

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.7
(451 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
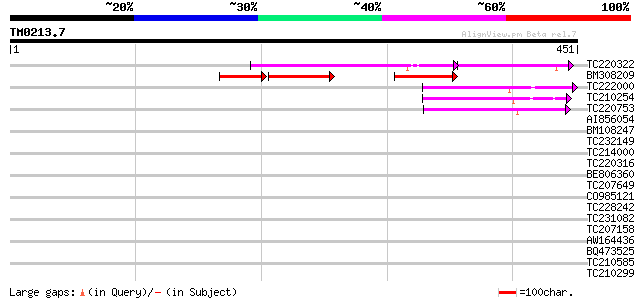
Score E
Sequences producing significant alignments: (bits) Value
TC220322 115 2e-42
BM308209 69 2e-22
TC222000 weakly similar to PIR|H84710|H84710 Mutator-like transp... 79 3e-15
TC210254 similar to GB|AAM26637.1|20855902|AY101516 AT3g06940/F1... 72 4e-13
TC220753 64 1e-10
AI856054 37 0.004
BM108247 weakly similar to PIR|H84710|H847 Mutator-like transpos... 38 0.011
TC232149 35 0.090
TC214000 33 0.26
TC220316 32 0.77
BE806360 similar to GP|6175165|gb| Mutator-like transposase {Ara... 31 1.0
TC207649 similar to UP|Q9SY66 (Q9SY66) F14N23.12 (At1g10240/F14N... 30 1.7
CO985121 30 2.9
TC228242 weakly similar to UP|Q9LIE5 (Q9LIE5) Far-red impaired r... 29 3.8
TC231082 weakly similar to UP|O83031 (O83031) Orf509e (Fragment)... 29 5.0
TC207158 29 5.0
AW164436 similar to GP|4519792|dbj| Asp1 {Arabidopsis thaliana},... 29 5.0
BQ473525 similar to GP|9988432|dbj| P0671B11.14 {Oryza sativa (j... 28 6.5
TC210585 similar to UP|Q9FX71 (Q9FX71) T6J4.1 protein, partial (... 28 6.5
TC210299 homologue to UP|EFT2_SOYBN (P46280) Elongation factor T... 28 6.5
>TC220322
Length = 1264
Score = 115 bits (288), Expect(2) = 2e-42
Identities = 67/169 (39%), Positives = 94/169 (54%), Gaps = 3/169 (1%)
Frame = +2
Query: 192 AKATYHQLWTEKMMELQALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNAT 251
AK+T + M L+ ++ +A+ +L + P +AW K F+ PK D + NN E FN
Sbjct: 8 AKSTTVAEFEGHMAHLKTINCQAWEYLNKWPKQAWTKAHFSTIPKVDNICNNTCEVFNFR 187
Query: 252 ILLVRDKPVLTMVDWIRSYIMGRFATMNEKLEKYKGEVMPKPLKRLEHEIEMVRYWVATR 311
IL R KP++TM++ IRSYIM A KL G + KRLE E W
Sbjct: 188 ILQYRCKPIITMLEEIRSYIMRTMAARKVKLSGKPGPLCLVQYKRLEKEFHFANQWTPIW 367
Query: 312 CGED---KYEVKHMVSQDQFVVNLAEHTCSCNFWELVGIPCRHAVAAIS 357
CG++ +YEV HM ++ VNLA TC+C W+L G+PCRHA+A I+
Sbjct: 368 CGDNMGLRYEV-HMWG-NKVEVNLAGWTCTCGVWQLTGMPCRHAIATIT 508
Score = 75.9 bits (185), Expect(2) = 2e-42
Identities = 39/99 (39%), Positives = 48/99 (48%), Gaps = 7/99 (7%)
Frame = +1
Query: 357 SHSSQRPEDFVHPYYRREAYMATYGHVITPINGQKLWPTTNDPYILPPLYKRAPGRPKKL 416
SH +PED H + EAY TY H I P+ G + W T + +PP + GRPKK
Sbjct: 508 SHKGGKPEDMCHEWLSIEAYNKTYQHFIEPVQGPQYWAQTQYTHPVPPYKRVQRGRPKKN 687
Query: 417 RRRDPHEDTTDNQTRLR-------SRCGQFGHNSRSCRN 448
R+R ED R RCGQ HN RSC+N
Sbjct: 688 RKRSVDEDNVTGHKLKRKLAEFTCGRCGQTNHNIRSCKN 804
>BM308209
Length = 435
Score = 68.6 bits (166), Expect(2) = 2e-22
Identities = 27/52 (51%), Positives = 41/52 (77%)
Frame = +2
Query: 207 LQALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLSESFNATILLVRDK 258
++ ++ EAY WL+++ T+ W KH F++Y KCDVLMNNL ++FN+TIL R+K
Sbjct: 122 IKEVNNEAYDWLIKVGTKLWRKHTFSYYTKCDVLMNNLCDAFNSTILCAREK 277
Score = 57.8 bits (138), Expect = 1e-08
Identities = 26/50 (52%), Positives = 32/50 (64%)
Frame = +3
Query: 307 WVATRCGEDKYEVKHMVSQDQFVVNLAEHTCSCNFWELVGIPCRHAVAAI 356
W AT G +EV H + D+F+VNL +CS N WE+ GI CRHA AAI
Sbjct: 285 WFATLSGHLVFEVSHSMLSDKFIVNLNSWSCS*NLWEITGILCRHACAAI 434
Score = 55.1 bits (131), Expect(2) = 2e-22
Identities = 23/37 (62%), Positives = 29/37 (78%)
Frame = +3
Query: 168 RHLYANFKKKFGGGVVIRNLMMAAAKATYHQLWTEKM 204
RH+Y NFK KF GG ++RNLMM AAKA++ + W EKM
Sbjct: 6 RHIYNNFKNKFVGGTLLRNLMMRAAKASFEEAWDEKM 116
>TC222000 weakly similar to PIR|H84710|H84710 Mutator-like transposase
[imported] - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (12%)
Length = 617
Score = 79.3 bits (194), Expect = 3e-15
Identities = 44/130 (33%), Positives = 63/130 (47%), Gaps = 7/130 (5%)
Frame = +2
Query: 329 VVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPIN 388
+VN+ H+CSC W+L GIPC HA AA+ + F + ++ TY I PI
Sbjct: 17 IVNIGSHSCSCRDWQLNGIPCSHAAAALISCRKDVYAFSQKCFTAASFRDTYAETIHPIP 196
Query: 389 GQKLWPTT-------NDPYILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLRSRCGQFGH 441
G+ W T N + PP +R PGRP+K R + + T SRC Q GH
Sbjct: 197 GKLEWSKTGNSSMDDNILVVRPPKLRRPPGRPEK--RMCVDDLNREKHTVHCSRCNQTGH 370
Query: 442 NSRSCRNPVV 451
R+C+ ++
Sbjct: 371 YKRTCKAEMI 400
>TC210254 similar to GB|AAM26637.1|20855902|AY101516 AT3g06940/F17A9_9
{Arabidopsis thaliana;} , partial (6%)
Length = 814
Score = 72.4 bits (176), Expect = 4e-13
Identities = 39/123 (31%), Positives = 59/123 (47%), Gaps = 4/123 (3%)
Frame = +1
Query: 329 VVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPIN 388
+V++ CSC W+L G+PC HA+A + P D+ Y+ E Y TY I P+
Sbjct: 64 IVDIDNWDCSCKGWQLTGVPCCHAIAVFECVGRSPYDYCSRYFTVENYRLTYAESIHPVP 243
Query: 389 GQKLWPTTNDP----YILPPLYKRAPGRPKKLRRRDPHEDTTDNQTRLRSRCGQFGHNSR 444
P + ++PP KR PGRPK ++ D Q + S+C GHN +
Sbjct: 244 NVDKPPVQGESTALVMVIPPPTKRPPGRPK--MKQVESIDIIKRQLQC-SKCKGLGHNRK 414
Query: 445 SCR 447
+C+
Sbjct: 415 TCK 423
>TC220753
Length = 1312
Score = 63.9 bits (154), Expect = 1e-10
Identities = 33/119 (27%), Positives = 55/119 (45%), Gaps = 2/119 (1%)
Frame = +2
Query: 330 VNLAEHTCSCNFWELVGIPCRHAVAAISHSSQRPEDFVHPYYRREAYMATYGHVITPING 389
V L E C ++ + +PC HA+AA + + +DFV P Y+ E Y H +
Sbjct: 215 VKLNEWLCDYGQFQALRLPCSHAIAACAFCNLNYDDFVDPVYKLENIFKVYQHHFHSLGS 394
Query: 390 QKLWPTTNDPYIL--PPLYKRAPGRPKKLRRRDPHEDTTDNQTRLRSRCGQFGHNSRSC 446
+ WP P+ + P ++ GRP R + +++ N+ + S C GHN +C
Sbjct: 395 EDTWPQYLGPHFMSDPSKRRQISGRPATTRIHNEMDESIPNRLKKCSLCKTEGHNQSNC 571
>AI856054
Length = 173
Score = 36.6 bits (83), Expect(2) = 0.004
Identities = 17/47 (36%), Positives = 32/47 (67%)
Frame = -1
Query: 244 LSESFNATILLVRDKPVLTMVDWIRSYIMGRFATMNEKLEKYKGEVM 290
+ ESFN+ IL R+K +++M++ I Y+M R+ T ++K+ +G V+
Sbjct: 143 ICESFNSVILEAREKLIVSMLEDIHIYLMERWDTNSKKVVAREGSVL 3
Score = 21.6 bits (44), Expect(2) = 0.004
Identities = 7/9 (77%), Positives = 9/9 (99%)
Frame = -3
Query: 236 KCDVLMNNL 244
KCDVL+NN+
Sbjct: 165 KCDVLLNNM 139
>BM108247 weakly similar to PIR|H84710|H847 Mutator-like transposase
[imported] - Arabidopsis thaliana, partial (4%)
Length = 265
Score = 37.7 bits (86), Expect = 0.011
Identities = 25/79 (31%), Positives = 40/79 (49%)
Frame = +2
Query: 186 NLMMAAAKATYHQLWTEKMMELQALHPEAYTWLMQIPTRAWCKHAFTFYPKCDVLMNNLS 245
+L+ A+ AT + EKM E++ + PEA WL Q W F + L +N+
Sbjct: 20 HLLWKASHATTTIAFKEKMGEIEEVSPEAAKWLQQFXPSQWALVHFK-GTRFGHLSSNI- 193
Query: 246 ESFNATILLVRDKPVLTMV 264
E FN IL R+ P++ ++
Sbjct: 194 EEFNKWILDTRELPIIXVI 250
>TC232149
Length = 1005
Score = 34.7 bits (78), Expect = 0.090
Identities = 33/141 (23%), Positives = 58/141 (40%), Gaps = 34/141 (24%)
Frame = -3
Query: 260 VLTMVDWIRSYIMGRFATMNEKLEKYKGEVMPKPLKRLEH-EIEMVR---YWVATRCGED 315
+L M + IR+YIM + + K+ ++P R+E ++E + WV G
Sbjct: 427 ILNMCEDIRTYIMCKMKSDKLKMATRLRPLVPMQQSRIEKGKVESNK*TANWVGDPDGT- 251
Query: 316 KYEVKHMVSQDQFVVNLAEHTCSCNFWELVGI---------------------------- 347
+++V H + V+L + +C W+L+GI
Sbjct: 250 RFKVCHY--DTRLNVDLQN*SFTCRMWQLIGIPLS*LNLSILDIMYCVTMNLMFFWFIYI 77
Query: 348 --PCRHAVAAISHSSQRPEDF 366
PCRHA+ AI ++ PE +
Sbjct: 76 GMPCRHAIVAIRYNYHIPEGY 14
>TC214000
Length = 659
Score = 33.1 bits (74), Expect = 0.26
Identities = 16/40 (40%), Positives = 24/40 (60%), Gaps = 6/40 (15%)
Frame = +3
Query: 415 KLRRRDPHEDTTDNQTRLRS------RCGQFGHNSRSCRN 448
+LRRR+ ED D++ R + RC + GHN R+C+N
Sbjct: 6 RLRRREVDED*NDHKLRGTNTTN*CWRCHKLGHNKRTCKN 125
>TC220316
Length = 988
Score = 31.6 bits (70), Expect = 0.77
Identities = 15/47 (31%), Positives = 27/47 (56%)
Frame = +1
Query: 307 WVATRCGEDKYEVKHMVSQDQFVVNLAEHTCSCNFWELVGIPCRHAV 353
++ RCG D KH+V+ + ++++ CSC +E G+ CRH +
Sbjct: 454 YMVRRCGNDME--KHVVTFNASNISIS---CSCQMFEYEGVLCRHVL 579
>BE806360 similar to GP|6175165|gb| Mutator-like transposase {Arabidopsis
thaliana}, partial (17%)
Length = 431
Score = 31.2 bits (69), Expect = 1.0
Identities = 21/72 (29%), Positives = 36/72 (49%), Gaps = 11/72 (15%)
Frame = +2
Query: 151 LTVFDD----LMGGVE-------HIFCLRHLYANFKKKFGGGVVIRNLMMAAAKATYHQL 199
LT+ D ++ GVE H FC++HL +F+K+F +++ NL+ AA A
Sbjct: 101 LTILSDRQKGIVDGVEASFPTAFHGFCMQHLSDSFRKEFNNTMLV-NLLWEAANALTELN 277
Query: 200 WTEKMMELQALH 211
++ L+ H
Sbjct: 278 LKQRFWRLKRYH 313
>TC207649 similar to UP|Q9SY66 (Q9SY66) F14N23.12 (At1g10240/F14N23_12),
partial (30%)
Length = 971
Score = 30.4 bits (67), Expect = 1.7
Identities = 14/40 (35%), Positives = 25/40 (62%), Gaps = 2/40 (5%)
Frame = +2
Query: 336 TCSCNFWELVGIPCRHAVAAISHSS--QRPEDFVHPYYRR 373
+CSC+ +E GI CRH++ +S + Q P+ ++ +RR
Sbjct: 428 SCSCHQFEFTGILCRHSLRVLSTGNCFQIPDRYLPIRWRR 547
>CO985121
Length = 535
Score = 29.6 bits (65), Expect = 2.9
Identities = 17/47 (36%), Positives = 22/47 (46%)
Frame = +2
Query: 403 PPLYKRAPGRPKKLRRRDPHEDTTDNQTRLRSRCGQFGHNSRSCRNP 449
PPLYK P +PKK + R T + T R H + +CR P
Sbjct: 5 PPLYK-TPKQPKKKKTRPTKTHTCNPNTTNNKREKNTPHVAPTCRTP 142
>TC228242 weakly similar to UP|Q9LIE5 (Q9LIE5) Far-red impaired response
protein; Mutator-like transposase-like protein;
phytochrome A signaling protein-like, partial (25%)
Length = 1354
Score = 29.3 bits (64), Expect = 3.8
Identities = 13/45 (28%), Positives = 24/45 (52%), Gaps = 2/45 (4%)
Frame = +2
Query: 333 AEHTCSCNFWELVGIPCRHAVAAISHSSQR--PEDFVHPYYRREA 375
+E +C C +E G CRHA+ + +S Q P ++ + ++A
Sbjct: 230 SELSCICRLFEYRGYLCRHALFVLQYSGQSVFPSQYILKRWTKDA 364
>TC231082 weakly similar to UP|O83031 (O83031) Orf509e (Fragment), partial
(13%)
Length = 1383
Score = 28.9 bits (63), Expect = 5.0
Identities = 17/52 (32%), Positives = 30/52 (57%), Gaps = 7/52 (13%)
Frame = +2
Query: 397 NDPYILPPLYKRAPGR------PKKLRRRDPH-EDTTDNQTRLRSRCGQFGH 441
+ P +PP ++AP R P+++RRR+PH + N+T ++R +F H
Sbjct: 182 HQPNRIPP--RQAPNRIHQNPRPRQIRRRNPHRRGSPHNRTTKKTRVFRFHH 331
>TC207158
Length = 1004
Score = 28.9 bits (63), Expect = 5.0
Identities = 13/38 (34%), Positives = 18/38 (47%)
Frame = -3
Query: 324 SQDQFVVNLAEHTCSCNFWELVGIPCRHAVAAISHSSQ 361
S Q V A+H CSCN +PC+H + H +
Sbjct: 885 SDGQSGVTAAKHYCSCNL-----LPCQHTTHLLMHHGE 787
>AW164436 similar to GP|4519792|dbj| Asp1 {Arabidopsis thaliana}, partial
(11%)
Length = 163
Score = 28.9 bits (63), Expect = 5.0
Identities = 13/27 (48%), Positives = 17/27 (62%)
Frame = -3
Query: 399 PYILPPLYKRAPGRPKKLRRRDPHEDT 425
PYILP L R P RP+ + R P E++
Sbjct: 155 PYILPSLRGRQPLRPRLVLRHFPREES 75
>BQ473525 similar to GP|9988432|dbj| P0671B11.14 {Oryza sativa (japonica
cultivar-group)}, partial (9%)
Length = 427
Score = 28.5 bits (62), Expect = 6.5
Identities = 10/13 (76%), Positives = 11/13 (83%)
Frame = +1
Query: 434 SRCGQFGHNSRSC 446
S CG FGHNSR+C
Sbjct: 202 SYCGNFGHNSRTC 240
>TC210585 similar to UP|Q9FX71 (Q9FX71) T6J4.1 protein, partial (30%)
Length = 440
Score = 28.5 bits (62), Expect = 6.5
Identities = 17/35 (48%), Positives = 19/35 (53%), Gaps = 2/35 (5%)
Frame = +2
Query: 404 PLYKRAPGRPKKLRRRDPHEDTT--DNQTRLRSRC 436
P KR P +P+ R RDPH D D RLR RC
Sbjct: 122 PQRKRMP-QPRAERLRDPHRDDAGRDLPPRLRRRC 223
>TC210299 homologue to UP|EFT2_SOYBN (P46280) Elongation factor Tu,
chloroplast precursor (EF-Tu), complete
Length = 1615
Score = 28.5 bits (62), Expect = 6.5
Identities = 20/61 (32%), Positives = 26/61 (41%), Gaps = 14/61 (22%)
Frame = +3
Query: 403 PPLYKRAPGRPKKLRRRDPHEDTTDNQ---TRLRSR-----------CGQFGHNSRSCRN 448
PPL R RP++LR++ H D + RLR R GQ S+ CR
Sbjct: 432 PPLRPRRLPRPRRLRQKHDHRRRPDGRRHPRRLRRRRPNAPNQRTHSLGQTSRRSQHCRL 611
Query: 449 P 449
P
Sbjct: 612 P 614
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.341 0.149 0.525
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,992,735
Number of Sequences: 63676
Number of extensions: 341715
Number of successful extensions: 2771
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 2729
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2763
length of query: 451
length of database: 12,639,632
effective HSP length: 100
effective length of query: 351
effective length of database: 6,272,032
effective search space: 2201483232
effective search space used: 2201483232
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0213.7


