

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0214.6
(398 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
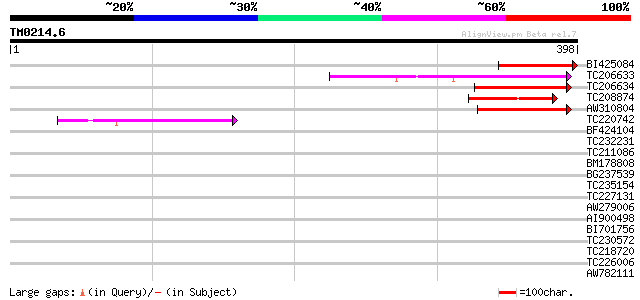
Score E
Sequences producing significant alignments: (bits) Value
BI425084 similar to GP|17381270|gb| AT3g24470/MXP5_4 {Arabidopsi... 120 1e-27
TC206633 111 5e-25
TC206634 78 8e-15
TC208874 69 4e-12
AW310804 68 8e-12
TC220742 52 5e-07
BF424104 39 0.005
TC232231 31 1.1
TC211086 similar to UP|Q7XJQ0 (Q7XJQ0) At2g37860 protein, partia... 31 1.1
BM178808 30 1.5
BG237539 30 1.5
TC235154 30 1.9
TC227131 similar to UP|Q8GSD7 (Q8GSD7) MRNA capping enzyme-like ... 30 2.5
AW279006 weakly similar to SP|P98204|ALA1 Phospholipid-transport... 29 3.3
AI900498 similar to GP|7239157|gb| homeodomain protein {Malus x ... 29 4.3
BI701756 homologue to GP|7239157|gb|A homeodomain protein {Malus... 29 4.3
TC230572 similar to UP|Q8LF32 (Q8LF32) Aspartyl aminopeptidase-l... 29 4.3
TC218720 similar to GB|AAO42946.1|28416831|BT004700 At3g18880 {A... 28 5.6
TC226006 similar to UP|REMO_SOLTU (P93788) Remorin (pp34), parti... 28 7.3
AW782111 28 7.3
>BI425084 similar to GP|17381270|gb| AT3g24470/MXP5_4 {Arabidopsis thaliana},
partial (12%)
Length = 425
Score = 120 bits (300), Expect = 1e-27
Identities = 51/55 (92%), Positives = 54/55 (97%)
Frame = +1
Query: 344 LLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLVAPIIWKSRQTDST 398
LLIGWNSHHSMRKWTIDVGWTSTWV+IVNEWLAVCVYLWML+APIIWK+RQT ST
Sbjct: 1 LLIGWNSHHSMRKWTIDVGWTSTWVKIVNEWLAVCVYLWMLIAPIIWKNRQTGST 165
>TC206633
Length = 959
Score = 111 bits (278), Expect = 5e-25
Identities = 58/187 (31%), Positives = 92/187 (49%), Gaps = 17/187 (9%)
Frame = +3
Query: 225 VLLQLMTSVSLHPKVNAGILTPGLMGLYVVFLCWNAIRSEPDGGSC--IRKSDSSTKTDW 282
+L + V+LHP VN +L ++ LY +LC++A+ SEP C + K + T
Sbjct: 3 ILAFVFAIVALHPAVNGSVLPASVISLYCTYLCYSALASEPRDYECNGLHKHSKAVSTGT 182
Query: 283 LSIISFIIAILAIVIATFSTGIDSKCFQ---------------FRKDDDIPAEDDVPYGY 327
L++ +L++V + G + ++D++ V Y Y
Sbjct: 183 LTL-GLATTVLSVVYSAVRAGSSAAVLSPPSSPRAGKPLLPLDAKEDEEKEKAKPVTYSY 359
Query: 328 GFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLVAP 387
FFH +F+ +MY AMLL GW++ +DVGW S WVRI+ W +YLW LVAP
Sbjct: 360 AFFHLIFSLASMYSAMLLTGWSTSVGESGKLVDVGWPSVWVRIITSWATALLYLWSLVAP 539
Query: 388 IIWKSRQ 394
I++ R+
Sbjct: 540 IMFPERE 560
>TC206634
Length = 410
Score = 77.8 bits (190), Expect = 8e-15
Identities = 31/68 (45%), Positives = 43/68 (62%)
Frame = +2
Query: 327 YGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLVA 386
Y FFH +F+ +MY AMLL GW++ +DVGW S WVRI+ W +YLW L+A
Sbjct: 11 YAFFHLIFSLASMYSAMLLTGWSTSVGESGKLVDVGWPSVWVRIITSWATALLYLWSLIA 190
Query: 387 PIIWKSRQ 394
PI++ R+
Sbjct: 191 PIMFPERE 214
>TC208874
Length = 769
Score = 68.9 bits (167), Expect = 4e-12
Identities = 32/62 (51%), Positives = 39/62 (62%)
Frame = +1
Query: 323 VPYGYGFFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLW 382
V Y Y FFH +FA +MY AMLL GW S IDVGWTS WVRI EW+ +Y+W
Sbjct: 334 VIYSYSFFHQIFALASMYSAMLLSGWTSTSES*D-LIDVGWTSVWVRIGTEWVTAGLYIW 510
Query: 383 ML 384
++
Sbjct: 511 IV 516
Score = 28.9 bits (63), Expect = 4.3
Identities = 9/25 (36%), Positives = 17/25 (68%)
Frame = +2
Query: 233 VSLHPKVNAGILTPGLMGLYVVFLC 257
++LHP+VN +L ++ LY ++C
Sbjct: 8 IALHPQVNGSLLPSAVISLYCAYVC 82
>AW310804
Length = 199
Score = 67.8 bits (164), Expect = 8e-12
Identities = 29/66 (43%), Positives = 41/66 (61%)
Frame = -1
Query: 329 FFHFVFATGAMYFAMLLIGWNSHHSMRKWTIDVGWTSTWVRIVNEWLAVCVYLWMLVAPI 388
F H +F+ G+MY AMLL GW + + + VGW S WVRI+ W+ +YLW VAPI
Sbjct: 199 FSHSIFSLGSMYCAMLLTGWFTSVAETGKLVHVGWLSVWVRIITFWVTALLYLWSFVAPI 20
Query: 389 IWKSRQ 394
++ R+
Sbjct: 19 MFLERE 2
>TC220742
Length = 881
Score = 52.0 bits (123), Expect = 5e-07
Identities = 35/132 (26%), Positives = 56/132 (41%), Gaps = 5/132 (3%)
Frame = +2
Query: 34 ARYVYALMFLVANLLAWAARDYGRGALTEMERLKGCNGGK-----DCLGAEGVLRVSLGC 88
AR Y +F + ++AW R+ A ME L N K + + VLRVSLG
Sbjct: 287 ARIAYCGLFAFSLVVAWILREV---AAPLMESLPWINHFKHTPSREWFETDAVLRVSLGN 457
Query: 89 FIFYFIMFLTTARTSKLNEVRDTWHSGWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHF 148
F+F+ I+ + + RD+ H G W +KI+ W + + F + +
Sbjct: 458 FLFFTILAVLMVGVKTQRDPRDSMHHGGWMMKIICWCLLVIFMFFVQMRSSASMKQYQRL 637
Query: 149 GAGVFLLIQLIS 160
F L++ S
Sbjct: 638 ARACFFLVKACS 673
>BF424104
Length = 409
Score = 38.5 bits (88), Expect = 0.005
Identities = 18/47 (38%), Positives = 28/47 (59%)
Frame = +1
Query: 112 WHSGWWSVKIVLWVGMTVIPFLLPSEFIQIYGEVAHFGAGVFLLIQL 158
+H G W+ KIV+W+ + V+ F LP I +YG + ++LL QL
Sbjct: 262 FHHGGWTAKIVIWLLLVVLAFFLPDAVILVYGIL------LYLLYQL 384
>TC232231
Length = 422
Score = 30.8 bits (68), Expect = 1.1
Identities = 13/24 (54%), Positives = 15/24 (62%)
Frame = -3
Query: 67 KGCNGGKDCLGAEGVLRVSLGCFI 90
+GCN K L A G+L LGCFI
Sbjct: 381 RGCNSRKGSLSAVGILYSDLGCFI 310
>TC211086 similar to UP|Q7XJQ0 (Q7XJQ0) At2g37860 protein, partial (73%)
Length = 1073
Score = 30.8 bits (68), Expect = 1.1
Identities = 15/58 (25%), Positives = 29/58 (49%), Gaps = 7/58 (12%)
Frame = +2
Query: 177 YAAKCQIHVMLFATTAYVVCLVGIILMFI-------WYAPKPSCLLNIFFIAWTLVLL 227
+A +C + F+ A + + ++L+ + + P SC+L IF + W L+LL
Sbjct: 275 FAIECWLIPHFFSKLAQRLSSIHVVLLLLKFKREAKTFGPNLSCILQIFLLGWLLMLL 448
>BM178808
Length = 422
Score = 30.4 bits (67), Expect = 1.5
Identities = 13/22 (59%), Positives = 15/22 (68%)
Frame = -3
Query: 69 CNGGKDCLGAEGVLRVSLGCFI 90
CN K LGA G+L +LGCFI
Sbjct: 408 CNSRKGSLGAVGILYSNLGCFI 343
>BG237539
Length = 199
Score = 30.4 bits (67), Expect = 1.5
Identities = 11/32 (34%), Positives = 18/32 (55%)
Frame = +2
Query: 234 SLHPKVNAGILTPGLMGLYVVFLCWNAIRSEP 265
+LHP N + + LY +LC++A+ EP
Sbjct: 41 ALHPAENGSVAPASKISLYCTYLCYSALAREP 136
>TC235154
Length = 437
Score = 30.0 bits (66), Expect = 1.9
Identities = 22/67 (32%), Positives = 33/67 (48%), Gaps = 5/67 (7%)
Frame = +1
Query: 34 ARYVYALMFLVANLLAWAARDYGRGALTEMERLKGCNGGK-----DCLGAEGVLRVSLGC 88
AR Y +F + ++AW R+ A ME L N K + + VLRVSLG
Sbjct: 241 ARIAYCGLFAFSLVVAWILREV---AAPLMESLPWINHFKHTPSREWFETDAVLRVSLGN 411
Query: 89 FIFYFIM 95
++F+ I+
Sbjct: 412 YLFFTIL 432
>TC227131 similar to UP|Q8GSD7 (Q8GSD7) MRNA capping enzyme-like protein,
partial (38%)
Length = 1812
Score = 29.6 bits (65), Expect = 2.5
Identities = 27/88 (30%), Positives = 42/88 (47%), Gaps = 1/88 (1%)
Frame = -2
Query: 158 LISIISFITWLNDC-CESEKYAAKCQIHVMLFATTAYVVCLVGIILMFIWYAPKPSCLLN 216
LI+IIS+I +L C CES I + + + V ++PKPS +L
Sbjct: 323 LIAIISYIRYLLSCFCES----GNVSITISPSSNVKWWV-----------FSPKPSFVLL 189
Query: 217 IFFIAWTLVLLQLMTSVSLHPKVNAGIL 244
+ WTL L+L+ S+ HP + I+
Sbjct: 188 QGNLIWTLRKLKLL-SIK*HPSIVISII 108
>AW279006 weakly similar to SP|P98204|ALA1 Phospholipid-transporting ATPase 1
(EC 3.6.3.1) (Aminophospholipid flippase 1). [Mouse-ear
cress], partial (7%)
Length = 424
Score = 29.3 bits (64), Expect = 3.3
Identities = 21/81 (25%), Positives = 35/81 (42%), Gaps = 13/81 (16%)
Frame = +3
Query: 325 YGYGFFHFVFATGAMYFAMLLIGWNS------------HHSMRKWTIDVGWTSTWVRIVN 372
YG G H + +F M+ W S ++ W++ WT + V +VN
Sbjct: 81 YGAGHRHEAYNMQLFWFTMIDTLWQSLVLFYIPVFIYKDSTIDIWSMGSLWTISVVILVN 260
Query: 373 EWLAVCVYLWMLVAPI-IWKS 392
LA+ + W LV+ + +W S
Sbjct: 261 VHLAMDINQWALVSHVAVWGS 323
>AI900498 similar to GP|7239157|gb| homeodomain protein {Malus x domestica},
partial (12%)
Length = 377
Score = 28.9 bits (63), Expect = 4.3
Identities = 13/23 (56%), Positives = 16/23 (69%)
Frame = -2
Query: 212 SCLLNIFFIAWTLVLLQLMTSVS 234
SCLLN+FF+ W L LQ +S S
Sbjct: 178 SCLLNLFFLNWPLPTLQNSSSNS 110
>BI701756 homologue to GP|7239157|gb|A homeodomain protein {Malus x
domestica}, partial (11%)
Length = 422
Score = 28.9 bits (63), Expect = 4.3
Identities = 13/23 (56%), Positives = 16/23 (69%)
Frame = -3
Query: 212 SCLLNIFFIAWTLVLLQLMTSVS 234
SCLLN+FF+ W L LQ +S S
Sbjct: 303 SCLLNLFFLNWPLPTLQNSSSNS 235
>TC230572 similar to UP|Q8LF32 (Q8LF32) Aspartyl aminopeptidase-like protein,
partial (25%)
Length = 507
Score = 28.9 bits (63), Expect = 4.3
Identities = 21/72 (29%), Positives = 30/72 (41%), Gaps = 1/72 (1%)
Frame = -3
Query: 220 IAWTLVLLQLMTSVSLHPKVNAGILTPGLMG-LYVVFLCWNAIRSEPDGGSCIRKSDSST 278
+ W LV L+ S H ++ P L L+ W A+ G +RKS S
Sbjct: 316 VEWFLVKKYLLPG*SFHGSLSESWWYPALRKRLFASSTAWKAV------GEALRKSIISE 155
Query: 279 KTDWLSIISFII 290
T WLS + I+
Sbjct: 154 ATAWLSSFAAIV 119
>TC218720 similar to GB|AAO42946.1|28416831|BT004700 At3g18880 {Arabidopsis
thaliana;} , partial (79%)
Length = 804
Score = 28.5 bits (62), Expect = 5.6
Identities = 9/28 (32%), Positives = 18/28 (64%)
Frame = -3
Query: 333 VFATGAMYFAMLLIGWNSHHSMRKWTID 360
V+ T + + ++L W+SH+ M WT++
Sbjct: 547 VWTTLSQFMKLVLQSWSSHYQMLAWTVE 464
>TC226006 similar to UP|REMO_SOLTU (P93788) Remorin (pp34), partial (64%)
Length = 991
Score = 28.1 bits (61), Expect = 7.3
Identities = 12/27 (44%), Positives = 19/27 (69%)
Frame = +1
Query: 187 LFATTAYVVCLVGIILMFIWYAPKPSC 213
L T+ ++V ++ I MFI+YA KP+C
Sbjct: 703 LINTSVFLVYMLEI*FMFIYYA*KPTC 783
>AW782111
Length = 418
Score = 28.1 bits (61), Expect = 7.3
Identities = 18/59 (30%), Positives = 32/59 (53%), Gaps = 4/59 (6%)
Frame = +2
Query: 183 IHVMLFATTAYVVCLVGIILMFIWYAPKPSCL----LNIFFIAWTLVLLQLMTSVSLHP 237
+H ++ T Y+ CL+ IL+ + + CL LNI ++ L+ LQL+T ++ P
Sbjct: 239 VHTIIVITEFYMNCLLQCILLLMHFI*G-LCLRY*KLNILSNSFLLLALQLLTKLTTQP 412
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.328 0.140 0.472
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,500,619
Number of Sequences: 63676
Number of extensions: 396103
Number of successful extensions: 3118
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 3082
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3115
length of query: 398
length of database: 12,639,632
effective HSP length: 99
effective length of query: 299
effective length of database: 6,335,708
effective search space: 1894376692
effective search space used: 1894376692
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0214.6


