

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0218.20
(631 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
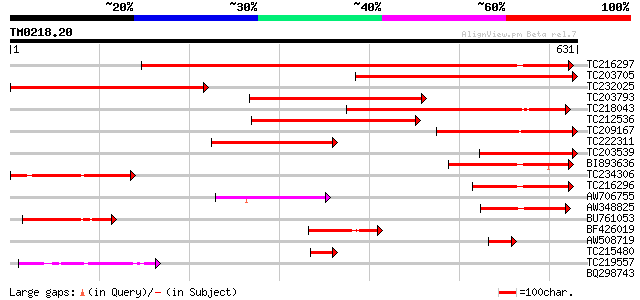
Sequences producing significant alignments: (bits) Value
TC216297 similar to UP|Q93X09 (Q93X09) Vacuolar sorting receptor... 744 0.0
TC203705 similar to UP|P93484 (P93484) BP-80 vacuolar sorting re... 498 e-141
TC232025 similar to UP|P93484 (P93484) BP-80 vacuolar sorting re... 405 e-113
TC203793 similar to PIR|T02602|T02602 vacuolar sorting receptor ... 398 e-111
TC218043 similar to UP|O49438 (O49438) Vacuolar sorting receptor... 379 e-105
TC212536 homologue to UP|P93484 (P93484) BP-80 vacuolar sorting ... 368 e-102
TC209167 similar to UP|P93484 (P93484) BP-80 vacuolar sorting re... 297 8e-81
TC222311 similar to UP|Q9FYH7 (Q9FYH7) F17F8.23, partial (21%) 225 4e-59
TC203539 similar to UP|Q9SDR8 (Q9SDR8) Vacuolar sorting receptor... 206 3e-53
BI893636 similar to GP|15487292|db vacuolar sorting receptor {Vi... 173 2e-43
TC234306 homologue to UP|Q93X09 (Q93X09) Vacuolar sorting recept... 163 2e-40
TC216296 similar to UP|Q93X09 (Q93X09) Vacuolar sorting receptor... 147 2e-35
AW706755 similar to GP|15487292|dbj vacuolar sorting receptor {V... 117 1e-26
AW348825 similar to GP|15487292|dbj vacuolar sorting receptor {V... 113 3e-25
BU761053 similar to GP|22530930|gb| vacuolar sorting receptor-li... 110 1e-24
BF426019 90 3e-18
AW508719 59 5e-09
TC215480 similar to GB|BAA96951.1|8777361|AB019233 protein carbo... 59 9e-09
TC219557 similar to UP|Q9M621 (Q9M621) ReMembR-H2 protein JR702 ... 47 2e-05
BQ298743 32 0.66
>TC216297 similar to UP|Q93X09 (Q93X09) Vacuolar sorting receptor, partial
(78%)
Length = 1641
Score = 744 bits (1922), Expect = 0.0
Identities = 330/481 (68%), Positives = 398/481 (82%)
Frame = +1
Query: 147 YIENITIPSALIEKSFGEKLKKSISGGEMVNVNLDWREAVPHPDDRVEYELWTNSNDECG 206
Y++ I+IPSALI KS G+ +K+++S GEMVN+NLDWRE++PHPDDRVEYELWTNSNDECG
Sbjct: 4 YVDKISIPSALISKSLGDSIKQALSDGEMVNINLDWRESLPHPDDRVEYELWTNSNDECG 183
Query: 207 VKCDMLMEFVKDFKGAAQILEKGGYTQFTPHYITWYCPKAFTLSKQCKSQCINHGRYCAP 266
KCD L+ F+KDFKG AQ LEK G+TQFTP YITW+CP+AF LSKQCKSQCIN+GRYCAP
Sbjct: 184 PKCDSLINFLKDFKGVAQQLEKKGFTQFTPRYITWFCPEAFLLSKQCKSQCINNGRYCAP 363
Query: 267 DPEQDFSTGYDGKDVVIENLRQLCVYKVTSEAKKPWLWWDYVTDFQIRCPMKEKKYNKKC 326
DPEQDFS GYDGKDVV++NLRQ C YKV +E+ KPW WWDYVTDF IRCPMKE KY+++C
Sbjct: 364 DPEQDFSRGYDGKDVVVQNLRQACFYKVANESGKPWQWWDYVTDFAIRCPMKENKYSEEC 543
Query: 327 ADAVIESLGLDNKKIERCMGDPDADSDNPVLKEEQDAQVGKGSRGDVTILPTLVVNNRQY 386
+D VI+SLG D KKI+ C+GDP AD +NPVLK EQDAQ+G+GSRGDVTILPTLV+NNRQY
Sbjct: 544 SDQVIKSLGADLKKIKDCVGDPHADVENPVLKAEQDAQIGQGSRGDVTILPTLVINNRQY 723
Query: 387 RGKLEKGAVMKAICSGFEETTEPAVCLSSDVETNECLENNGGCWKDKAANITACKDTFRG 446
RGKL + +V+KAICSG+ ETTEP++CL+SD+ETNECLENNGGCW+DK++NITAC+DTFRG
Sbjct: 724 RGKLSRPSVLKAICSGYLETTEPSICLTSDLETNECLENNGGCWQDKSSNITACRDTFRG 903
Query: 447 RVCECPLVDGVQFKGDGYTTCEASGPGRCKIKNGGCWHEARNGHAFSACVDDGEVKCQCP 506
RVCECP+V V+F GDGYT CEASG C+ NGGCW A+ G A+SAC+DD C CP
Sbjct: 904 RVCECPIVQNVKFFGDGYTHCEASGSLSCEFNNGGCWKGAQGGRAYSACLDDYRKGCTCP 1083
Query: 507 AGFKGDGVKNCQDVDECKEKKACQCPECSCKNTWGSYDCSCSGDLLYIRDHDTCISKTAS 566
GF+GDGV++C+D+DEC EK +CQCP C CKNTWGSY+C C L Y R++DTC + +
Sbjct: 1084PGFRGDGVQSCEDIDECNEKTSCQCPGCKCKNTWGSYECKCKSGLFYSRENDTCFGEYS- 1260
Query: 567 QGGRSAWAAFWVIIAGLVLSAGGGYLIYKYRIRSYMDSEIRAIMAQYMPLDSQAEVVNHV 626
++ WVII LV++ GGY YKYRI+ YMDSEIR IMAQYMPLDSQ +V N V
Sbjct: 1261----ASVLNIWVIILVLVVAVAGGYAFYKYRIQRYMDSEIRTIMAQYMPLDSQPDVSNQV 1428
Query: 627 N 627
+
Sbjct: 1429H 1431
>TC203705 similar to UP|P93484 (P93484) BP-80 vacuolar sorting receptor,
partial (38%)
Length = 1045
Score = 498 bits (1283), Expect = e-141
Identities = 224/247 (90%), Positives = 236/247 (94%)
Frame = +1
Query: 385 QYRGKLEKGAVMKAICSGFEETTEPAVCLSSDVETNECLENNGGCWKDKAANITACKDTF 444
QYRGKLEKGAVMKAIC+GFEETTEPAVCLSSDVETNECLENNGGCW+DK ANITACKDTF
Sbjct: 1 QYRGKLEKGAVMKAICAGFEETTEPAVCLSSDVETNECLENNGGCWQDKVANITACKDTF 180
Query: 445 RGRVCECPLVDGVQFKGDGYTTCEASGPGRCKIKNGGCWHEARNGHAFSACVDDGEVKCQ 504
RGRVCECPLVDGVQFKGDGYT C ASGPGRCKI NGGCWHEARNGHA+SAC DDG VKCQ
Sbjct: 181 RGRVCECPLVDGVQFKGDGYTICAASGPGRCKINNGGCWHEARNGHAYSACSDDGGVKCQ 360
Query: 505 CPAGFKGDGVKNCQDVDECKEKKACQCPECSCKNTWGSYDCSCSGDLLYIRDHDTCISKT 564
CPAGFKGDGVKNC+D+DECKEKKACQCPECSCKNTWGSYDC+CSGDLLYIRDHDTCISKT
Sbjct: 361 CPAGFKGDGVKNCEDIDECKEKKACQCPECSCKNTWGSYDCTCSGDLLYIRDHDTCISKT 540
Query: 565 ASQGGRSAWAAFWVIIAGLVLSAGGGYLIYKYRIRSYMDSEIRAIMAQYMPLDSQAEVVN 624
ASQ GRSAWAAFWVI+ GLV++A G YL+YKYRIRSYMDSEIRAIMAQYMPLDSQ EVVN
Sbjct: 541 ASQEGRSAWAAFWVILVGLVVAAAGAYLVYKYRIRSYMDSEIRAIMAQYMPLDSQGEVVN 720
Query: 625 HVNDERA 631
HV++ERA
Sbjct: 721 HVHEERA 741
>TC232025 similar to UP|P93484 (P93484) BP-80 vacuolar sorting receptor,
partial (31%)
Length = 832
Score = 405 bits (1042), Expect = e-113
Identities = 200/221 (90%), Positives = 207/221 (93%)
Frame = +1
Query: 1 MKLRRFSLGFLLGFLLLLSVTPSSMARFVVEKNSLTVTSPEDIKGTHDSAIGNFGIPQYG 60
MKLRR SL LGFL L P SMA+FVVEKNSLTVTSP++IKGTHDSAIGNFGIPQYG
Sbjct: 169 MKLRRSSLCVFLGFLALFLSPPPSMAKFVVEKNSLTVTSPDNIKGTHDSAIGNFGIPQYG 348
Query: 61 GSMAGNVVYPKDNHKGCKEFDESGISFKSKPGALPTIVLLDRGNCFFALKVWNAQKAGAS 120
GSMAGNV+YPKDN KGCKEFDE GISFKSKPGALPTIVLLDRGNCFFALKVWNAQKAGAS
Sbjct: 349 GSMAGNVLYPKDNKKGCKEFDEYGISFKSKPGALPTIVLLDRGNCFFALKVWNAQKAGAS 528
Query: 121 AVLVADDIEEKLITMDTPEEDGSSAKYIENITIPSALIEKSFGEKLKKSISGGEMVNVNL 180
AVLV+DDIEEKLITMDTPEEDGSSAKYIENITIPSALIEKSFGEKLK +IS G+MVNVNL
Sbjct: 529 AVLVSDDIEEKLITMDTPEEDGSSAKYIENITIPSALIEKSFGEKLKVAISNGDMVNVNL 708
Query: 181 DWREAVPHPDDRVEYELWTNSNDECGVKCDMLMEFVKDFKG 221
DWREAVPHPDDRVEYELWTNSNDECGVKCDMLMEFVKDFKG
Sbjct: 709 DWREAVPHPDDRVEYELWTNSNDECGVKCDMLMEFVKDFKG 831
>TC203793 similar to PIR|T02602|T02602 vacuolar sorting receptor protein
homolog At2g14740 - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (31%)
Length = 711
Score = 398 bits (1022), Expect = e-111
Identities = 185/197 (93%), Positives = 193/197 (97%)
Frame = +2
Query: 267 DPEQDFSTGYDGKDVVIENLRQLCVYKVTSEAKKPWLWWDYVTDFQIRCPMKEKKYNKKC 326
DPEQDFSTGYDGKDVVIENLRQLCV+KV +E KKPW+WWDYVTDFQIRCPMKEKKYNKKC
Sbjct: 11 DPEQDFSTGYDGKDVVIENLRQLCVFKVANETKKPWVWWDYVTDFQIRCPMKEKKYNKKC 190
Query: 327 ADAVIESLGLDNKKIERCMGDPDADSDNPVLKEEQDAQVGKGSRGDVTILPTLVVNNRQY 386
ADAVIESLGLD KKIERCMGDP+ADS+NPVLKEEQDAQVGKGSRGDVTILPTLVVNNRQY
Sbjct: 191 ADAVIESLGLDIKKIERCMGDPNADSENPVLKEEQDAQVGKGSRGDVTILPTLVVNNRQY 370
Query: 387 RGKLEKGAVMKAICSGFEETTEPAVCLSSDVETNECLENNGGCWKDKAANITACKDTFRG 446
RGKLEKGAVMKAIC+GFEETTEPAVCLSSDVETNECLENNGGCW+DK ANITACKDTFRG
Sbjct: 371 RGKLEKGAVMKAICAGFEETTEPAVCLSSDVETNECLENNGGCWQDKVANITACKDTFRG 550
Query: 447 RVCECPLVDGVQFKGDG 463
RVCECPLVDGVQFKG+G
Sbjct: 551 RVCECPLVDGVQFKGEG 601
>TC218043 similar to UP|O49438 (O49438) Vacuolar sorting receptor-like
protein, partial (36%)
Length = 1124
Score = 379 bits (974), Expect = e-105
Identities = 173/249 (69%), Positives = 201/249 (80%)
Frame = +2
Query: 376 LPTLVVNNRQYRGKLEKGAVMKAICSGFEETTEPAVCLSSDVETNECLENNGGCWKDKAA 435
LPTLV+NN QYRGKLE+ AV+KA+CSGF+ETTEP+VCLS DVETNECLE NGGCW+DK A
Sbjct: 2 LPTLVINNVQYRGKLERTAVLKAVCSGFKETTEPSVCLSGDVETNECLERNGGCWQDKHA 181
Query: 436 NITACKDTFRGRVCECPLVDGVQFKGDGYTTCEASGPGRCKIKNGGCWHEARNGHAFSAC 495
NITACKDTFRGRVCECP+V+GVQ+KGDGYTTCEA GP RC I NGGCW E + G FSAC
Sbjct: 182 NITACKDTFRGRVCECPVVNGVQYKGDGYTTCEAFGPARCSINNGGCWSETKKGLTFSAC 361
Query: 496 VDDGEVKCQCPAGFKGDGVKNCQDVDECKEKKACQCPECSCKNTWGSYDCSCSGDLLYIR 555
D CQCP GF+GDG C+DVDECKE+ ACQC CSCKNTWGSYDC C G+LLYI+
Sbjct: 362 SDSKVNGCQCPVGFRGDGTNKCEDVDECKERSACQCDGCSCKNTWGSYDCKCKGNLLYIK 541
Query: 556 DHDTCISKTASQGGRSAWAAFWVIIAGLVLSAGGGYLIYKYRIRSYMDSEIRAIMAQYMP 615
+ D CI ++ S+ GR + AF V+IA +V + GY+ YKYR+RSYMDSEI AIM+QYMP
Sbjct: 542 EQDACIERSESKFGR--FLAF-VVIAVVVGAGLAGYVFYKYRLRSYMDSEIMAIMSQYMP 712
Query: 616 LDSQAEVVN 624
LD Q VV+
Sbjct: 713 LDQQNNVVH 739
>TC212536 homologue to UP|P93484 (P93484) BP-80 vacuolar sorting receptor,
partial (30%)
Length = 567
Score = 368 bits (944), Expect = e-102
Identities = 168/188 (89%), Positives = 183/188 (96%)
Frame = +3
Query: 270 QDFSTGYDGKDVVIENLRQLCVYKVTSEAKKPWLWWDYVTDFQIRCPMKEKKYNKKCADA 329
QDF+TGYDGKDVVIENLRQLCV+KV +E +KPW+WWDYVTDFQIRCPMKEKKYNK+CA+A
Sbjct: 3 QDFTTGYDGKDVVIENLRQLCVFKVANETEKPWVWWDYVTDFQIRCPMKEKKYNKECANA 182
Query: 330 VIESLGLDNKKIERCMGDPDADSDNPVLKEEQDAQVGKGSRGDVTILPTLVVNNRQYRGK 389
VI+SLGL+ +KIE+CMGDPDAD+DNPVLKEEQDAQ+GKGSRGDVTILPTLVVNNRQYRGK
Sbjct: 183 VIKSLGLNTEKIEKCMGDPDADTDNPVLKEEQDAQIGKGSRGDVTILPTLVVNNRQYRGK 362
Query: 390 LEKGAVMKAICSGFEETTEPAVCLSSDVETNECLENNGGCWKDKAANITACKDTFRGRVC 449
LEKGAV+KAICSGFEETTEP VCLSSDVETNECL NNGGCW+DKAANITACKDTFRGRVC
Sbjct: 363 LEKGAVLKAICSGFEETTEPTVCLSSDVETNECLTNNGGCWQDKAANITACKDTFRGRVC 542
Query: 450 ECPLVDGV 457
ECPLVDGV
Sbjct: 543 ECPLVDGV 566
>TC209167 similar to UP|P93484 (P93484) BP-80 vacuolar sorting receptor,
partial (25%)
Length = 783
Score = 297 bits (761), Expect = 8e-81
Identities = 130/156 (83%), Positives = 145/156 (92%)
Frame = +1
Query: 476 KIKNGGCWHEARNGHAFSACVDDGEVKCQCPAGFKGDGVKNCQDVDECKEKKACQCPECS 535
KI NGGCWH+A+NGHAFSAC+D+G VKCQCP GF+GDGVKNCQD+DECKEKKACQCP+CS
Sbjct: 10 KINNGGCWHDAQNGHAFSACLDNGGVKCQCPTGFRGDGVKNCQDIDECKEKKACQCPDCS 189
Query: 536 CKNTWGSYDCSCSGDLLYIRDHDTCISKTASQGGRSAWAAFWVIIAGLVLSAGGGYLIYK 595
CKNTWGSYDCSCSGDLLY+RDHDTCISKT SQ GRS WAAFW+I+ G+V+ AGG +L+YK
Sbjct: 190 CKNTWGSYDCSCSGDLLYMRDHDTCISKTGSQ-GRSTWAAFWLILLGVVMIAGGAFLVYK 366
Query: 596 YRIRSYMDSEIRAIMAQYMPLDSQAEVVNHVNDERA 631
YRIR YMDSEIRAIMAQYMPLDSQAEV NHVND+RA
Sbjct: 367 YRIRQYMDSEIRAIMAQYMPLDSQAEVPNHVNDQRA 474
>TC222311 similar to UP|Q9FYH7 (Q9FYH7) F17F8.23, partial (21%)
Length = 421
Score = 225 bits (574), Expect = 4e-59
Identities = 96/140 (68%), Positives = 116/140 (82%)
Frame = +1
Query: 225 ILEKGGYTQFTPHYITWYCPKAFTLSKQCKSQCINHGRYCAPDPEQDFSTGYDGKDVVIE 284
ILE+GGYT FTPHYITW+CP F LS QCKSQCINHGRYCAPDPE+DF GY+GKDVV E
Sbjct: 1 ILERGGYTLFTPHYITWFCPPPFILSSQCKSQCINHGRYCAPDPEKDFGEGYEGKDVVYE 180
Query: 285 NLRQLCVYKVTSEAKKPWLWWDYVTDFQIRCPMKEKKYNKKCADAVIESLGLDNKKIERC 344
NLRQLCV++V +E+ + W+WWDYVTDF +RC MKEK+Y+K CA+ V++SL L KI++C
Sbjct: 181 NLRQLCVHRVANESNRSWVWWDYVTDFHVRCSMKEKRYSKDCAEEVMKSLDLPVDKIKKC 360
Query: 345 MGDPDADSDNPVLKEEQDAQ 364
MGDP+AD +N VLK EQ Q
Sbjct: 361 MGDPEADVENEVLKNEQQVQ 420
>TC203539 similar to UP|Q9SDR8 (Q9SDR8) Vacuolar sorting receptor protein
(Fragment), partial (66%)
Length = 648
Score = 206 bits (524), Expect = 3e-53
Identities = 92/108 (85%), Positives = 101/108 (93%)
Frame = +1
Query: 524 KEKKACQCPECSCKNTWGSYDCSCSGDLLYIRDHDTCISKTASQGGRSAWAAFWVIIAGL 583
+EK CQCPECSCKNTWGSYDC+CSGDLLYIRDHDTCISKTASQ GRSAWAAFWVI+ GL
Sbjct: 7 EEKNVCQCPECSCKNTWGSYDCTCSGDLLYIRDHDTCISKTASQEGRSAWAAFWVILIGL 186
Query: 584 VLSAGGGYLIYKYRIRSYMDSEIRAIMAQYMPLDSQAEVVNHVNDERA 631
V++A G YL+YKYRIRSYMDSEIRAIMAQYMPLDSQ E+VNHV++ERA
Sbjct: 187 VVAAAGAYLVYKYRIRSYMDSEIRAIMAQYMPLDSQGEIVNHVSEERA 330
>BI893636 similar to GP|15487292|db vacuolar sorting receptor {Vigna mungo},
partial (22%)
Length = 437
Score = 173 bits (438), Expect = 2e-43
Identities = 79/144 (54%), Positives = 99/144 (67%), Gaps = 5/144 (3%)
Frame = +2
Query: 489 GHAFSACVDDGEVKCQCPAGFKGDGVKNCQDVDECKEKKACQCPECSCKNTWGSYDCSCS 548
G A+SAC+DD C CP GF+GDGV++C+D+DEC EK +CQCP C CKNTWGSY+C C
Sbjct: 5 GRAYSACLDDYRKGCTCPPGFRGDGVQSCEDIDECNEKTSCQCPGCKCKNTWGSYECKCK 184
Query: 549 GDLLYIRDHDTCISKTASQGGRSAWAAFWVIIAGLVLSAGGGYLIYKYRI-----RSYMD 603
L Y R++DTC + + ++ WVII LV++ GGY YKYRI + YMD
Sbjct: 185 SGLFYSRENDTCFGEYS-----ASVLNIWVIILVLVVAVAGGYAFYKYRIQLIYVQRYMD 349
Query: 604 SEIRAIMAQYMPLDSQAEVVNHVN 627
SEIR IMAQYMPLDSQ +V N V+
Sbjct: 350 SEIRTIMAQYMPLDSQPDVSNQVH 421
>TC234306 homologue to UP|Q93X09 (Q93X09) Vacuolar sorting receptor, partial
(21%)
Length = 639
Score = 163 bits (412), Expect = 2e-40
Identities = 81/140 (57%), Positives = 103/140 (72%)
Frame = +1
Query: 1 MKLRRFSLGFLLGFLLLLSVTPSSMARFVVEKNSLTVTSPEDIKGTHDSAIGNFGIPQYG 60
M + SL +G LLL + RFVVEKNSL VTSP+ +KGT++ AIGNFG+P+YG
Sbjct: 238 MGTTKLSLLLCVGVLLLRCC----VGRFVVEKNSLKVTSPKSLKGTYECAIGNFGVPKYG 405
Query: 61 GSMAGNVVYPKDNHKGCKEFDESGISFKSKPGALPTIVLLDRGNCFFALKVWNAQKAGAS 120
G++ G+V+YPK N KGC F S ++F+SKPG PT +L+DRG+C+F LK WNAQ GA+
Sbjct: 406 GTLVGSVLYPKVNQKGCTNF--SDVNFQSKPGGSPTFLLVDRGDCYFTLKAWNAQNGGAA 579
Query: 121 AVLVADDIEEKLITMDTPEE 140
A+LVADD E LITMDTPEE
Sbjct: 580 AILVADDKAETLITMDTPEE 639
>TC216296 similar to UP|Q93X09 (Q93X09) Vacuolar sorting receptor, partial
(18%)
Length = 648
Score = 147 bits (370), Expect = 2e-35
Identities = 66/112 (58%), Positives = 81/112 (71%)
Frame = +3
Query: 516 NCQDVDECKEKKACQCPECSCKNTWGSYDCSCSGDLLYIRDHDTCISKTASQGGRSAWAA 575
+C+D+DECKEK ACQCP C CKNTWGSY+C C L Y R++DTC + + ++
Sbjct: 3 SCEDIDECKEKTACQCPGCKCKNTWGSYECKCKSGLFYSRENDTCFGEYS-----ASVLN 167
Query: 576 FWVIIAGLVLSAGGGYLIYKYRIRSYMDSEIRAIMAQYMPLDSQAEVVNHVN 627
WVII LV++ GGY YKYRI+ YMDSEIRAIMAQYMPLD+Q EV N V+
Sbjct: 168 IWVIILVLVVAVAGGYAFYKYRIQRYMDSEIRAIMAQYMPLDNQPEVSNQVH 323
>AW706755 similar to GP|15487292|dbj vacuolar sorting receptor {Vigna mungo},
partial (12%)
Length = 421
Score = 117 bits (294), Expect = 1e-26
Identities = 57/132 (43%), Positives = 79/132 (59%), Gaps = 4/132 (3%)
Frame = +2
Query: 230 GYTQFTPHYITWYCPKAFTLSKQCKSQCINHGR----YCAPDPEQDFSTGYDGKDVVIEN 285
G+T FT ++T + P+ F L+KQ + + +P K + + N
Sbjct: 2 GFTNFTLGFLTGFGPEPFFLTKQANFNALTMEKGLCVLISPTKYSQIEGSISEKILFVLN 181
Query: 286 LRQLCVYKVTSEAKKPWLWWDYVTDFQIRCPMKEKKYNKKCADAVIESLGLDNKKIERCM 345
LR C YKV +E+ KPW WWDYVTDF IRCPMKE K++++C D VI+SLG+D +K + C+
Sbjct: 182 LRPACFYKVANESGKPWQWWDYVTDFSIRCPMKENKFSEECVDQVIKSLGVDLEKNKDCV 361
Query: 346 GDPDADSDNPVL 357
GDP AD N VL
Sbjct: 362 GDPHADI*NAVL 397
>AW348825 similar to GP|15487292|dbj vacuolar sorting receptor {Vigna mungo},
partial (15%)
Length = 475
Score = 113 bits (282), Expect = 3e-25
Identities = 52/100 (52%), Positives = 65/100 (65%)
Frame = -2
Query: 525 EKKACQCPECSCKNTWGSYDCSCSGDLLYIRDHDTCISKTASQGGRSAWAAFWVIIAGLV 584
EK +CQCP C CKN+WGSY+C C L Y R++DTC + ++ + WVII LV
Sbjct: 456 EKTSCQCPGCKCKNSWGSYECKCKSGLFYSRENDTCFGEYSA-----SVLNIWVIILVLV 292
Query: 585 LSAGGGYLIYKYRIRSYMDSEIRAIMAQYMPLDSQAEVVN 624
++ GGY YK RI+ YMDSEIR IMAQ MPLD +V N
Sbjct: 291 VAVAGGYAFYKXRIQRYMDSEIRTIMAQXMPLDXXPDVSN 172
>BU761053 similar to GP|22530930|gb| vacuolar sorting receptor-like protein
{Arabidopsis thaliana}, partial (12%)
Length = 446
Score = 110 bits (276), Expect = 1e-24
Identities = 60/104 (57%), Positives = 70/104 (66%)
Frame = +1
Query: 15 LLLLSVTPSSMARFVVEKNSLTVTSPEDIKGTHDSAIGNFGIPQYGGSMAGNVVYPKDNH 74
LLLL V ARFVVEK+S+TV SP +K D AIGNFG+P YGG + G+VVYP
Sbjct: 139 LLLLVVVVVVDARFVVEKSSITVLSPHKLKAKRDGAIGNFGLPDYGGFIVGSVVYPAKGS 318
Query: 75 KGCKEFDESGISFKSKPGALPTIVLLDRGNCFFALKVWNAQKAG 118
GC+ F E FK + PTIVLLDRG C+FALKVW+AQ AG
Sbjct: 319 HGCENF-EGDKPFKIQ-SYRPTIVLLDRGECYFALKVWHAQLAG 444
>BF426019
Length = 408
Score = 89.7 bits (221), Expect = 3e-18
Identities = 50/83 (60%), Positives = 61/83 (73%)
Frame = +3
Query: 333 SLGLDNKKIERCMGDPDADSDNPVLKEEQDAQVGKGSRGDVTILPTLVVNNRQYRGKLEK 392
S GL+ +KIE+CMGDPDAD+DNPVLKEEQDAQV + G + ++V+ YR L K
Sbjct: 69 SAGLNTEKIEKCMGDPDADTDNPVLKEEQDAQVLSFNNGYLVHCISVVI----YR-CLTK 233
Query: 393 GAVMKAICSGFEETTEPAVCLSS 415
+ AICSGFEETTEPAVCLS+
Sbjct: 234 LVFIIAICSGFEETTEPAVCLSN 302
>AW508719
Length = 116
Score = 59.3 bits (142), Expect = 5e-09
Identities = 21/31 (67%), Positives = 26/31 (83%)
Frame = +1
Query: 534 CSCKNTWGSYDCSCSGDLLYIRDHDTCISKT 564
CSCKNTWGSYDC C G+LLYI++ D CI ++
Sbjct: 22 CSCKNTWGSYDCKCKGNLLYIKEQDVCIERS 114
>TC215480 similar to GB|BAA96951.1|8777361|AB019233 protein carboxyl
methylase-like {Arabidopsis thaliana;} , partial (21%)
Length = 955
Score = 58.5 bits (140), Expect = 9e-09
Identities = 25/30 (83%), Positives = 29/30 (96%)
Frame = +1
Query: 335 GLDNKKIERCMGDPDADSDNPVLKEEQDAQ 364
GL+ +KIE+CMGDPDAD+DNPVLKEEQDAQ
Sbjct: 598 GLNTEKIEKCMGDPDADTDNPVLKEEQDAQ 687
>TC219557 similar to UP|Q9M621 (Q9M621) ReMembR-H2 protein JR702 (Fragment),
partial (42%)
Length = 838
Score = 47.4 bits (111), Expect = 2e-05
Identities = 47/158 (29%), Positives = 70/158 (43%)
Frame = +1
Query: 11 LLGFLLLLSVTPSSMARFVVEKNSLTVTSPEDIKGTHDSAIGNFGIPQYGGSMAGNVVYP 70
LL F L+S+ + ++ V+ N++T++ D NF P GS ++Y
Sbjct: 238 LLLFFSLVSLCAMAASKVVLIGNNITLS--------FDDIEANFA-PTVKGSGEYGILYL 390
Query: 71 KDNHKGCKEFDESGISFKSKPGALPTIVLLDRGNCFFALKVWNAQKAGASAVLVADDIEE 130
+ C E + P A L+ RG C F KV AQKAG AV+V D+ E
Sbjct: 391 AEPLDACTELTNK---VEQLPNASSPFALVVRGGCSFEEKVRIAQKAGFKAVIVYDNEEG 561
Query: 131 KLITMDTPEEDGSSAKYIENITIPSALIEKSFGEKLKK 168
++ G+SA I I + + K+ GE LKK
Sbjct: 562 GILVAMA----GNSA----GIRIHAVFVSKASGEILKK 651
>BQ298743
Length = 427
Score = 32.3 bits (72), Expect = 0.66
Identities = 17/30 (56%), Positives = 18/30 (59%), Gaps = 1/30 (3%)
Frame = +2
Query: 519 DVDECKEKKACQCPECS-CKNTWGSYDCSC 547
DVDEC EK C E S C N+ G Y CSC
Sbjct: 119 DVDECNEKTH-NCTEGSICSNSPGIYSCSC 205
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.136 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,019,409
Number of Sequences: 63676
Number of extensions: 531002
Number of successful extensions: 2641
Number of sequences better than 10.0: 54
Number of HSP's better than 10.0 without gapping: 2592
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2623
length of query: 631
length of database: 12,639,632
effective HSP length: 103
effective length of query: 528
effective length of database: 6,081,004
effective search space: 3210770112
effective search space used: 3210770112
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0218.20


