

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.11
(579 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
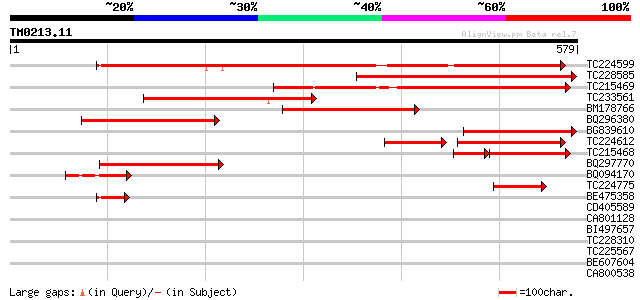
Sequences producing significant alignments: (bits) Value
TC224599 homologue to UP|G6PD_MEDSA (Q42919) Glucose-6-phosphate... 457 e-129
TC228585 homologue to GB|CAC05439.1|9955555|ATT19L5 glucose-6-ph... 392 e-109
TC215469 similar to UP|GPD4_ARATH (Q93ZW0) Glucose-6-phosphate 1... 359 2e-99
TC233561 homologue to UP|Q9LL88 (Q9LL88) Plastidic glucose 6-pho... 286 1e-77
BM178766 homologue to SP|Q43839|G6PC Glucose-6-phosphate 1-dehyd... 263 2e-70
BQ296380 similar to SP|Q43727|GPD1 Glucose-6-phosphate 1-dehydro... 249 2e-66
BG839610 similar to GP|1174336|emb glucose-6-phosphate 1-dehydro... 218 5e-57
TC224612 homologue to UP|Q9ZSR1 (Q9ZSR1) Cytoplasmic glucose-6-p... 106 3e-44
TC215468 similar to UP|GPD4_ARATH (Q93ZW0) Glucose-6-phosphate 1... 107 3e-33
BQ297770 139 5e-33
BQ094170 65 6e-11
TC224775 UP|O81978 (O81978) Glucose-6-phosphate-dehydrogenase (... 60 3e-09
BE475358 similar to GP|18086470|gb| AT3g27300/K17E12_12 {Arabido... 44 3e-04
CD405589 GP|10834744|gb plastidic glucose-6-phosphate dehydrogen... 40 0.003
CA801128 31 1.7
BI497657 similar to PIR|T47453|T47 oxidosqualene cyclase-like pr... 30 3.0
TC228310 similar to UP|Q6Z916 (Q6Z916) Pentatricopeptide (PPR) r... 29 6.6
TC225567 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp pro... 29 6.6
BE607604 similar to PIR|T52610|T526 glucose-6-phosphate 1-dehydr... 29 6.6
CA800538 similar to GP|22652121|gb| BEL1-related homeotic protei... 28 8.6
>TC224599 homologue to UP|G6PD_MEDSA (Q42919) Glucose-6-phosphate
1-dehydrogenase, cytoplasmic isoform (G6PD) , partial
(96%)
Length = 1866
Score = 457 bits (1175), Expect = e-129
Identities = 251/489 (51%), Positives = 331/489 (67%), Gaps = 10/489 (2%)
Frame = +3
Query: 89 TGSNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFL-VFGFARTKMTSEELRNMISR 147
TGS LSI V+GASGDLAKKK FPALF L+ + LP + + +FG+ARTK++ +ELRN +
Sbjct: 192 TGS-LSIVVLGASGDLAKKKTFPALFHLYLQGFLPPDEVHIFGYARTKISDDELRNRLRG 368
Query: 148 TLTCRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRS-----NRL 202
L + + +++FL+ Y SG Y+SE F LD ++ E E K S RL
Sbjct: 369 YLVPKKGASPQQLEDVEKFLQLIKYVSGSYDSEDGFRLLDKEISEHEYLKNSVEGLSRRL 548
Query: 203 FYLSIPPNIFVNV---VRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYLTED 259
FYL++PP+++ +V ++ + S GWTRV+VEKPFG+D ES+ EL+ + + E
Sbjct: 549 FYLALPPSVYPSVCKMIKTCCMNKSDLGGWTRVVVEKPFGKDLESAEELSTEIGKLFEEP 728
Query: 260 QIFRIDHYLGKELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYG 319
QI+RIDHYLGKELV+NL VLRF+N F PLW+R I NVQ +F EDFGT+GRGGYFD YG
Sbjct: 729 QIYRIDHYLGKELVQNLLVLRFANRFFLPLWNRDNIDNVQIVFREDFGTDGRGGYFDQYG 908
Query: 320 IIRDIMQNHLLQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKG 379
IIRDI+QNHLLQ+L L AME PVSL E IR+EKVKVL S+ P+ + VV GQY+
Sbjct: 909 IIRDIIQNHLLQVLCLVAMEKPVSLRPEHIRDEKVKVLESVLPINDDEVVLGQYE----- 1073
Query: 380 GKSYPGYADDPSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRH 439
GY DDP+VP S TPTFA L I N RW+GVPF++KAGKAL++++AEIRVQF+
Sbjct: 1074-----GYKDDPTVPDESNTPTFATVVLRIHNERWEGVPFILKAGKALNSRKAEIRVQFKD 1238
Query: 440 VPGNLYKRNFGADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYP 499
VPG+++K + NE V+R+QP EAIY+K+ K PGL M +S+L+L Y RY
Sbjct: 1239VPGDIFKCK-----KQGRNEFVIRLQPSEAIYMKLTVKQPGLEMSTVQSELDLSYGQRYQ 1403
Query: 500 -TEIPDAYERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGP 558
IP+AYERL+LD I G+++ F+R DEL A+W +FTPLL +++ + P Y GSRGP
Sbjct: 1404GVTIPEAYERLILDTIRGDQQHFVRRDELKASWQIFTPLLHRIDEGEFKPLPYKPGSRGP 1583
Query: 559 IGAHYLAAK 567
A L K
Sbjct: 1584AEADQLLEK 1610
>TC228585 homologue to GB|CAC05439.1|9955555|ATT19L5 glucose-6-phosphate
1-dehydrogenase {Arabidopsis thaliana;} , partial (37%)
Length = 1035
Score = 392 bits (1007), Expect = e-109
Identities = 184/224 (82%), Positives = 206/224 (91%)
Frame = +2
Query: 355 KVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADDPSVPKGSLTPTFAAAALFIDNARWD 414
KVLRSMRPL+L+++V GQYK H++GG +YP Y DD +VP GSLTPTFAAAALFIDNARWD
Sbjct: 2 KVLRSMRPLRLDDMVIGQYKSHTRGGVTYPAYVDDKTVPSGSLTPTFAAAALFIDNARWD 181
Query: 415 GVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNFGADMDKATNELVLRVQPDEAIYLKV 474
GVPFLMKAGKALH KRAEIRVQFRHVPGNLY RNFG D+D+ATNELV+RVQPDEAIYLK+
Sbjct: 182 GVPFLMKAGKALHNKRAEIRVQFRHVPGNLYNRNFGTDLDRATNELVIRVQPDEAIYLKI 361
Query: 475 NNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLLLDAIEGERRLFIRSDELDAAWALF 534
NNK+PGLGM+LDRS+LNL Y ARY EIPDAYERLLLDAIEGERRLFIRSDELDAAW+LF
Sbjct: 362 NNKVPGLGMKLDRSNLNLHYAARYSKEIPDAYERLLLDAIEGERRLFIRSDELDAAWSLF 541
Query: 535 TPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNVRWGDTGND 578
TP+LKE+E+KKI PE YP+GSRGP+GAHYLAA+HNVRWGD G D
Sbjct: 542 TPVLKELEEKKIIPEYYPYGSRGPVGAHYLAARHNVRWGDLGTD 673
>TC215469 similar to UP|GPD4_ARATH (Q93ZW0) Glucose-6-phosphate
1-dehydrogenase 4, chloroplast precursor (G6PD4)
(G6PDH4) , partial (45%)
Length = 979
Score = 359 bits (922), Expect = 2e-99
Identities = 179/303 (59%), Positives = 226/303 (74%)
Frame = +2
Query: 270 KELVENLSVLRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHL 329
+ L+ENL+VLRFSNLVFEPLWSR YI NVQ I SED G YF YGIIRDI+ +H+
Sbjct: 2 RNLIENLTVLRFSNLVFEPLWSRTYIDNVQVILSEDLAVHP-GRYFSGYGIIRDIVHSHV 178
Query: 330 LQILALFAMEPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADD 389
LQ +AL AMEPP+SLD EDIRNEK+KVLRS+R L+ ++V+ GQYK + GG +
Sbjct: 179 LQTIALLAMEPPISLDGEDIRNEKLKVLRSIRKLEPKDVILGQYK--TSGGAKVDACLN- 349
Query: 390 PSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPGNLYKRNF 449
LTPT+ AAAL+IDNARWDGVPFL+K G L + EIR+QFR+VPGN+Y
Sbjct: 350 ------GLTPTYFAAALYIDNARWDGVPFLIKTGLGLIKHQMEIRIQFRNVPGNVYHECI 511
Query: 450 GADMDKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERL 509
G +MD+A NEL+LR PDEAI ++VNNK+PGLG++LD S+LNLLY+ +Y E+PD+YE L
Sbjct: 512 GHNMDRAVNELILRDVPDEAILVRVNNKVPGLGLQLDSSELNLLYKDKYNMEVPDSYEHL 691
Query: 510 LLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHN 569
LLD I+G+ LF+RSDEL AAW + TP+L E++ ++ ELY G RGP+GA+YL AKH
Sbjct: 692 LLDVIDGDSHLFMRSDELAAAWNILTPILNEIDKDNMSVELYEMGGRGPVGAYYLWAKHG 871
Query: 570 VRW 572
VRW
Sbjct: 872 VRW 880
>TC233561 homologue to UP|Q9LL88 (Q9LL88) Plastidic glucose 6-phosphate
dehydrogenase , partial (30%)
Length = 631
Score = 286 bits (733), Expect = 1e-77
Identities = 144/202 (71%), Positives = 158/202 (77%), Gaps = 25/202 (12%)
Frame = +3
Query: 137 TSEELRNMISRTLTCRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGG 196
T ELRNM+S+TLTCRIDKR NC +KMDQFLKRCFYHSG Y+S+ +F+ LD K+KE EGG
Sbjct: 3 TDAELRNMVSKTLTCRIDKRENCNEKMDQFLKRCFYHSGQYDSQENFAALDKKLKEHEGG 182
Query: 197 KRSNRLFYLSIPPNIFVNVVRCASLKASSKKGWTRVIVEKPFGRDSESSNELTRCLKQYL 256
+ SNRLFYLSIPPNIF++ V+CASL ASS GWTRVIVEKPFGRDSESS LT+ LKQYL
Sbjct: 183 RTSNRLFYLSIPPNIFIDAVKCASLSASSGNGWTRVIVEKPFGRDSESSAALTKSLKQYL 362
Query: 257 TEDQIF-------------------------RIDHYLGKELVENLSVLRFSNLVFEPLWS 291
TEDQIF RIDHYLGKELVENLSVLRFSNL+FEPLWS
Sbjct: 363 TEDQIFRFRTKTDHFQL*DCKIHFIKWEVLYRIDHYLGKELVENLSVLRFSNLIFEPLWS 542
Query: 292 RAYIRNVQFIFSEDFGTEGRGG 313
R YIRNVQ IFSEDFGTEGRGG
Sbjct: 543 RQYIRNVQLIFSEDFGTEGRGG 608
>BM178766 homologue to SP|Q43839|G6PC Glucose-6-phosphate 1-dehydrogenase
chloroplast precursor (EC 1.1.1.49) (G6PD). [Potato],
partial (24%)
Length = 421
Score = 263 bits (671), Expect = 2e-70
Identities = 128/140 (91%), Positives = 132/140 (93%)
Frame = +2
Query: 279 LRFSNLVFEPLWSRAYIRNVQFIFSEDFGTEGRGGYFDNYGIIRDIMQNHLLQILALFAM 338
LRFSNLVFEPLWSR YIRNVQ IFSEDFGTEGRGGYFD+YGIIR IMQNHLLQILALFAM
Sbjct: 2 LRFSNLVFEPLWSRNYIRNVQIIFSEDFGTEGRGGYFDHYGIIRHIMQNHLLQILALFAM 181
Query: 339 EPPVSLDAEDIRNEKVKVLRSMRPLQLENVVTGQYKGHSKGGKSYPGYADDPSVPKGSLT 398
E PVSL AEDIRNEKVKVLRSMRPL+LENVV GQYKGHSKGGKS+P Y DDP+VPKGSLT
Sbjct: 182 ETPVSLAAEDIRNEKVKVLRSMRPLELENVVVGQYKGHSKGGKSHPAYTDDPTVPKGSLT 361
Query: 399 PTFAAAALFIDNARWDGVPF 418
PTFAAAALFIDNARWDGVPF
Sbjct: 362 PTFAAAALFIDNARWDGVPF 421
>BQ296380 similar to SP|Q43727|GPD1 Glucose-6-phosphate 1-dehydrogenase 1
chloroplast precursor (EC 1.1.1.49) (G6PD1) (G6PDH1).,
partial (21%)
Length = 425
Score = 249 bits (636), Expect = 2e-66
Identities = 122/141 (86%), Positives = 129/141 (90%)
Frame = +2
Query: 74 PQPVDGPFLSSDSECTGSNLSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFAR 133
PQP++GPF DSE TGSNLSITVVGASGDLAKKKIFPALFALFYED LPENFLVFGFAR
Sbjct: 2 PQPLEGPFPFPDSEYTGSNLSITVVGASGDLAKKKIFPALFALFYEDWLPENFLVFGFAR 181
Query: 134 TKMTSEELRNMISRTLTCRIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEK 193
TKMT EELRNMIS+TLTCRIDKR NCEDKMDQFLKRCFYHSG YNSE FS+LD K++EK
Sbjct: 182 TKMTYEELRNMISKTLTCRIDKRENCEDKMDQFLKRCFYHSGQYNSEDHFSELDSKLREK 361
Query: 194 EGGKRSNRLFYLSIPPNIFVN 214
EGGK SNRLFYLSIPPNIFV+
Sbjct: 362 EGGKLSNRLFYLSIPPNIFVD 424
>BG839610 similar to GP|1174336|emb glucose-6-phosphate 1-dehydrogenase
{Arabidopsis thaliana}, partial (22%)
Length = 529
Score = 218 bits (556), Expect = 5e-57
Identities = 102/115 (88%), Positives = 110/115 (94%)
Frame = +3
Query: 464 VQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYPTEIPDAYERLLLDAIEGERRLFIR 523
VQPDEAIYLK+ NK+PGLGMRLDRSDLNLL+RARYP EIPDAYERLLLDAIEGERRLFIR
Sbjct: 3 VQPDEAIYLKIXNKVPGLGMRLDRSDLNLLFRARYPREIPDAYERLLLDAIEGERRLFIR 182
Query: 524 SDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAKHNVRWGDTGND 578
SDELDAAWALFTPL KE+E+KKIAPELYP+GSRGP+GAHYLAA+HNVRWGD G D
Sbjct: 183 SDELDAAWALFTPLXKEIENKKIAPELYPYGSRGPVGAHYLAARHNVRWGDLGED 347
>TC224612 homologue to UP|Q9ZSR1 (Q9ZSR1) Cytoplasmic glucose-6-phosphate
1-dehydrogenase , partial (38%)
Length = 791
Score = 106 bits (265), Expect(2) = 3e-44
Identities = 54/111 (48%), Positives = 75/111 (66%), Gaps = 1/111 (0%)
Frame = +2
Query: 458 NELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDLNLLYRARYP-TEIPDAYERLLLDAIEG 516
NE V+R+QP EA+Y+K+ K PGL M +S+L+L Y RY IP+AYERL+LD I+G
Sbjct: 215 NEFVIRLQPSEAMYMKLTVKQPGLEMSTVQSELDLSYGQRYQGVTIPEAYERLILDTIKG 394
Query: 517 ERRLFIRSDELDAAWALFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAK 567
+++ F+R DEL A+W +FTPLL +++ + P Y GSRGP A L K
Sbjct: 395 DQQHFVRRDELKASWQIFTPLLHRIDEGEFKPLPYKPGSRGPAEADQLLEK 547
Score = 90.9 bits (224), Expect(2) = 3e-44
Identities = 40/64 (62%), Positives = 52/64 (80%)
Frame = +1
Query: 383 YPGYADDPSVPKGSLTPTFAAAALFIDNARWDGVPFLMKAGKALHTKRAEIRVQFRHVPG 442
Y GY DDP+VP S TPTFA L I N RW+GVPF++KAGKAL++++AEIRVQF+ VPG
Sbjct: 16 YEGYKDDPTVPDESNTPTFATVVLRIHNERWEGVPFILKAGKALNSRKAEIRVQFKDVPG 195
Query: 443 NLYK 446
+++K
Sbjct: 196 DIFK 207
>TC215468 similar to UP|GPD4_ARATH (Q93ZW0) Glucose-6-phosphate
1-dehydrogenase 4, chloroplast precursor (G6PD4)
(G6PDH4) , partial (18%)
Length = 804
Score = 107 bits (267), Expect(2) = 3e-33
Identities = 47/82 (57%), Positives = 62/82 (75%)
Frame = +3
Query: 491 NLLYRARYPTEIPDAYERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAPEL 550
NLLY+ +Y E+PD+YE LLLD I+G+ LF+RSDEL AAW + TP+L E++ ++ EL
Sbjct: 303 NLLYKDKYNMEVPDSYEHLLLDVIDGDSHLFMRSDELAAAWNILTPILNEIDKDNMSVEL 482
Query: 551 YPHGSRGPIGAHYLAAKHNVRW 572
Y G RGP+GA+YL AKH VRW
Sbjct: 483 YEMGGRGPVGAYYLWAKHGVRW 548
Score = 53.1 bits (126), Expect(2) = 3e-33
Identities = 23/37 (62%), Positives = 32/37 (86%)
Frame = +2
Query: 454 DKATNELVLRVQPDEAIYLKVNNKIPGLGMRLDRSDL 490
D+A NEL+LR PDEAI ++VN+K+PGLG++LD S+L
Sbjct: 191 DRAVNELILRDVPDEAILVRVNHKVPGLGLQLDSSEL 301
>BQ297770
Length = 421
Score = 139 bits (349), Expect = 5e-33
Identities = 61/127 (48%), Positives = 96/127 (75%)
Frame = +3
Query: 92 NLSITVVGASGDLAKKKIFPALFALFYEDCLPENFLVFGFARTKMTSEELRNMISRTLTC 151
+L I V+GA+G+LAK+KIFPALFAL+Y LPEN +FG++R +T E+L+++I+ TLTC
Sbjct: 27 SLCIAVIGATGELAKRKIFPALFALYYSGFLPENVGIFGYSRKDITDEDLQSIIASTLTC 206
Query: 152 RIDKRANCEDKMDQFLKRCFYHSGLYNSEGSFSDLDCKMKEKEGGKRSNRLFYLSIPPNI 211
R+D + NC+DK++ FL R +Y +G Y+++ S L+ +M++ EGG ++NR+FYLS+P
Sbjct: 207 RVDHQENCDDKLNAFLSRTYYINGGYDNKYGMSMLNARMEQIEGGSKTNRIFYLSVPQEA 386
Query: 212 FVNVVRC 218
++V C
Sbjct: 387 LLDVASC 407
>BQ094170
Length = 421
Score = 65.5 bits (158), Expect = 6e-11
Identities = 37/67 (55%), Positives = 46/67 (68%)
Frame = +1
Query: 58 SHDGLAGSSLTKEDSKPQPVDGPFLSSDSECTGSNLSITVVGASGDLAKKKIFPALFALF 117
SH L L+ + P + F ++E S++SITVVGASGDLAKKKIFPALFAL+
Sbjct: 235 SHKQLKADLLSLKS--PDESEDGFEKDENE---SSVSITVVGASGDLAKKKIFPALFALY 399
Query: 118 YEDCLPE 124
YEDCLP+
Sbjct: 400 YEDCLPK 420
>TC224775 UP|O81978 (O81978) Glucose-6-phosphate-dehydrogenase (Fragment) ,
complete
Length = 424
Score = 60.1 bits (144), Expect = 3e-09
Identities = 27/55 (49%), Positives = 39/55 (70%), Gaps = 1/55 (1%)
Frame = +3
Query: 495 RARYP-TEIPDAYERLLLDAIEGERRLFIRSDELDAAWALFTPLLKEVEDKKIAP 548
R RY IP+AYERL+LD I G+++ F+R DEL A+W +FTPLL +++ + P
Sbjct: 3 RQRYQGVTIPEAYERLILDTIRGDQQHFVRRDELKASWQIFTPLLHRIDEGEFKP 167
>BE475358 similar to GP|18086470|gb| AT3g27300/K17E12_12 {Arabidopsis
thaliana}, partial (8%)
Length = 170
Score = 43.5 bits (101), Expect = 3e-04
Identities = 24/34 (70%), Positives = 27/34 (78%)
Frame = +1
Query: 89 TGSNLSITVVGASGDLAKKKIFPALFALFYEDCL 122
TGS LSI V+GASGDLAKKK FPALF L+ + L
Sbjct: 49 TGS-LSIVVLGASGDLAKKKTFPALFNLYRQGFL 147
>CD405589 GP|10834744|gb plastidic glucose-6-phosphate dehydrogenase
{Cucurbita pepo}, partial (9%)
Length = 591
Score = 40.0 bits (92), Expect = 0.003
Identities = 18/20 (90%), Positives = 19/20 (95%)
Frame = -1
Query: 260 QIFRIDHYLGKELVENLSVL 279
+ FRIDHYLGKELVENLSVL
Sbjct: 558 KFFRIDHYLGKELVENLSVL 499
>CA801128
Length = 438
Score = 30.8 bits (68), Expect = 1.7
Identities = 25/64 (39%), Positives = 36/64 (56%), Gaps = 1/64 (1%)
Frame = +1
Query: 184 SDLDCKMKEKEGGKRSNRLFYLSIPPNIFVNVVRCASLKASSKKG-WTRVIVEKPFGRDS 242
+DL K + + RS+R SIP N+ + V+ +A+SKK WTR+I +KP R S
Sbjct: 13 TDLAIKWRREAINHRSSRT---SIPNNLSLLVI*A---QATSKKCFWTRMIFKKPIMRFS 174
Query: 243 ESSN 246
S N
Sbjct: 175 MSRN 186
>BI497657 similar to PIR|T47453|T47 oxidosqualene cyclase-like protein -
Arabidopsis thaliana, partial (17%)
Length = 407
Score = 30.0 bits (66), Expect = 3.0
Identities = 16/42 (38%), Positives = 23/42 (54%), Gaps = 6/42 (14%)
Frame = -1
Query: 181 GSFSDLDCKMKEKEG------GKRSNRLFYLSIPPNIFVNVV 216
G+FS +D K+ +G K R FY+ PP+IFV V+
Sbjct: 293 GTFSTIDVFSKQSQGHIVESSAKHCARPFYMRSPPSIFVLVI 168
>TC228310 similar to UP|Q6Z916 (Q6Z916) Pentatricopeptide (PPR)
repeat-containing protein-like, partial (48%)
Length = 600
Score = 28.9 bits (63), Expect = 6.6
Identities = 16/36 (44%), Positives = 21/36 (57%), Gaps = 3/36 (8%)
Frame = +3
Query: 21 PQLYPV--WFSCI-RPGNLPRNHFQLKSSNGHPLNA 53
PQ YP +F C+ RP LPR+ +NGHP +A
Sbjct: 222 PQFYPCLSFFRCLHRPSLLPRHSRSSIPTNGHPESA 329
>TC225567 homologue to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor ,
partial (61%)
Length = 2077
Score = 28.9 bits (63), Expect = 6.6
Identities = 20/99 (20%), Positives = 45/99 (45%), Gaps = 5/99 (5%)
Frame = +3
Query: 226 KKGWTRVIVEKPFGRDSESSNEL----TRCLKQYLTEDQIFRIDHYLGKELVENLSVLRF 281
+KG ++ + + S N + T LKQY + + R+D + + L V
Sbjct: 1236 EKGGRKIGFDLDYDEKDSSYNRIKSLVTEELKQYFRPEFLNRLDEMIVFRQLTKLEVKEI 1415
Query: 282 SNLVFEPLWSRAYIRNVQFIFSEDFGTE-GRGGYFDNYG 319
++++ + ++ R +++++ +E F GY +YG
Sbjct: 1416 ADIMLKEVFDRLKVKDIELQVTERFRDRVVEEGYNPSYG 1532
>BE607604 similar to PIR|T52610|T526 glucose-6-phosphate 1-dehydrogenase (EC
1.1.1.49) [imported] - Arabidopsis thaliana, partial
(10%)
Length = 414
Score = 28.9 bits (63), Expect = 6.6
Identities = 15/47 (31%), Positives = 22/47 (45%), Gaps = 7/47 (14%)
Frame = +2
Query: 533 LFTPLLKEVEDKKIAPELYPHGSRGPIGAHYLAAK-------HNVRW 572
+FTPLL ++ + Y G+RGP+ A L + H RW
Sbjct: 5 IFTPLLHRIDKGEFKSTPYQPGTRGPVEADKLLLEKAGYVQTHGYRW 145
>CA800538 similar to GP|22652121|gb| BEL1-related homeotic protein 14
{Solanum tuberosum}, partial (7%)
Length = 433
Score = 28.5 bits (62), Expect = 8.6
Identities = 18/66 (27%), Positives = 34/66 (51%)
Frame = +3
Query: 22 QLYPVWFSCIRPGNLPRNHFQLKSSNGHPLNAVSSHSHDGLAGSSLTKEDSKPQPVDGPF 81
Q + + S P + F+L+ + HP + VSS S +G G ++ + + DG +
Sbjct: 153 QGFSLSLSSTNPSTIGLQSFELRQTGHHP-DFVSSSSREGFFGKPVSLQQQQMLSQDG-Y 326
Query: 82 LSSDSE 87
+SS+S+
Sbjct: 327 VSSNSK 344
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.138 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,328,039
Number of Sequences: 63676
Number of extensions: 352389
Number of successful extensions: 1512
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 1496
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1501
length of query: 579
length of database: 12,639,632
effective HSP length: 102
effective length of query: 477
effective length of database: 6,144,680
effective search space: 2931012360
effective search space used: 2931012360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0213.11


