

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0225.8
(423 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
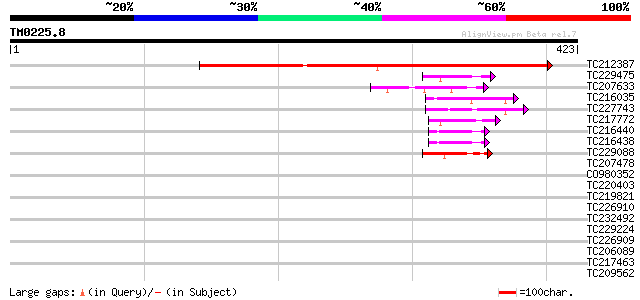
Sequences producing significant alignments: (bits) Value
TC212387 weakly similar to UP|Q9ZW89 (Q9ZW89) F5A8.9 protein, pa... 294 4e-80
TC229475 similar to UP|Q944Q9 (Q944Q9) AT4g31450/F3L17_20, parti... 43 2e-04
TC207633 similar to UP|Q94AB7 (Q94AB7) AT5g41350/MYC6_6, partial... 43 3e-04
TC216035 similar to GB|AAP13393.1|30023720|BT006285 At4g27470 {A... 43 3e-04
TC227743 similar to PIR|E86326|E86326 protein F18O14.3 [imported... 42 4e-04
TC217772 similar to GB|AAM51588.1|21436021|AY116954 AT5g19430/F7... 42 4e-04
TC216440 similar to UP|Q8GUU2 (Q8GUU2) RES protein, partial (68%) 42 5e-04
TC216438 homologue to UP|Q8GUU2 (Q8GUU2) RES protein, partial (14%) 42 5e-04
TC229088 similar to UP|Q40221 (Q40221) Protein containing C-term... 42 7e-04
TC207478 similar to UP|Q6R567 (Q6R567) Ring domain containing pr... 41 0.001
CO980352 41 0.001
TC220403 similar to UP|Q7X7W8 (Q7X7W8) Makorin ring-zinc-finger ... 40 0.002
TC219821 weakly similar to UP|PJA1_MOUSE (O55176) Ubiquitin prot... 40 0.002
TC226910 weakly similar to UP|Q8LPJ6 (Q8LPJ6) Pspzf zinc finger ... 40 0.002
TC232492 weakly similar to UP|Q8VYW5 (Q8VYW5) AT5g45290/K9E15_7,... 40 0.002
TC229224 weakly similar to UP|Q9LUR1 (Q9LUR1) RING zinc finger p... 40 0.002
TC226909 similar to PIR|H96764|H96764 protein RING zinc finger p... 40 0.002
TC206089 UP|Q39902 (Q39902) C-terminal zinc-finger (Fragment), c... 40 0.002
TC217463 similar to PIR|E86378|E86378 protein F21J9.10 [imported... 40 0.002
TC209562 similar to UP|Q8GT74 (Q8GT74) NEP1-interacting protein ... 40 0.003
>TC212387 weakly similar to UP|Q9ZW89 (Q9ZW89) F5A8.9 protein, partial (16%)
Length = 1107
Score = 294 bits (753), Expect = 4e-80
Identities = 161/275 (58%), Positives = 189/275 (68%), Gaps = 11/275 (4%)
Frame = +1
Query: 142 VLEDSCLMKKYEETSLHSSRRLKKVKRNISCDNEVSTAAGPSRKEKKFVKKVVDEVVSDP 201
VLEDSCLMKK+EE+S +SSR ++ KR+I E +T A PSRK ++ +VD+VV P
Sbjct: 7 VLEDSCLMKKHEESSSYSSRLSRRGKRHICNGKENTTVARPSRKGRRLASDLVDKVVLGP 186
Query: 202 VILDLSPDDRLCRMDRDRLHTEAAATSSLSGGVDVNDLQENSSGPDARLCNQNRAMNGSS 261
ILDL DD L RMD + T+A ATSSLS GV+ N + +NS GP+A L NQ+R + G S
Sbjct: 187 SILDLITDDHLFRMDSQQ--TDAEATSSLSVGVNNNIILQNSEGPNAGLSNQSRTIIGGS 360
Query: 262 DGIEQITDSNQI----------EDPQTVEQTSIDLCSSDG-KFIDGDQVDNVAGSPISNE 310
+ IEQI DS+ I EDP V QTSIDLCSSD KF DGDQ DNVAG P S E
Sbjct: 361 NDIEQIKDSSHISKPRNSTSFIEDPLPVPQTSIDLCSSDDEKFTDGDQADNVAGLPTSTE 540
Query: 311 WSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVEHA 370
SCVIC +FSST G+LPC H FC CIQ WA SM K STCPLCKASF K VEHA
Sbjct: 541 MSCVICLTEFSSTRGILPCGHRFCFPCIQNWADHTTSMRKTSTCPLCKASFMMIKKVEHA 720
Query: 371 ATADKELNSQTVPCDNLASVTFISMDQELPDNSFE 405
ATAD+++ SQT+PCDN ASV FI +DQ LPDN FE
Sbjct: 721 ATADQKIYSQTIPCDNSASVIFIPVDQNLPDNIFE 825
>TC229475 similar to UP|Q944Q9 (Q944Q9) AT4g31450/F3L17_20, partial (21%)
Length = 741
Score = 43.1 bits (100), Expect = 2e-04
Identities = 20/56 (35%), Positives = 32/56 (56%), Gaps = 2/56 (3%)
Frame = +2
Query: 309 NEWSCVICFEDF--SSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFA 362
++ C IC E++ + +G L CEH + + CIQ+W + + CP+CKAS A
Sbjct: 284 DDTKCSICQEEYVAADEVGSLQCEHAYHVACIQQWLQLK------NWCPICKASVA 433
>TC207633 similar to UP|Q94AB7 (Q94AB7) AT5g41350/MYC6_6, partial (33%)
Length = 825
Score = 42.7 bits (99), Expect = 3e-04
Identities = 31/96 (32%), Positives = 49/96 (50%), Gaps = 8/96 (8%)
Frame = +3
Query: 270 SNQIEDPQTVE---QTSIDLCSSDGKFIDGDQVDNVAGSPIS---NEWSCVICFEDFSST 323
S ++E+P+ E QT ++L S+ G I+ + +G PI E +C IC E++ +
Sbjct: 477 SAKLEEPKESECKVQTDLELDSAKGSEIELAK----SGKPIDLVEEEDACPICLEEYDAE 644
Query: 324 LGVLP--CEHTFCLTCIQKWAHQRISMGKISTCPLC 357
L C+H F L CI +W M + TCP+C
Sbjct: 645 NPKLATNCDHHFHLACILEW------MERSETCPVC 734
>TC216035 similar to GB|AAP13393.1|30023720|BT006285 At4g27470 {Arabidopsis
thaliana;} , partial (32%)
Length = 966
Score = 42.7 bits (99), Expect = 3e-04
Identities = 26/77 (33%), Positives = 38/77 (48%), Gaps = 8/77 (10%)
Frame = +1
Query: 311 WSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAH---QRISMGKISTCPLCKASFAWYKIV 367
+ C IC DF+ V C H +C CI KW H ++ + CP+CKA + +V
Sbjct: 199 FDCNICL-DFAHEPVVTLCGHLYCWPCIYKWLHVQSDSLAPDEHPQCPVCKADISNSTMV 375
Query: 368 E-----HAATADKELNS 379
HAATA+ + +S
Sbjct: 376 PLYGRGHAATAEGKTSS 426
>TC227743 similar to PIR|E86326|E86326 protein F18O14.3 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(73%)
Length = 1212
Score = 42.4 bits (98), Expect = 4e-04
Identities = 25/79 (31%), Positives = 37/79 (46%), Gaps = 2/79 (2%)
Frame = +2
Query: 311 WSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKIVE-- 368
+ C ICFE + L C H FC C+ KW H + CP+CKA K+V
Sbjct: 212 FECNICFELAQDPIITL-CGHLFCWPCLYKWLHFH---SQSRECPVCKALVEEEKLVPLY 379
Query: 369 HAATADKELNSQTVPCDNL 387
+ + S+++P DN+
Sbjct: 380 GRGKSSTDPRSKSIPGDNI 436
>TC217772 similar to GB|AAM51588.1|21436021|AY116954 AT5g19430/F7K24_180
{Arabidopsis thaliana;} , partial (61%)
Length = 1136
Score = 42.4 bits (98), Expect = 4e-04
Identities = 21/56 (37%), Positives = 27/56 (47%), Gaps = 2/56 (3%)
Frame = +2
Query: 313 CVICFEDF--SSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFAWYKI 366
C IC + T V CEH +C+TCI WA R + TCP CK F + +
Sbjct: 311 CAICLDKIVLQETALVKGCEHAYCVTCILHWATYREKV----TCPQCKHPFEFLNV 466
>TC216440 similar to UP|Q8GUU2 (Q8GUU2) RES protein, partial (68%)
Length = 1655
Score = 42.0 bits (97), Expect = 5e-04
Identities = 21/46 (45%), Positives = 25/46 (53%)
Frame = +1
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCK 358
C+ +ED + L VLPC H F TCI KW +TCPLCK
Sbjct: 1117 CISSYED-GAELHVLPCNHHFHSTCIVKWLKMN------ATCPLCK 1233
>TC216438 homologue to UP|Q8GUU2 (Q8GUU2) RES protein, partial (14%)
Length = 650
Score = 42.0 bits (97), Expect = 5e-04
Identities = 21/46 (45%), Positives = 25/46 (53%)
Frame = +3
Query: 313 CVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCK 358
C+ +ED + L VLPC H F TCI KW +TCPLCK
Sbjct: 147 CISSYED-GAELHVLPCNHHFHSTCIVKWLKMN------ATCPLCK 263
>TC229088 similar to UP|Q40221 (Q40221) Protein containing C-terminal
RING-finger, partial (63%)
Length = 1240
Score = 41.6 bits (96), Expect = 7e-04
Identities = 22/55 (40%), Positives = 35/55 (63%), Gaps = 3/55 (5%)
Frame = +3
Query: 309 NEWSCVICFEDFSST--LGVLP-CEHTFCLTCIQKWAHQRISMGKISTCPLCKAS 360
+E +CVIC E++ + +G L C H + ++CI+KW +SM K+ CP+CK S
Sbjct: 891 DEGNCVICLEEYKNMDDVGTLQTCGHDYHVSCIKKW----LSMKKL--CPICKVS 1037
>TC207478 similar to UP|Q6R567 (Q6R567) Ring domain containing protein,
partial (51%)
Length = 1103
Score = 40.8 bits (94), Expect = 0.001
Identities = 33/123 (26%), Positives = 51/123 (40%), Gaps = 4/123 (3%)
Frame = +1
Query: 249 RLCNQNRAMNGSSDGIEQITDSNQIEDPQTVEQTSIDLCSSDGKFIDGDQVDNVAGSPIS 308
RLC R M E + + +ED ++E C SD D D+ N +G
Sbjct: 88 RLCKIKREMALDQYFDEAVPQLDSLEDKSSLETWK---CGSDD-IADSDR--NASGG--- 240
Query: 309 NEWSCVICFEDFSSTLGVLPCEHTFCLTCIQKWAHQRISMG----KISTCPLCKASFAWY 364
+ C IC E + L C H +C CI KW + + + + CP+CK+ +
Sbjct: 241 --FDCNICLECVQDPVVTL-CGHLYCWPCIYKWLNLQTASSENEEEKQQCPVCKSEISQS 411
Query: 365 KIV 367
+V
Sbjct: 412 SLV 420
>CO980352
Length = 639
Score = 40.8 bits (94), Expect = 0.001
Identities = 20/52 (38%), Positives = 28/52 (53%), Gaps = 2/52 (3%)
Frame = -2
Query: 313 CVICFEDF--SSTLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFA 362
C IC E++ +G L CEH F + CIQ+W + + CP+CK S A
Sbjct: 497 CCICQEEYVVGDEVGDLQCEHRFHVVCIQEWMRLK------NWCPVCKVSAA 360
>TC220403 similar to UP|Q7X7W8 (Q7X7W8) Makorin ring-zinc-finger protein
(Makorin RING finger protein), partial (62%)
Length = 1109
Score = 40.4 bits (93), Expect = 0.002
Identities = 27/86 (31%), Positives = 37/86 (42%), Gaps = 25/86 (29%)
Frame = +1
Query: 308 SNEWSCVICFEDFSST-------LGVLP-CEHTFCLTCIQKWAHQRISMG---------- 349
S E C +C E S G+LP C+H FCL+CI+ W + + G
Sbjct: 481 SQEVQCNVCLERVLSKPKPADCKFGLLPECDHAFCLSCIRNWRNSAPTSGMDIINAGTAN 660
Query: 350 KISTCPLC-KASF------AWYKIVE 368
+ TCP+C K S+ WY E
Sbjct: 661 TVRTCPVCLKLSYFVIPSGIWYSTKE 738
>TC219821 weakly similar to UP|PJA1_MOUSE (O55176) Ubiquitin protein ligase
Praja1 , partial (4%)
Length = 1135
Score = 40.4 bits (93), Expect = 0.002
Identities = 21/49 (42%), Positives = 28/49 (56%), Gaps = 3/49 (6%)
Frame = +3
Query: 313 CVICFEDFSS--TLGVLP-CEHTFCLTCIQKWAHQRISMGKISTCPLCK 358
CVIC D+ L ++P C HTF L+CI W + K STCP+C+
Sbjct: 564 CVICLADYREREVLRIMPKCGHTFHLSCIDIW------LRKQSTCPVCR 692
>TC226910 weakly similar to UP|Q8LPJ6 (Q8LPJ6) Pspzf zinc finger
protein-like, partial (8%)
Length = 665
Score = 40.4 bits (93), Expect = 0.002
Identities = 20/52 (38%), Positives = 29/52 (55%), Gaps = 2/52 (3%)
Frame = +3
Query: 313 CVICFEDFSS--TLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFA 362
C IC E++ + LG L CEH + CI++WA Q+ + CP+CK A
Sbjct: 114 CSICQEEYEAGNELGRLNCEHIYHFQCIKQWAAQK------NFCPVCKQQVA 251
>TC232492 weakly similar to UP|Q8VYW5 (Q8VYW5) AT5g45290/K9E15_7, partial
(6%)
Length = 582
Score = 40.0 bits (92), Expect = 0.002
Identities = 19/63 (30%), Positives = 30/63 (47%), Gaps = 2/63 (3%)
Frame = +1
Query: 300 DNVAGSPISNEWSCVICFEDFSST--LGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLC 357
++ + S + + C +C + + VLPC H F C+ KW G+ TCPLC
Sbjct: 148 ESASASNFNEDSWCCVCLSRLKAKDEIRVLPCSHKFHKICVNKWL-----KGRHKTCPLC 312
Query: 358 KAS 360
+ S
Sbjct: 313 RFS 321
>TC229224 weakly similar to UP|Q9LUR1 (Q9LUR1) RING zinc finger protein-like,
partial (38%)
Length = 937
Score = 40.0 bits (92), Expect = 0.002
Identities = 20/50 (40%), Positives = 25/50 (50%), Gaps = 3/50 (6%)
Frame = +1
Query: 313 CVICFEDFS--STLGVLP-CEHTFCLTCIQKWAHQRISMGKISTCPLCKA 359
C +C +F T VLP C H+F + CI W H TCPLC+A
Sbjct: 367 CAVCLSEFEPGETGRVLPKCNHSFHIECIDMWFHSH------DTCPLCRA 498
>TC226909 similar to PIR|H96764|H96764 protein RING zinc finger protein
F25P22.18 [imported] - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (32%)
Length = 1522
Score = 40.0 bits (92), Expect = 0.002
Identities = 20/60 (33%), Positives = 32/60 (53%), Gaps = 2/60 (3%)
Frame = +3
Query: 305 SPISNEWSCVICFEDFSS--TLGVLPCEHTFCLTCIQKWAHQRISMGKISTCPLCKASFA 362
S + + C IC E++ + LG L CEH++ CI++W Q+ + CP+CK A
Sbjct: 1002 SKLQVDKECSICQEEYEAGDELGRLNCEHSYHFQCIKQWVAQK------NFCPVCKQQVA 1163
>TC206089 UP|Q39902 (Q39902) C-terminal zinc-finger (Fragment), complete
Length = 1401
Score = 40.0 bits (92), Expect = 0.002
Identities = 22/54 (40%), Positives = 33/54 (60%), Gaps = 3/54 (5%)
Frame = +3
Query: 310 EWSCVICFEDFSST--LGVLP-CEHTFCLTCIQKWAHQRISMGKISTCPLCKAS 360
E +C IC E++ + +G L C H + + CI+KW +SM K+ CP+CKAS
Sbjct: 1029 EEACAICLEEYKNMDDVGTLKACGHDYHVGCIRKW----LSMKKV--CPICKAS 1172
>TC217463 similar to PIR|E86378|E86378 protein F21J9.10 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(46%)
Length = 687
Score = 40.0 bits (92), Expect = 0.002
Identities = 27/80 (33%), Positives = 37/80 (45%), Gaps = 1/80 (1%)
Frame = +3
Query: 282 TSIDLCSSDGKFIDGDQVDNVAGSPISNEWSCVICFEDFSSTLGVLP-CEHTFCLTCIQK 340
+SID S K I+GD+ + + E C IC E T VLP C H C+ C +K
Sbjct: 162 SSIDGSSFGKKMIEGDE--KLINIDLEREDECGICLEP--CTKMVLPNCCHAMCIKCYRK 329
Query: 341 WAHQRISMGKISTCPLCKAS 360
W + +CP C+ S
Sbjct: 330 W------NTRSESCPFCRGS 371
>TC209562 similar to UP|Q8GT74 (Q8GT74) NEP1-interacting protein 2, partial
(91%)
Length = 1352
Score = 39.7 bits (91), Expect = 0.003
Identities = 25/79 (31%), Positives = 36/79 (44%), Gaps = 12/79 (15%)
Frame = +3
Query: 292 KFIDGDQVDNVAGSPISNE---------WSCVICFEDF--SSTLGVLP-CEHTFCLTCIQ 339
K + GD V+ + I+ + SC +C +DF T+ LP C H F L CI
Sbjct: 894 KGLSGDLVEKIPKIKITTDNNFDASGDRVSCSVCLQDFMLGETVRSLPHCHHMFHLPCID 1073
Query: 340 KWAHQRISMGKISTCPLCK 358
KW + +CPLC+
Sbjct: 1074KWLFRH------GSCPLCR 1112
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.133 0.393
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,346,717
Number of Sequences: 63676
Number of extensions: 284936
Number of successful extensions: 1541
Number of sequences better than 10.0: 204
Number of HSP's better than 10.0 without gapping: 1512
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1516
length of query: 423
length of database: 12,639,632
effective HSP length: 100
effective length of query: 323
effective length of database: 6,272,032
effective search space: 2025866336
effective search space used: 2025866336
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0225.8


