

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0226.11
(231 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
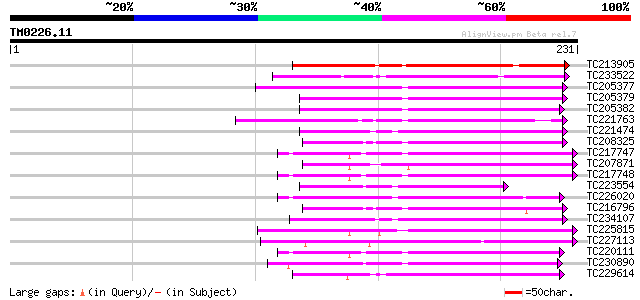
Score E
Sequences producing significant alignments: (bits) Value
TC213905 107 3e-24
TC233522 87 8e-18
TC205377 85 3e-17
TC205379 79 1e-15
TC205382 79 1e-15
TC221763 similar to UP|O49561 (O49561) Gibberellin 20-oxidase-li... 76 1e-14
TC221474 homologue to UP|O04281 (O04281) Gibberellin 20-oxidase ... 73 1e-13
TC208325 similar to GB|AAQ65160.1|34365697|BT010537 At4g10500 {A... 71 5e-13
TC217747 similar to UP|Q9FFF6 (Q9FFF6) Leucoanthocyanidin dioxyg... 70 8e-13
TC207871 similar to UP|O82017 (O82017) Adventitious rooting rela... 69 1e-12
TC217748 69 2e-12
TC223554 weakly similar to UP|Q9FXW0 (Q9FXW0) Gibberellin 3b-hyd... 67 5e-12
TC226020 similar to UP|Q9SP53 (Q9SP53) Flavanone 3-hydroxylase, ... 67 9e-12
TC216796 similar to GB|AAQ65160.1|34365697|BT010537 At4g10500 {A... 66 1e-11
TC234107 similar to UP|G2O2_PEA (Q9XHM5) Gibberellin 2-beta-diox... 65 2e-11
TC225815 65 2e-11
TC227113 65 2e-11
TC220111 similar to PIR|T49209|T49209 leucoanthocyanidin dioxyge... 64 4e-11
TC230890 similar to UP|Q8LA31 (Q8LA31) Naringenin 3-dioxygenase ... 64 8e-11
TC229614 similar to UP|O48631 (O48631) Ethylene-forming-enzyme-l... 64 8e-11
>TC213905
Length = 530
Score = 107 bits (268), Expect = 3e-24
Identities = 59/113 (52%), Positives = 72/113 (63%)
Frame = +1
Query: 116 DTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWS 175
+ ++G HTD + TIL Q+ V+ L V T +G WI SS + SF+VM D AWS
Sbjct: 1 EPQLGFVAHTDKSFTTILHQNHVNALMVETTNGN-WIDVDFSSPT-SFVVMAGDALMAWS 174
Query: 176 NGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFD 228
N R+ SP H VMM GNE RYS LF +G I+K PEEL+DEEHPL YKPFD
Sbjct: 175 NDRIKSPNHMVMMNGNETRYSLGLFAFYRG--ILKVPEELIDEEHPLQYKPFD 327
>TC233522
Length = 800
Score = 86.7 bits (213), Expect = 8e-18
Identities = 49/121 (40%), Positives = 69/121 (56%)
Frame = +1
Query: 108 KYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMM 167
KY+ + +T +G+ HTD+ LTIL Q +++GLE+ KDG +W +K + V+
Sbjct: 22 KYRTXKGGETNLGVHPHTDSGFLTILNQ-KLNGLEIQLKDG-EW--FKVDASPNMXAVLA 189
Query: 168 NDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPF 227
D F WSN R+ H+V M RY L + G +++ EELVDEEHPL YKPF
Sbjct: 190 GDAFMVWSNDRIRGCVHQVFMNSKVDRYCLGL--LSYAGKVMEPEEELVDEEHPLRYKPF 363
Query: 228 D 228
D
Sbjct: 364 D 366
>TC205377
Length = 1345
Score = 84.7 bits (208), Expect = 3e-17
Identities = 44/127 (34%), Positives = 68/127 (52%)
Frame = +3
Query: 101 TYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSES 160
T F++ Y D +G+ H D LTIL Q +V GLEV K ++WI KP+ ++
Sbjct: 681 TSFIRLNHYPPCPYPDLALGVGRHKDPGALTILAQDEVGGLEVRRKADQEWIRVKPTPDA 860
Query: 161 QSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEH 220
+I+ + D WSN S HRV++ + R+S FF P +K EEL++E++
Sbjct: 861 --YIINIGDTVQVWSNDAYESVDHRVVVNSEKERFSIPFFFFPAHDTEVKPLEELINEQN 1034
Query: 221 PLLYKPF 227
P Y+P+
Sbjct: 1035PSKYRPY 1055
>TC205379
Length = 774
Score = 79.3 bits (194), Expect = 1e-15
Identities = 40/109 (36%), Positives = 60/109 (54%)
Frame = +2
Query: 119 VGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGR 178
+G+ H D LT+L Q +V GLEV K ++WI KP+ ++ +I+ + D WSN
Sbjct: 161 LGVGRHKDGGALTVLAQDEVGGLEVKRKADQEWIRVKPTPDA--YIINVGDLIQVWSNDA 334
Query: 179 LHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPF 227
S HRVM+ + R+S FF P +K EEL +E +P Y+P+
Sbjct: 335 YESVEHRVMVNSEKERFSIPFFFNPAHDIEVKPLEELTNEHNPSKYRPY 481
>TC205382
Length = 613
Score = 79.3 bits (194), Expect = 1e-15
Identities = 40/108 (37%), Positives = 61/108 (56%)
Frame = +2
Query: 119 VGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGR 178
+G+ H D LTIL Q +V GLEV K ++WI KP+ ++ +I+ + D WSN
Sbjct: 296 LGVARHKDPGALTILAQDEVGGLEVKRKADQEWIGVKPTLDA--YIINVGDIIQVWSNDA 469
Query: 179 LHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKP 226
S HRV++ + R+S FF P +K EEL++E++P Y+P
Sbjct: 470 YESVEHRVVVNSEKERFSIPFFFYPAHETEVKPLEELINEQNPSKYRP 613
>TC221763 similar to UP|O49561 (O49561) Gibberellin 20-oxidase-like protein,
partial (54%)
Length = 782
Score = 76.3 bits (186), Expect = 1e-14
Identities = 51/135 (37%), Positives = 69/135 (50%)
Frame = +1
Query: 93 FRYASQPITYFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWI 152
F+ P T +L+ +Y GL HTD+ LTIL+Q QV GL+ L KD KWI
Sbjct: 43 FKENCLPNTCYLRLNRYPPCPIGFGIHGLMPHTDSDFLTILYQDQVGGLQ-LVKDS-KWI 216
Query: 153 SYKPSSESQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAP 212
+ KP+ ++ I+ + D F AWSNG S HRVM R+S A FF P +I++
Sbjct: 217 AVKPNPDA--LIINIGDLFQAWSNGVYKSVEHRVMTNPKLERFSMAYFFCPSNDTVIESC 390
Query: 213 EELVDEEHPLLYKPF 227
E P Y+ F
Sbjct: 391 RE------PSFYRKF 417
>TC221474 homologue to UP|O04281 (O04281) Gibberellin 20-oxidase , partial
(34%)
Length = 410
Score = 72.8 bits (177), Expect = 1e-13
Identities = 43/109 (39%), Positives = 56/109 (50%)
Frame = +3
Query: 119 VGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGR 178
+G H D LTIL Q QV GL+V + +W S KP + +F+V + D F A SNGR
Sbjct: 3 LGTGPHCDPTSLTILHQDQVGGLQVCVDN--EWHSIKP--DLNAFVVNVGDTFMALSNGR 170
Query: 179 LHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPF 227
S HR ++ R S A F P+ ++ P ELVD P LY F
Sbjct: 171 YKSCLHRAVVNSQTTRKSLAFFLCPRSDKVVSPPCELVDNLSPRLYPDF 317
>TC208325 similar to GB|AAQ65160.1|34365697|BT010537 At4g10500 {Arabidopsis
thaliana;} , partial (32%)
Length = 640
Score = 70.9 bits (172), Expect = 5e-13
Identities = 43/108 (39%), Positives = 57/108 (51%)
Frame = +3
Query: 120 GLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRL 179
GL H D +TIL Q+QV GL+VL DG KW++ P + FIV + D SN R
Sbjct: 54 GLPAHADPNAITILLQNQVPGLQVL-HDG-KWLTVNPVPNT--FIVNIADQIQVISNDRY 221
Query: 180 HSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPF 227
S HR ++ + R S F+ P +IK +LVD+EHP Y F
Sbjct: 222 KSVLHRALVNCEKERMSIPTFYCPSPDALIKPAPQLVDKEHPAQYTNF 365
>TC217747 similar to UP|Q9FFF6 (Q9FFF6) Leucoanthocyanidin dioxygenase-like
protein (AT5g05600/MOP10_14), partial (53%)
Length = 713
Score = 70.1 bits (170), Expect = 8e-13
Identities = 46/123 (37%), Positives = 65/123 (52%), Gaps = 1/123 (0%)
Frame = +1
Query: 110 KGPQTSDTKVGLDTHTDTAILTILFQSQ-VDGLEVLTKDGRKWISYKPSSESQSFIVMMN 168
K PQ D GL H+D +TIL V GL+V + G +WI+ KP + FI+ +
Sbjct: 187 KCPQP-DLTFGLSPHSDPGGMTILLPDDFVSGLQV--RRGDEWITVKPVPNA--FIINIG 351
Query: 169 DCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFD 228
D SN S HRV++ N+ R S ALF+ P+ +I+ +ELV EE P LY P
Sbjct: 352 DQIQVLSNAIYKSVEHRVIVNSNKDRVSLALFYNPRSDLLIQPAKELVTEEKPALYSPMT 531
Query: 229 RED 231
++
Sbjct: 532 YDE 540
>TC207871 similar to UP|O82017 (O82017) Adventitious rooting related
oxygenase, partial (41%)
Length = 611
Score = 69.3 bits (168), Expect = 1e-12
Identities = 45/114 (39%), Positives = 58/114 (50%), Gaps = 2/114 (1%)
Frame = +2
Query: 120 GLDTHTDTAILTILFQSQ-VDGLEVLTKDGRKWISYKPSSESQ-SFIVMMNDCFHAWSNG 177
G+ HTD+ LTIL + V GLEVL S+ P S +V + D WSNG
Sbjct: 26 GVQIHTDSGFLTILQDDENVGGLEVLHSS----TSFVPIPLFPGSLLVNLGDIARVWSNG 193
Query: 178 RLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFDRED 231
R + HRV R+S A F + ++AP ELVD +HP LY+PF ED
Sbjct: 194 RFCNLTHRVQCKEATKRFSIATFMIAPRNRNVEAPAELVDHDHPRLYQPFIYED 355
>TC217748
Length = 1003
Score = 68.9 bits (167), Expect = 2e-12
Identities = 44/123 (35%), Positives = 64/123 (51%), Gaps = 1/123 (0%)
Frame = +1
Query: 110 KGPQTSDTKVGLDTHTDTAILTILFQSQ-VDGLEVLTKDGRKWISYKPSSESQSFIVMMN 168
K PQ D GL H+D +TIL V GL+V + G +W+ KP + F++ +
Sbjct: 352 KCPQP-DLTFGLSPHSDPGGMTILLSDDFVSGLQV--RRGDEWVIVKPVPNA--FVINIG 516
Query: 169 DCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPFD 228
D SN S HRV++ N+ R S ALF+ P+ +I+ +ELV EE P LY P
Sbjct: 517 DQIQVLSNAIYKSVEHRVIVNSNKDRVSLALFYNPRSDLLIQPAKELVTEERPALYSPMT 696
Query: 229 RED 231
++
Sbjct: 697 YDE 705
>TC223554 weakly similar to UP|Q9FXW0 (Q9FXW0) Gibberellin 3b-hydroxylase
No3, partial (36%)
Length = 614
Score = 67.4 bits (163), Expect = 5e-12
Identities = 33/85 (38%), Positives = 48/85 (55%)
Frame = +3
Query: 119 VGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGR 178
+GL HTDT++ TIL QS++ GL++ K+G++W+ P + +V D H SN R
Sbjct: 177 MGLAPHTDTSLFTILHQSRITGLQIF-KEGKEWVPVHP--HPNTLVVHTGDLLHIISNAR 347
Query: 179 LHSPFHRVMMTGNEARYSAALFFVP 203
S HRV + RYS A F+ P
Sbjct: 348 FRSALHRVTVNSTRERYSVAYFYSP 422
>TC226020 similar to UP|Q9SP53 (Q9SP53) Flavanone 3-hydroxylase, partial
(93%)
Length = 1441
Score = 66.6 bits (161), Expect = 9e-12
Identities = 39/117 (33%), Positives = 62/117 (52%)
Frame = +3
Query: 110 KGPQTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMND 169
K PQ D +GL HTD +T+L Q QV GL+ +G+ WI+ +P +F+V + D
Sbjct: 678 KCPQP-DLTLGLKRHTDPGTITLLLQDQVGGLQATRDNGKTWITVQP--VEAAFVVNLGD 848
Query: 170 CFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKP 226
H SNGR + H+ ++ N +R S A F P + P ++ + E P++ +P
Sbjct: 849 HAHYLSNGRFKNADHQAVVNSNHSRLSIATFQNPAPNATV-YPLKIREGEKPVMEEP 1016
>TC216796 similar to GB|AAQ65160.1|34365697|BT010537 At4g10500 {Arabidopsis
thaliana;} , partial (72%)
Length = 1454
Score = 66.2 bits (160), Expect = 1e-11
Identities = 41/109 (37%), Positives = 57/109 (51%), Gaps = 1/109 (0%)
Frame = +1
Query: 120 GLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAWSNGRL 179
GL HTD ++TIL Q +V GL+VL KDG KW++ P + F+V + D SN +
Sbjct: 745 GLPGHTDPTVITILLQDEVPGLQVL-KDG-KWVAVNPIPNT--FVVNVGDQIQVISNDKY 912
Query: 180 HSPFHRVMMTGNEARYSAALFFVPKGGCII-KAPEELVDEEHPLLYKPF 227
S HR ++ N+ R S F+ P II AP+ + HP Y F
Sbjct: 913 KSVLHRAVVNCNKDRISIPTFYFPSNDAIIGPAPQLIHHHHHPPQYNNF 1059
>TC234107 similar to UP|G2O2_PEA (Q9XHM5) Gibberellin 2-beta-dioxygenase 2
(Gibberellin 2-beta-hydroxylase 2) (Gibberellin
2-oxidase 2) (GA 2-oxidase 2) , partial (40%)
Length = 687
Score = 65.5 bits (158), Expect = 2e-11
Identities = 40/113 (35%), Positives = 55/113 (48%)
Frame = +3
Query: 115 SDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAW 174
++ +G H+D ILTI+ + VDGL++ T DG WI P + VM+ D
Sbjct: 33 NNNNIGXXEHSDPQILTIMRSNNVDGLQISTHDGL-WIPVPP--DPNEXXVMVGDALQVL 203
Query: 175 SNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKPF 227
+NGR S HRV+ +AR S F P I +V +P LYKPF
Sbjct: 204 TNGRFASVRHRVLTNTTKARMSMMYFAAPPLNRWITPLPMMVTPHNPSLYKPF 362
>TC225815
Length = 945
Score = 65.5 bits (158), Expect = 2e-11
Identities = 40/132 (30%), Positives = 66/132 (49%), Gaps = 2/132 (1%)
Frame = +3
Query: 102 YFLKTMKYKGPQTSDTKVGLDTHTDTAILTILFQSQ-VDGLEVLTKDGR-KWISYKPSSE 159
+ L+T+K+ G H+DT +T+L + V GLE++ G K + P +
Sbjct: 153 FILRTIKFSFTPDVIGSTGAQLHSDTGFITLLQDDEHVSGLEMMDDFGSFKAVPPIPGA- 329
Query: 160 SQSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEE 219
F+ ++ D H WSNG+ + HRV+ RYS F + ++AP++LV+ +
Sbjct: 330 ---FLCIVGDVGHVWSNGKFWNARHRVICKETGTRYSFGAFMLSPRDGNVEAPKKLVEVD 500
Query: 220 HPLLYKPFDRED 231
H Y+PF ED
Sbjct: 501 HVQRYRPFKYED 536
>TC227113
Length = 1270
Score = 65.5 bits (158), Expect = 2e-11
Identities = 41/131 (31%), Positives = 60/131 (45%), Gaps = 2/131 (1%)
Frame = +2
Query: 103 FLKTMKYKGPQTSDTKV-GLDTHTDTAILTILFQSQVDGLEVLT-KDGRKWISYKPSSES 160
FL+ + Y G SD ++ G H+D ++T+L V GL++ K + +
Sbjct: 512 FLRLLHYPGELGSDEQICGASPHSDYGMITLLMTDGVPGLQICKDKVNQPQVWEDVPHVE 691
Query: 161 QSFIVMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEH 220
+ IV + D W+N S HRVM TG E RYS A FF P C+++ E E
Sbjct: 692 GALIVNIGDMMERWTNCLYRSTLHRVMPTGKE-RYSVAFFFDPASDCVVECFESCCSESS 868
Query: 221 PLLYKPFDRED 231
P + P D
Sbjct: 869 PPRFSPIRSGD 901
>TC220111 similar to PIR|T49209|T49209 leucoanthocyanidin dioxygenase-like
protein - Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (43%)
Length = 786
Score = 64.3 bits (155), Expect = 4e-11
Identities = 43/118 (36%), Positives = 62/118 (52%), Gaps = 1/118 (0%)
Frame = +3
Query: 110 KGPQTSDTKVGLDTHTDTAILTILFQSQ-VDGLEVLTKDGRKWISYKPSSESQSFIVMMN 168
K PQ D +GL H+D +TIL + V GL+V + G WI+ KP + FI+ M
Sbjct: 186 KCPQP-DLTLGLSPHSDPGGMTILLPDENVSGLQV--RRGEDWITVKPVPNA--FIINMG 350
Query: 169 DCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKP 226
D SN S HRV++ ++ R S A F+ P+ I+ +ELV ++ P LY P
Sbjct: 351 DQIQVLSNAIYKSIEHRVIVNSDKDRVSLAFFYNPRSDIPIQPAKELVTKDRPALYPP 524
>TC230890 similar to UP|Q8LA31 (Q8LA31) Naringenin 3-dioxygenase like
protein, partial (64%)
Length = 677
Score = 63.5 bits (153), Expect = 8e-11
Identities = 39/121 (32%), Positives = 62/121 (51%), Gaps = 1/121 (0%)
Frame = +1
Query: 106 TMKYKGP-QTSDTKVGLDTHTDTAILTILFQSQVDGLEVLTKDGRKWISYKPSSESQSFI 164
T+ Y P D +GL +H+D +T+L Q V GL+VL K KW++ +P S++ +
Sbjct: 79 TISYYPPCPEPDLTLGLQSHSDMGAITLLIQDDVGGLQVL-KGSDKWVTVQPLSDA--VL 249
Query: 165 VMMNDCFHAWSNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLY 224
V++ D +NG+ S HR + + AR S A F P I EL+++ P Y
Sbjct: 250 VLLADQTEIITNGKYRSCEHRAITNPDRARLSVATFHDPAKTAKISPASELINDSSPAKY 429
Query: 225 K 225
+
Sbjct: 430 R 432
>TC229614 similar to UP|O48631 (O48631) Ethylene-forming-enzyme-like
dioxygenase, partial (52%)
Length = 1289
Score = 63.5 bits (153), Expect = 8e-11
Identities = 38/112 (33%), Positives = 61/112 (53%), Gaps = 1/112 (0%)
Frame = +1
Query: 116 DTKVGLDTHTDTAILTILFQS-QVDGLEVLTKDGRKWISYKPSSESQSFIVMMNDCFHAW 174
D +G+ H D + +T L Q +V+GL+VL D +W +K + ++ + D
Sbjct: 673 DHVLGVKPHADGSTITFLLQDKEVEGLQVLKDD--QW--FKVPIIPDALLINVGDQIEIM 840
Query: 175 SNGRLHSPFHRVMMTGNEARYSAALFFVPKGGCIIKAPEELVDEEHPLLYKP 226
SNG SP HRV++ + R + A+F VP IK ++LV+E P+LY+P
Sbjct: 841 SNGIFRSPVHRVVINKAKERLTVAMFCVPDSEKEIKPVDKLVNESRPVLYRP 996
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.135 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,344,350
Number of Sequences: 63676
Number of extensions: 140010
Number of successful extensions: 761
Number of sequences better than 10.0: 122
Number of HSP's better than 10.0 without gapping: 720
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 726
length of query: 231
length of database: 12,639,632
effective HSP length: 94
effective length of query: 137
effective length of database: 6,654,088
effective search space: 911610056
effective search space used: 911610056
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0226.11


