHSHAYFHKAAAGAIRCYIKLHDSPPKSTAEEDNDMSKLLPSQKKK 1373
>BG652839 similar to GP|13365578|db P0686E09.16 {Oryza sativa (japonica
cultivar-group)}, partial (19%)
Length = 404
Score = 259 bits (663), Expect = 2e-69
Identities = 124/134 (92%), Positives = 129/134 (95%)
Frame = +2
Query: 168 VKRMLQADQVALAEKTAVLFTKEGDQHNNLHDMQCMWYELASGESFFRQGDLGRALKKFL 227
VKRMLQADQVALAEKTAVLFTK+GDQHNNLHDMQCMWYELASGES+FRQGDLGRALKKFL
Sbjct: 2 VKRMLQADQVALAEKTAVLFTKDGDQHNNLHDMQCMWYELASGESYFRQGDLGRALKKFL 181
Query: 228 GVEKHYADINEDQFDFHSYCLRKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLH 287
VEKHYADI EDQFDFHSYCLRKMTL TY+EMLKFQDQLHSH+YFHKAAAGAIR YIKLH
Sbjct: 182 AVEKHYADITEDQFDFHSYCLRKMTLCTYVEMLKFQDQLHSHAYFHKAAAGAIRGYIKLH 361
Query: 288 DSPPKSTAEEDEEM 301
DSPPKSTAEED+ M
Sbjct: 362 DSPPKSTAEEDDNM 403
>BG316234 homologue to GP|1653666|dbj| chloride channel protein
{Synechocystis sp. PCC 6803}, partial (1%)
Length = 421
Score = 99.0 bits (245), Expect(2) = 1e-39
Identities = 48/64 (75%), Positives = 54/64 (84%)
Frame = +2
Query: 419 DSHRCLVCQLHEKTLFEANNSFLEKHKDSLMHRAAFAETLYILDPNRKSEAVKLIEESTN 478
++ R + QLH K+LFE NNSFLEKH+DSL HRAAF ETLYILDPNR+SEAVKLIE S N
Sbjct: 86 EAERQTISQLHGKSLFETNNSFLEKHEDSLTHRAAFGETLYILDPNRRSEAVKLIEGSPN 265
Query: 479 NIVP 482
NIVP
Sbjct: 266 NIVP 277
Score = 83.6 bits (205), Expect(2) = 1e-39
Identities = 38/47 (80%), Positives = 41/47 (86%)
Frame = +1
Query: 476 STNNIVPRNGALGPIREWKLKDCIAVHKLLGTVLLDQDAALRWKVRC 522
S N + RNG LGPIREWKL DC+AVHKLLGTVL+DQDAALRWKVRC
Sbjct: 280 SCNILFNRNGVLGPIREWKLIDCVAVHKLLGTVLVDQDAALRWKVRC 420
>TC208783
Length = 769
Score = 87.4 bits (215), Expect = 2e-17
Identities = 46/73 (63%), Positives = 52/73 (71%)
Frame = +2
Query: 511 DQDAALRWKVRCAEYFPYSRYFEGSRSSASSNTALKQLSKNSENETLNHSVCTQNVGSIT 570
DQDAA RWK+RCAE FPYS YFEG SSAS N+A Q+ K++E + NH V N S T
Sbjct: 2 DQDAASRWKMRCAELFPYSTYFEGICSSASPNSAFNQIRKSTETGSSNHWVGDHNAES-T 178
Query: 571 SNGKLEAFKQLAI 583
SNGKLEAFK L I
Sbjct: 179 SNGKLEAFKDLTI 217
>BI969696
Length = 725
Score = 37.7 bits (86), Expect = 0.014
Identities = 25/55 (45%), Positives = 30/55 (54%)
Frame = -2
Query: 529 SRYFEGSRSSASSNTALKQLSKNSENETLNHSVCTQNVGSITSNGKLEAFKQLAI 583
S EG S AS A Q+ K+SE+ + V N S TSNG LEAFK L+I
Sbjct: 721 STXXEGXXSCASHICAXTQMCKSSESGSYTQWVGDHNAES-TSNGNLEAFKDLSI 560
>CD402593
Length = 639
Score = 34.7 bits (78), Expect = 0.12
Identities = 20/47 (42%), Positives = 30/47 (63%), Gaps = 3/47 (6%)
Frame = -2
Query: 308 QKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVK---PVD 351
+KKK ++K++K + + KKG ++E+SA V K GK HV PVD
Sbjct: 395 EKKKKKKKKKKKKKKKKKGTRTNHKEISAVRV-KVGKPHVAVRGPVD 258
>TC205780 weakly similar to UP|Q7TPA6 (Q7TPA6) Ab1-042, partial (16%)
Length = 454
Score = 34.3 bits (77), Expect = 0.16
Identities = 17/62 (27%), Positives = 34/62 (54%)
Frame = +3
Query: 273 HKAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNE 332
++A A+ IKL + K+ + E+ K ++KKM +K++K E + KK ++ +
Sbjct: 264 YRARGDALIESIKLREEMEKAEKDRKEKREKKREKKEKKMEKKEKKKEKKEKKERRKEEK 443
Query: 333 EL 334
E+
Sbjct: 444 EI 449
>TC205401 similar to GB|AAP37850.1|30725656|BT008491 At4g26630 {Arabidopsis
thaliana;} , partial (16%)
Length = 912
Score = 33.1 bits (74), Expect = 0.35
Identities = 34/145 (23%), Positives = 59/145 (40%), Gaps = 11/145 (7%)
Frame = +3
Query: 289 SPPK----------STAEED-EEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSAS 337
SPPK S ++ED +E K+ +KK + ++K K +EEK E+++
Sbjct: 141 SPPKRAPKKPSSNLSKSDEDSDESPKVFSRKKKNEKGGKQKTATPTKSASEEKTEKVTRG 320
Query: 338 GVSKSGKRHVKPVDPDPHGEKLLQVDDPLSEAIKYLKLLQKNSPDSLETHLLSFELYTRK 397
K K +P D Q+ D + E +K + D L+ F++
Sbjct: 321 KGKK--KEKSRPSDN--------QLRDAICEILKEVNFNTATFTDILKKLAKQFDMDLTP 470
Query: 398 QKVLLTFQAVKQLLRLDAEHPDSHR 422
+K + ++L +L E D R
Sbjct: 471 RKASIKSMIQEELTKLADEADDEDR 545
>TC215424 similar to UP|Q9MA04 (Q9MA04) F20B17.16, partial (26%)
Length = 1030
Score = 33.1 bits (74), Expect = 0.35
Identities = 23/79 (29%), Positives = 36/79 (45%)
Frame = +3
Query: 268 SHSYFHKAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGA 327
+H HK +K ++ P AE DEE +KK+ +KQR+ E ++
Sbjct: 387 NHGIAHKQHKQQPPVPVKKMNNGPPGRAETDEEKR----LRKKREFEKQRQEEKHRQQLK 554
Query: 328 EEKNEELSASGVSKSGKRH 346
E +N L + + SGK H
Sbjct: 555 ESQNTVLQKTHMLSSGKGH 611
>TC206039
Length = 834
Score = 31.6 bits (70), Expect = 1.0
Identities = 20/77 (25%), Positives = 35/77 (44%)
Frame = -1
Query: 282 CYIKLHDSPPKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSK 341
CYI K ++ ++ K +KKK ++K++K + KK ++K G +K
Sbjct: 294 CYISALKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKN*KKKKKKKKKKKXXXGGGGGTK 115
Query: 342 SGKRHVKPVDPDPHGEK 358
++ K P P G K
Sbjct: 114 KXQKEGKKGAP-PGGAK 67
>TC216150 weakly similar to UP|MNN4_YEAST (P36044) MNN4 protein, partial (5%)
Length = 928
Score = 31.2 bits (69), Expect = 1.3
Identities = 25/93 (26%), Positives = 42/93 (44%), Gaps = 4/93 (4%)
Frame = +3
Query: 291 PKSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSK--SGKRHVK 348
PK + EE K P ++ + ++++ K E K+ EEK EE K K+
Sbjct: 402 PKKEEPKKEEEKKEEPKKEGEAKKEEEKKEEPKKE--EEKKEETKKEEEKKEEEKKKEAP 575
Query: 349 PVDPDPHGE--KLLQVDDPLSEAIKYLKLLQKN 379
P PDP E K + +P Y++ +++N
Sbjct: 576 PPPPDPVLELVKAYRAYNPHMTTYYYVQSMEEN 674
>TC205778 weakly similar to UP|Q7TPA6 (Q7TPA6) Ab1-042, partial (16%)
Length = 2225
Score = 31.2 bits (69), Expect = 1.3
Identities = 13/38 (34%), Positives = 25/38 (65%)
Frame = +3
Query: 315 KQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDP 352
K+ KA+ + +KG + +NE+L SG+++ + V V+P
Sbjct: 1380 KETKADGKLQKGEDCENEQLERSGITEEFDQPVTSVEP 1493
>CA818874
Length = 444
Score = 30.8 bits (68), Expect = 1.8
Identities = 14/50 (28%), Positives = 27/50 (54%)
Frame = +3
Query: 292 KSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKKGAEEKNEELSASGVSK 341
K ++ ++ K +KKK ++K++K + + KKG K + + GV K
Sbjct: 171 KKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKGGAPKKKPILTPGVGK 320
Score = 30.0 bits (66), Expect = 3.0
Identities = 15/58 (25%), Positives = 30/58 (50%)
Frame = +3
Query: 308 QKKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHGEKLLQVDDP 365
+KKK ++K++K + + KK ++K ++ K G KP+ G+K+ + P
Sbjct: 171 KKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKGGAPKKKPILTPGVGKKIFFLKGP 344
>CF921716
Length = 367
Score = 30.4 bits (67), Expect = 2.3
Identities = 20/58 (34%), Positives = 31/58 (52%), Gaps = 7/58 (12%)
Frame = +2
Query: 308 QKKKMRQKQR-------KAEARAKKGAEEKNEELSASGVSKSGKRHVKPVDPDPHGEK 358
QKKK ++K+R K + + KKG ++K +E + GK+ +P P P GEK
Sbjct: 59 QKKKTQKKKRGGPPTEKKKKKKKKKGGKKKKKE-GGKNKTPPGKKKRRP--PTPRGEK 223
>TC222324
Length = 698
Score = 30.4 bits (67), Expect = 2.3
Identities = 16/44 (36%), Positives = 24/44 (54%)
Frame = -1
Query: 400 VLLTFQAVKQLLRLDAEHPDSHRCLVCQLHEKTLFEANNSFLEK 443
+LL F A++ L LD+ + H C++CQ H L A +S K
Sbjct: 551 LLLNFSALE--LELDSTNKYMHSCMLCQRHPTRLTSAQHSVSSK 426
>TC214512 homologue to UP|O65334 (O65334) SAR DNA-binding protein-1, partial
(7%)
Length = 1535
Score = 30.4 bits (67), Expect = 2.3
Identities = 17/72 (23%), Positives = 34/72 (46%)
Frame = +2
Query: 247 CLRKMTLRTYLEMLKFQDQLHSHSYFHKAAAGAIRCYIKLHDSPPKSTAEEDEEMSKLLP 306
C +K+ +TY+ M ++QD L H Y + + + K + +++
Sbjct: 590 CSKKIPHKTYICMFRWQDTLDMHVYSKNTSQKYAHV*VAKYLMKKKKKKKTNKKKK*TND 769
Query: 307 AQKKKMRQKQRK 318
+KKK R+K++K
Sbjct: 770 NEKKKRRRKKKK 805
>TC219469 similar to UP|Q9ZS03 (Q9ZS03) RNA helicase (Fragment), partial
(66%)
Length = 1062
Score = 30.4 bits (67), Expect = 2.3
Identities = 20/78 (25%), Positives = 38/78 (48%), Gaps = 1/78 (1%)
Frame = +3
Query: 309 KKKMRQKQRKAEARAKKGAEEKNEELSASGVSKSGKRHV-KPVDPDPHGEKLLQVDDPLS 367
+ K R+KQRK +AKK A+EK + + + + K ++ ++ ++ L
Sbjct: 783 RDKSREKQRKKNLQAKKEAKEKEPKPQKPKKTPNAPTDMRKKTARQRRAQQTMEDEEELM 962
Query: 368 EAIKYLKLLQKNSPDSLE 385
+ + LK L+K + D E
Sbjct: 963 QEYRLLKKLKKGTIDENE 1016
>TC228604 weakly similar to UP|Q93X59 (Q93X59) Fructan 1-exohydrolase IIb
precursor , partial (33%)
Length = 828
Score = 29.6 bits (65), Expect = 3.9
Identities = 44/199 (22%), Positives = 84/199 (42%), Gaps = 11/199 (5%)
Frame = +1
Query: 329 EKNEELSASGVSKSGKRHVKPVDPDPHGEKLLQVDDPLSEAIKYLKLLQKNSPDSLETHL 388
E+ E+L VS +G++ + + G QVD + + L+ + +++HL
Sbjct: 142 EEVEKLRDKQVSITGEKLIGGSTIEVSGITASQVDVEVLFELPELENAEWLDESEVDSHL 321
Query: 389 LSFELYTRKQKVLLTFQAVKQLLRLDAEH----------PDSHRCLVCQLHEKTLFEANN 438
L E Y + ++ F + EH P+ + CL+C + +
Sbjct: 322 LCSEEYASRSGIIGPFGLLALASEDQTEHTAIFFRIYRAPNRYLCLMCS-------DQSR 480
Query: 439 SFLEKHKDSLMHRAAFAETLYILDPNRKSEAVK-LIEESTNNIVPRNGALGPIREWKLKD 497
S L + D + T++ +DPN K+ +++ LI+ S I+ G G I
Sbjct: 481 SSLRQDLDKTPY-----GTIFDIDPNVKTISLRSLIDRS---IIESFGEKGRI------- 615
Query: 498 CIAVHKLLGTVLLDQDAAL 516
CI ++ ++ +D+DA L
Sbjct: 616 CI-TSRVYPSLAIDKDAHL 669
>CK605837
Length = 498
Score = 29.6 bits (65), Expect = 3.9
Identities = 16/57 (28%), Positives = 30/57 (52%), Gaps = 4/57 (7%)
Frame = +3
Query: 292 KSTAEEDEEMSKLLPAQKKKMRQKQRKAEARAKK----GAEEKNEELSASGVSKSGK 344
K ++ ++ K +KKK ++K++K + + KK G +K + A G +K GK
Sbjct: 84 KKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKGPRGGTPKKKRKFGAPGGAKKGK 254
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.133 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,191,032
Number of Sequences: 63676
Number of extensions: 324873
Number of successful extensions: 1755
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 1729
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1747
length of query: 583
length of database: 12,639,632
effective HSP length: 102
effective length of query: 481
effective length of database: 6,144,680
effective search space: 2955591080
effective search space used: 2955591080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0226.2


