

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0225.5
(295 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
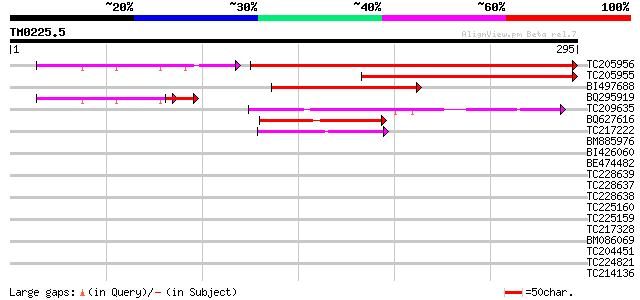
TC205956 homologue to UP|Q94BX6 (Q94BX6) K16N12.18/K16N12.18 (At... 318 1e-87
TC205955 homologue to UP|Q94BX6 (Q94BX6) K16N12.18/K16N12.18 (At... 214 4e-56
BI497688 similar to GP|14517500|gb| K16N12.18/K16N12.18 {Arabido... 117 6e-27
BQ295919 similar to GP|14517500|gb| K16N12.18/K16N12.18 {Arabido... 72 5e-16
TC209635 weakly similar to UP|Q7QF03 (Q7QF03) AgCP13350 (Fragmen... 69 2e-12
BQ627616 weakly similar to GP|18161138|gb| serine protease {Pyro... 56 2e-08
TC217222 similar to GB|BAB10901.1|10177507|AB010699 serine prote... 45 5e-05
BM885976 40 0.002
BI426060 similar to GP|10177507|db serine protease-like protein ... 40 0.002
BE474482 homologue to GP|14517500|gb| K16N12.18/K16N12.18 {Arabi... 38 0.006
TC228639 similar to UP|Q8VZD4 (Q8VZD4) At1g28320/F3H9_2, partial... 38 0.006
TC228637 similar to UP|Q8VZD4 (Q8VZD4) At1g28320/F3H9_2, partial... 35 0.040
TC228638 similar to UP|Q8VZD4 (Q8VZD4) At1g28320/F3H9_2, partial... 34 0.069
TC225160 UP|Q948X9 (Q948X9) Beta-conglycinin alpha-subunit, comp... 32 0.34
TC225159 homologue to UP|O22120 (O22120) Alpha subunit of beta c... 32 0.34
TC217328 weakly similar to GB|AAO42905.1|28416749|BT004659 At5g5... 32 0.45
BM086069 31 0.58
TC204451 similar to UP|TL38_ARATH (P82869) Peptidyl-prolyl cis-t... 31 0.76
TC224821 homologue to UP|Q94C02 (Q94C02) Myo-inositol-1-phosphat... 30 0.99
TC214136 similar to UP|Q84L12 (Q84L12) Peptide methionine sulfox... 30 0.99
>TC205956 homologue to UP|Q94BX6 (Q94BX6) K16N12.18/K16N12.18
(At3g27925/K16N12.18), partial (77%)
Length = 1489
Score = 318 bits (816), Expect = 1e-87
Identities = 160/170 (94%), Positives = 166/170 (97%)
Frame = +1
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKF 185
++ +DAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIP+DTVNGIVDQLVKF
Sbjct: 736 VIQTDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPVDTVNGIVDQLVKF 915
Query: 186 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 245
GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT
Sbjct: 916 GKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDIIT 1095
Query: 246 SVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDES 295
SVN KVT+GSDLYRILDQCKVGDK+ VEVLRGDHKEKIPVILEPKPDES
Sbjct: 1096SVNDKKVTNGSDLYRILDQCKVGDKLIVEVLRGDHKEKIPVILEPKPDES 1245
Score = 89.0 bits (219), Expect = 2e-18
Identities = 68/143 (47%), Positives = 75/143 (51%), Gaps = 37/143 (25%)
Frame = +1
Query: 15 HSHYSPHFTPNSLSLLRSSTPK----------TCFNSILILCTSIALSFT----NADSAY 60
H S H +L LLR P T S+L+LCTS+ALSFT +ADSA
Sbjct: 61 HLSKSLHLYTPTLFLLRPPNPTLPIIPLFPKLTIPKSLLLLCTSLALSFTLLLSDADSAA 240
Query: 61 AFVVTPPRKLQSDELAT----------VVYITNLAVKMRL-------------RWMCWRF 97
AFVVT PRKLQSDELAT VVYITNLAVK W
Sbjct: 241 AFVVTSPRKLQSDELATVRLFQENTPSVVYITNLAVKQDAFTLDVLEVPQGSGSGFVWD- 417
Query: 98 LKEGNIVTNYHVIPGASDLSLDL 120
KEG+IVTNYHVI GASDL + L
Sbjct: 418 -KEGHIVTNYHVIRGASDLKVTL 483
>TC205955 homologue to UP|Q94BX6 (Q94BX6) K16N12.18/K16N12.18
(At3g27925/K16N12.18), partial (25%)
Length = 631
Score = 214 bits (545), Expect = 4e-56
Identities = 108/112 (96%), Positives = 109/112 (96%)
Frame = +2
Query: 184 KFGKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDI 243
KFGKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDI
Sbjct: 2 KFGKVTRPILGIKFAPDQSVEQLGVSGVLVLDAPANGPAGKAGLQSTKRDSYGRLILGDI 181
Query: 244 ITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLRGDHKEKIPVILEPKPDES 295
ITSVN KVT+GSDLYRILDQCKVGDKV VEVLRGDHKEKIPVILEPKPDES
Sbjct: 182 ITSVNDKKVTNGSDLYRILDQCKVGDKVIVEVLRGDHKEKIPVILEPKPDES 337
>BI497688 similar to GP|14517500|gb| K16N12.18/K16N12.18 {Arabidopsis
thaliana}, partial (17%)
Length = 423
Score = 117 bits (293), Expect = 6e-27
Identities = 62/78 (79%), Positives = 66/78 (84%)
Frame = +3
Query: 137 NSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFGKVTRPILGIK 196
NSG LLDSSG LIGIN +IYSPS ASSGVGF I +D VNGIVDQLVKFGK+TR ILGIK
Sbjct: 189 NSGWLLLDSSGKLIGINISIYSPSRASSGVGFCILVDIVNGIVDQLVKFGKITRHILGIK 368
Query: 197 FAPDQSVEQLGVSGVLVL 214
FAPDQ V+QLGVS V VL
Sbjct: 369 FAPDQFVDQLGVSAVFVL 422
>BQ295919 similar to GP|14517500|gb| K16N12.18/K16N12.18 {Arabidopsis
thaliana}, partial (14%)
Length = 421
Score = 71.6 bits (174), Expect(2) = 5e-16
Identities = 52/99 (52%), Positives = 55/99 (55%), Gaps = 26/99 (26%)
Frame = +1
Query: 15 HSHYSPHFTPNSLSLLRSSTPK------------TCFNSILILCTSIALSFT----NADS 58
H S H L LLR PK T NS+L+LCTS+ALSFT ADS
Sbjct: 88 HLSKSLHLQTPILFLLRPPNPKPTLPIIPLFPKLTIPNSLLLLCTSLALSFTLLLSPADS 267
Query: 59 AYAFVVTPPRKLQSDELAT----------VVYITNLAVK 87
A AFVVT PRKLQSDELAT VVYITNLAVK
Sbjct: 268 AAAFVVTSPRKLQSDELATVRLFQENTPSVVYITNLAVK 384
Score = 30.0 bits (66), Expect(2) = 5e-16
Identities = 11/17 (64%), Positives = 13/17 (75%)
Frame = +2
Query: 82 TNLAVKMRLRWMCWRFL 98
T L+ +MRL W CWRFL
Sbjct: 371 TLLSSRMRLHWTCWRFL 421
>TC209635 weakly similar to UP|Q7QF03 (Q7QF03) AgCP13350 (Fragment), partial
(16%)
Length = 921
Score = 69.3 bits (168), Expect = 2e-12
Identities = 55/177 (31%), Positives = 83/177 (46%), Gaps = 12/177 (6%)
Frame = +2
Query: 125 QLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVK 184
+ + +D AIN GNSGGPL++ G +IG+N A+ G+ FS+P+D+V I++ K
Sbjct: 134 EYLQTDCAINMGNSGGPLVNMDGEIIGVN---IMKVAAADGLSFSVPIDSVCKILEHFKK 304
Query: 185 FGKVTRPILGIKFAP--DQSVEQLGV---------SGVLVLDAPANGPAGKAGLQSTKRD 233
G+V RP LG+K + + QL G+LV P +AG
Sbjct: 305 SGRVIRPWLGLKMLDLNEMIIAQLKKQDPSFPNVNKGILVPMVTPRSPGERAG------- 463
Query: 234 SYGRLILGDIITSVNGTKVTSGSDLYRILDQCKVGDKVTVEVLR-GDHKEKIPVILE 289
GD++ +G V ++ +L KVG + V V R GD + VI E
Sbjct: 464 ----FFPGDVVIEFDGRPVERLKEVIEVLGD-KVGVPIKVLVKRAGDKLVTLTVIPE 619
>BQ627616 weakly similar to GP|18161138|gb| serine protease {Pyrobaculum
aerophilum}, partial (17%)
Length = 431
Score = 56.2 bits (134), Expect = 2e-08
Identities = 28/66 (42%), Positives = 44/66 (66%)
Frame = +1
Query: 131 AAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFGKVTR 190
A++ GNSGGPL++ G +IG+N + A+ G+ FS+P+D+V I++ K G+V R
Sbjct: 226 ASVL*GNSGGPLVNMDGEIIGVNIMKVA---AADGLSFSVPIDSVCKILEHFKKSGRVIR 396
Query: 191 PILGIK 196
P LG+K
Sbjct: 397 PWLGLK 414
>TC217222 similar to GB|BAB10901.1|10177507|AB010699 serine protease-like
protein {Arabidopsis thaliana;} , partial (35%)
Length = 623
Score = 44.7 bits (104), Expect = 5e-05
Identities = 26/69 (37%), Positives = 36/69 (51%), Gaps = 1/69 (1%)
Frame = +1
Query: 130 DAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFGKVT 189
DAAIN GNSGGP + GN +GI A + +G+ IP + + K G T
Sbjct: 322 DAAINSGNSGGPAFNDKGNCVGIAFQSLKHEDAEN-IGYVIPTPVIMHFIQDYEKNGGYT 498
Query: 190 R-PILGIKF 197
PILG+++
Sbjct: 499 GFPILGVEW 525
>BM885976
Length = 421
Score = 39.7 bits (91), Expect = 0.002
Identities = 24/64 (37%), Positives = 37/64 (57%)
Frame = +3
Query: 124 LQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLV 183
L + DAAINPGNSGGP + S + G+ A + SGA + +G+ IP+ + + +
Sbjct: 30 LMAIQIDAAINPGNSGGPAIMGS-KVAGV--AFQNLSGAEN-IGYIIPVPVIEHFISGVE 197
Query: 184 KFGK 187
+ GK
Sbjct: 198 ENGK 209
>BI426060 similar to GP|10177507|db serine protease-like protein {Arabidopsis
thaliana}, partial (23%)
Length = 426
Score = 39.7 bits (91), Expect = 0.002
Identities = 24/69 (34%), Positives = 34/69 (48%), Gaps = 1/69 (1%)
Frame = +2
Query: 130 DAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVGFSIPLDTVNGIVDQLVKFGKVT 189
DAAIN GNSGGP + G +GI + +G+ IP + + K G T
Sbjct: 59 DAAINSGNSGGPAFNDKGKCVGIAFQSLKHEDVEN-IGYVIPTPVILYFIRDYEKNGAYT 235
Query: 190 R-PILGIKF 197
PILG+++
Sbjct: 236 GFPILGVEW 262
>BE474482 homologue to GP|14517500|gb| K16N12.18/K16N12.18 {Arabidopsis
thaliana}, partial (19%)
Length = 401
Score = 37.7 bits (86), Expect = 0.006
Identities = 20/30 (66%), Positives = 21/30 (69%)
Frame = +3
Query: 99 KEGNIVTNYHVIPGASDLSLDLTTLLQLVS 128
KEG+IVTNYHVI GASDL L L L S
Sbjct: 36 KEGHIVTNYHVIRGASDLKLLCAGSLLLTS 125
>TC228639 similar to UP|Q8VZD4 (Q8VZD4) At1g28320/F3H9_2, partial (21%)
Length = 746
Score = 37.7 bits (86), Expect = 0.006
Identities = 21/57 (36%), Positives = 36/57 (62%), Gaps = 2/57 (3%)
Frame = +3
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGAS--SGVGFSIPLDTVNGIVD 180
++ + AAI+PG SGG +++S G++IG+ T+ SG + + FSIP + IV+
Sbjct: 516 MLETTAAIHPGASGGAIINSDGHMIGLVTSNARHSGGAIIPQLNFSIPSAALAPIVN 686
>TC228637 similar to UP|Q8VZD4 (Q8VZD4) At1g28320/F3H9_2, partial (15%)
Length = 867
Score = 35.0 bits (79), Expect = 0.040
Identities = 18/48 (37%), Positives = 31/48 (64%), Gaps = 2/48 (4%)
Frame = +2
Query: 126 LVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGAS--SGVGFSIP 171
++ + A I+PG SGG +++S G++IG+ T+ SG + + FSIP
Sbjct: 266 MLETTATIHPGASGGAVINSDGHMIGLVTSNARHSGGAVIPHLNFSIP 409
>TC228638 similar to UP|Q8VZD4 (Q8VZD4) At1g28320/F3H9_2, partial (9%)
Length = 696
Score = 34.3 bits (77), Expect = 0.069
Identities = 21/52 (40%), Positives = 31/52 (59%), Gaps = 2/52 (3%)
Frame = +1
Query: 131 AAINPGNSGGPLLDSSGNLIGINTAIYSPSGAS--SGVGFSIPLDTVNGIVD 180
AAI+PG SGG +S G++IG+ T+ SG + + FSIP + IV+
Sbjct: 31 AAIHPGASGGAXXNSDGHMIGLVTSNARHSGGAIIPQLNFSIPSAALAPIVN 186
>TC225160 UP|Q948X9 (Q948X9) Beta-conglycinin alpha-subunit, complete
Length = 2152
Score = 32.0 bits (71), Expect = 0.34
Identities = 23/86 (26%), Positives = 43/86 (49%), Gaps = 1/86 (1%)
Frame = -3
Query: 108 HVIPGASDLSLDLTTLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVG 167
H++ G+ S +T L + S ++ G +G P+ D+ +I + S G GVG
Sbjct: 1013 HIVCGS*GDSQGITRLKSVGVSVVVVHQGKNGSPVKDND-EVISV-----SMVGEKEGVG 852
Query: 168 FSIPLDTVNGIVDQLVK-FGKVTRPI 192
F + L N +V +++K +G + P+
Sbjct: 851 FGVELQ--NAVVSEILKLWGALVEPL 780
>TC225159 homologue to UP|O22120 (O22120) Alpha subunit of beta conglycinin
(Fragment), partial (50%)
Length = 849
Score = 32.0 bits (71), Expect = 0.34
Identities = 23/86 (26%), Positives = 43/86 (49%), Gaps = 1/86 (1%)
Frame = -1
Query: 108 HVIPGASDLSLDLTTLLQLVSSDAAINPGNSGGPLLDSSGNLIGINTAIYSPSGASSGVG 167
H++ G+ S +T L + S ++ G +G P+ D+ +I + S G GVG
Sbjct: 474 HIVCGS*GDSQGITRLKSVGVSVVVVHQGKNGSPVKDND-EVISV-----SMVGEKEGVG 313
Query: 168 FSIPLDTVNGIVDQLVK-FGKVTRPI 192
F + L N +V +++K +G + P+
Sbjct: 312 FGVELQ--NAVVSEILKLWGALVEPL 241
>TC217328 weakly similar to GB|AAO42905.1|28416749|BT004659 At5g53650
{Arabidopsis thaliana;} , partial (79%)
Length = 563
Score = 31.6 bits (70), Expect = 0.45
Identities = 15/42 (35%), Positives = 24/42 (56%)
Frame = -3
Query: 3 QHYKHHKQRLHYHSHYSPHFTPNSLSLLRSSTPKTCFNSILI 44
+H ++R HSH+ PHF P SLSL +S +++L+
Sbjct: 135 RHCSPGEERRQSHSHHIPHFPPFSLSLYYTSQILLLLSTMLL 10
>BM086069
Length = 429
Score = 31.2 bits (69), Expect = 0.58
Identities = 22/81 (27%), Positives = 37/81 (45%), Gaps = 3/81 (3%)
Frame = -2
Query: 7 HHKQRLHYHSHYSPHFTPNSLSLLRSSTPKTCFNSILILCTS---IALSFTNADSAYAFV 63
HH R H H HY H ++ ++ P T +I+I+ T+ ++ TN A +
Sbjct: 353 HHCDRQHCHHHYHCHCHHHTATICYHHHPMT--GTIIIIATNTVDTTITSTNTTFVTATI 180
Query: 64 VTPPRKLQSDELATVVYITNL 84
T P + Q +L+ + T L
Sbjct: 179 GTNPLETQWGQLSPQTFKTLL 117
>TC204451 similar to UP|TL38_ARATH (P82869) Peptidyl-prolyl cis-trans
isomerase TLP38, chloroplast precursor (PPIase)
(Rotamase) (Thylakoid lumen PPIase of 38 kDa) (p38) ,
partial (33%)
Length = 715
Score = 30.8 bits (68), Expect = 0.76
Identities = 16/46 (34%), Positives = 25/46 (53%), Gaps = 3/46 (6%)
Frame = +3
Query: 131 AAINPGNSGGPLLDSSGN---LIGINTAIYSPSGASSGVGFSIPLD 173
A + PGN ++D + N L IN A+ S +G G+S+PL+
Sbjct: 21 APLTPGNFAKLVMDGAYNGAKLNCINQAVISDNGVDKNTGYSVPLE 158
>TC224821 homologue to UP|Q94C02 (Q94C02) Myo-inositol-1-phosphate synthase
, complete
Length = 1826
Score = 30.4 bits (67), Expect = 0.99
Identities = 21/66 (31%), Positives = 29/66 (43%), Gaps = 3/66 (4%)
Frame = -2
Query: 7 HHKQRLHYHSHYSPHFTPNSLSLLRSSTPKTCFNSILILC---TSIALSFTNADSAYAFV 63
H KQ L + ++ P PNS S PK L+LC ++++S A V
Sbjct: 484 HWKQALEWGIYFLPLERPNSNS*CLGQRPKVIGLLNLVLCCP*NTLSVSNNTTGEG*AIV 305
Query: 64 VTPPRK 69
TPP K
Sbjct: 304 STPPHK 287
>TC214136 similar to UP|Q84L12 (Q84L12) Peptide methionine sulfoxide
reductase, partial (71%)
Length = 1164
Score = 30.4 bits (67), Expect = 0.99
Identities = 15/37 (40%), Positives = 19/37 (50%)
Frame = +1
Query: 14 YHSHYSPHFTPNSLSLLRSSTPKTCFNSILILCTSIA 50
YH + PHF + L L S P FN L+L SI+
Sbjct: 67 YHCYRIPHFLLHLLLLCHFSAPLNLFNFFLLLAISIS 177
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,267,166
Number of Sequences: 63676
Number of extensions: 188767
Number of successful extensions: 1903
Number of sequences better than 10.0: 67
Number of HSP's better than 10.0 without gapping: 1746
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1847
length of query: 295
length of database: 12,639,632
effective HSP length: 97
effective length of query: 198
effective length of database: 6,463,060
effective search space: 1279685880
effective search space used: 1279685880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0225.5


