

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0213.1
(305 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
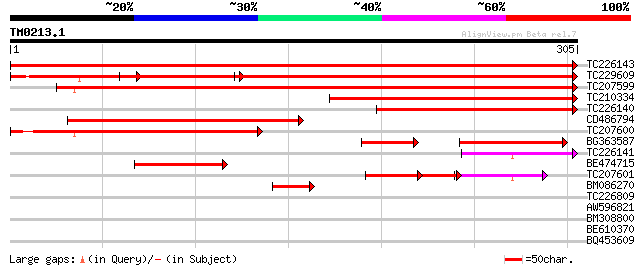
Score E
Sequences producing significant alignments: (bits) Value
TC226143 similar to UP|O82200 (O82200) Expressed protein, partia... 578 e-166
TC229609 similar to UP|Q6XGZ9 (Q6XGZ9) Myo-inositol oxygenase, p... 325 e-112
TC207599 weakly similar to UP|O82200 (O82200) Expressed protein,... 402 e-112
TC210334 weakly similar to UP|MIOX_PIG (Q8WN98) Inositol oxygena... 228 3e-60
TC226140 weakly similar to UP|MIOX_PIG (Q8WN98) Inositol oxygena... 218 4e-57
CD486794 204 3e-53
TC207600 similar to UP|O82200 (O82200) Expressed protein, partia... 160 9e-40
BG363587 92 4e-29
TC226141 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, ... 102 2e-22
BE474715 78 4e-15
TC207601 similar to UP|Q9MA30 (Q9MA30) T5E21.2, partial (20%) 57 7e-14
BM086270 41 8e-04
TC226809 similar to GB|AAP37699.1|30725354|BT008340 At5g20700 {A... 33 0.16
AW596821 28 4.0
BM308800 27 8.8
BE610370 27 8.8
BQ453609 homologue to GP|13605607|gb| T3B23.2/T3B23.2 {Arabidops... 27 8.8
>TC226143 similar to UP|O82200 (O82200) Expressed protein, partial (84%)
Length = 1560
Score = 578 bits (1491), Expect = e-166
Identities = 268/306 (87%), Positives = 291/306 (94%), Gaps = 1/306 (0%)
Frame = +2
Query: 1 MTIIIDQPDHFGSDVEGKTVPIDEKELVLDGGFVMPHTNSFGHTFRDYN-AESERQEGVE 59
MTI+I+Q DHFG++VE KT PI+E+ELVLDGGF++ TNSFGH FR YN AESER EGVE
Sbjct: 395 MTILIEQSDHFGAEVEAKTEPINERELVLDGGFLITQTNSFGHMFRAYNDAESERHEGVE 574
Query: 60 NFYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLNEVVDESDPDLDEPQIEHLLQ 119
NFYRKNH YQS+DFVKKMREEY KLNRVEMS+WECCELLNEVVDESDPDLDEPQIEHLLQ
Sbjct: 575 NFYRKNHTYQSIDFVKKMREEYAKLNRVEMSIWECCELLNEVVDESDPDLDEPQIEHLLQ 754
Query: 120 TAEAIRKDYPNEDWLHLTGLIHDLGKVLLLPVFGGLPQWAVVGDTFPVGCRFDESIVHHK 179
TAEAIR+DYP+EDWLHLTGLIHDLGKVL LP FGGLPQWAVVGD FPVGCRFDESIVHHK
Sbjct: 755 TAEAIRRDYPDEDWLHLTGLIHDLGKVLNLPSFGGLPQWAVVGDIFPVGCRFDESIVHHK 934
Query: 180 HFKENPDYNNPAYNTRYGMYSEKCGLNNVLMSWGHDDYMYLVAKENKTTLPSAALFIIRY 239
+FKENPDYNNPAYNT+YG+YSE+CGLNNV+MSWGHDDYMYLVAKEN TTLPSAALFIIRY
Sbjct: 935 YFKENPDYNNPAYNTKYGIYSERCGLNNVMMSWGHDDYMYLVAKENDTTLPSAALFIIRY 1114
Query: 240 HSFYALHREGAYKHLMNEEDFENLEWLHIFNKYDLYSKSKVRIDVEKVKPYYLSLIEKYF 299
HSFYALH+EGAYKHLMN+EDFENL+WLHIFNKYDLYSKSK+RIDVE+V+PYYLSLIEKYF
Sbjct: 1115HSFYALHKEGAYKHLMNDEDFENLKWLHIFNKYDLYSKSKIRIDVERVRPYYLSLIEKYF 1294
Query: 300 PAKLKW 305
PAKLKW
Sbjct: 1295PAKLKW 1312
>TC229609 similar to UP|Q6XGZ9 (Q6XGZ9) Myo-inositol oxygenase, partial (85%)
Length = 1196
Score = 325 bits (832), Expect(2) = e-112
Identities = 145/184 (78%), Positives = 164/184 (88%)
Frame = +2
Query: 122 EAIRKDYPNEDWLHLTGLIHDLGKVLLLPVFGGLPQWAVVGDTFPVGCRFDESIVHHKHF 181
+ IRKDYPNEDWLHLT LIHDLGK+L LP FG LPQWAVVGDTFP+GC FDES VHHK+F
Sbjct: 461 KTIRKDYPNEDWLHLTALIHDLGKILALPSFGELPQWAVVGDTFPLGCAFDESNVHHKYF 640
Query: 182 KENPDYNNPAYNTRYGMYSEKCGLNNVLMSWGHDDYMYLVAKENKTTLPSAALFIIRYHS 241
K+NPDY PAY+T+ G+Y+E CGL+N++MSWGHDDYMY+VAK N TTLPSA LFIIRYHS
Sbjct: 641 KDNPDYKCPAYSTKNGIYTEGCGLDNIVMSWGHDDYMYMVAKANGTTLPSAGLFIIRYHS 820
Query: 242 FYALHREGAYKHLMNEEDFENLEWLHIFNKYDLYSKSKVRIDVEKVKPYYLSLIEKYFPA 301
FY LH+EGAY H MNEED ENL+WL IFNKYDLYSKSKV +DVEKVKPYY+SLIEKYFPA
Sbjct: 821 FYPLHKEGAYTHFMNEEDVENLKWLKIFNKYDLYSKSKVLVDVEKVKPYYVSLIEKYFPA 1000
Query: 302 KLKW 305
K++W
Sbjct: 1001KVRW 1012
Score = 100 bits (248), Expect(2) = e-112
Identities = 46/67 (68%), Positives = 56/67 (82%)
Frame = +1
Query: 60 NFYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLNEVVDESDPDLDEPQIEHLLQ 119
NF N ++ DFVK+MREEYGKL++ EM +WECCELLNE+VDESDPDLDEPQI+HLLQ
Sbjct: 274 NFIGCNTSTKTYDFVKRMREEYGKLDKAEMGIWECCELLNELVDESDPDLDEPQIQHLLQ 453
Query: 120 TAEAIRK 126
+AE +K
Sbjct: 454 SAEDHQK 474
Score = 74.7 bits (182), Expect = 5e-14
Identities = 42/79 (53%), Positives = 52/79 (65%), Gaps = 8/79 (10%)
Frame = +3
Query: 1 MTIIIDQPDHFGSDVEGKTVPIDE-KELVLDGGFVMP-------HTNSFGHTFRDYNAES 52
MTI+I+QP VEG V +E ELVL+GGF +P NSFGH+FR+Y+AES
Sbjct: 75 MTILIEQPA-LELQVEGNNVHAEETNELVLEGGFPLPKDGYMAPEINSFGHSFREYDAES 251
Query: 53 ERQEGVENFYRKNHIYQSV 71
ERQ+GVE FYR HI Q +
Sbjct: 252 ERQKGVEEFYRLQHINQDI 308
>TC207599 weakly similar to UP|O82200 (O82200) Expressed protein, partial
(84%)
Length = 1287
Score = 402 bits (1032), Expect = e-112
Identities = 185/291 (63%), Positives = 225/291 (76%), Gaps = 11/291 (3%)
Frame = +3
Query: 26 ELVLDGGF-----------VMPHTNSFGHTFRDYNAESERQEGVENFYRKNHIYQSVDFV 74
+LVLDGGF +P NS+GH+F +Y ESER++ VE YR+ H Q+ +F
Sbjct: 9 DLVLDGGFSLLKSPSHVGFTVPDINSYGHSFTNYYDESERKKSVEELYRQQHTNQTYEFA 188
Query: 75 KKMREEYGKLNRVEMSMWECCELLNEVVDESDPDLDEPQIEHLLQTAEAIRKDYPNEDWL 134
M +EYGKLN+V++ +W CCE+L+++V +DPDL + QI H LQT E DYPNE WL
Sbjct: 189 NNMTQEYGKLNKVQLPIWPCCEMLDKIVHATDPDLQKSQIXHALQTXEPAXNDYPNEHWL 368
Query: 135 HLTGLIHDLGKVLLLPVFGGLPQWAVVGDTFPVGCRFDESIVHHKHFKENPDYNNPAYNT 194
HLT LIHDLGKVLLLP FGGLPQWAVVGD+FP+GC F E+ H K+FK+NPD NNPAYNT
Sbjct: 369 HLTALIHDLGKVLLLPSFGGLPQWAVVGDSFPLGCAFHEANTHFKYFKDNPDSNNPAYNT 548
Query: 195 RYGMYSEKCGLNNVLMSWGHDDYMYLVAKENKTTLPSAALFIIRYHSFYALHREGAYKHL 254
+ G+Y E CGL+NV+MSWGHD+YMYLV K N TTLPS ALFIIRYHSF+ L++ GAYKHL
Sbjct: 549 KKGIYDEGCGLDNVMMSWGHDEYMYLVTKGNNTTLPSEALFIIRYHSFHPLYQTGAYKHL 728
Query: 255 MNEEDFENLEWLHIFNKYDLYSKSKVRIDVEKVKPYYLSLIEKYFPAKLKW 305
+EED +NL+WL IF+KYDLYSKS V IDVEKVKPYY+SLI+KYFP KL W
Sbjct: 729 YSEEDVKNLKWLEIFHKYDLYSKSNVLIDVEKVKPYYMSLIKKYFPPKLIW 881
>TC210334 weakly similar to UP|MIOX_PIG (Q8WN98) Inositol oxygenase
(Myo-inositol oxygenase) (Aldehyde reductase-like 6) ,
partial (20%)
Length = 820
Score = 228 bits (580), Expect = 3e-60
Identities = 103/133 (77%), Positives = 116/133 (86%)
Frame = +3
Query: 173 ESIVHHKHFKENPDYNNPAYNTRYGMYSEKCGLNNVLMSWGHDDYMYLVAKENKTTLPSA 232
ES VHHKHFK+ PD NP YNT+ G+Y E GL+NV+MSWGHD+YMYLVAKEN TTLP
Sbjct: 3 ESNVHHKHFKDYPDNTNPTYNTKNGIYKEGIGLDNVVMSWGHDEYMYLVAKENGTTLPPV 182
Query: 233 ALFIIRYHSFYALHREGAYKHLMNEEDFENLEWLHIFNKYDLYSKSKVRIDVEKVKPYYL 292
ALFII+YHSFYALHR GAY HLMNEED ENL+WL IF+KYDLYSKSKV +D+EKVKPYY+
Sbjct: 183 ALFIIKYHSFYALHRAGAYTHLMNEEDIENLKWLKIFSKYDLYSKSKVLVDIEKVKPYYM 362
Query: 293 SLIEKYFPAKLKW 305
SLIEKYFPAKL+W
Sbjct: 363 SLIEKYFPAKLRW 401
>TC226140 weakly similar to UP|MIOX_PIG (Q8WN98) Inositol oxygenase
(Myo-inositol oxygenase) (Aldehyde reductase-like 6) ,
partial (19%)
Length = 495
Score = 218 bits (554), Expect = 4e-57
Identities = 99/108 (91%), Positives = 106/108 (97%)
Frame = +3
Query: 198 MYSEKCGLNNVLMSWGHDDYMYLVAKENKTTLPSAALFIIRYHSFYALHREGAYKHLMNE 257
+YSE+CGLNNV+MSWGHDDYMYLVAKEN TTLPSAALFIIRYHSFYALHREGAYKHL+N+
Sbjct: 9 IYSERCGLNNVMMSWGHDDYMYLVAKENSTTLPSAALFIIRYHSFYALHREGAYKHLLND 188
Query: 258 EDFENLEWLHIFNKYDLYSKSKVRIDVEKVKPYYLSLIEKYFPAKLKW 305
ED ENL+WLHIFNKYDLYSKSKVRIDVE+VKPYYLSLIEKYFPAKLKW
Sbjct: 189 EDVENLKWLHIFNKYDLYSKSKVRIDVERVKPYYLSLIEKYFPAKLKW 332
>CD486794
Length = 569
Score = 204 bits (520), Expect = 3e-53
Identities = 92/127 (72%), Positives = 109/127 (85%)
Frame = +2
Query: 32 GFVMPHTNSFGHTFRDYNAESERQEGVENFYRKNHIYQSVDFVKKMREEYGKLNRVEMSM 91
GF N+FG++FR+Y+AESERQ VE YR HI Q+ D+ K+MREEYGKL+++EM +
Sbjct: 188 GFNATDMNAFGNSFRNYDAESERQRMVEELYRMQHINQTYDYGKRMREEYGKLDKLEMGI 367
Query: 92 WECCELLNEVVDESDPDLDEPQIEHLLQTAEAIRKDYPNEDWLHLTGLIHDLGKVLLLPV 151
WECCELLNEV D+SDPDLDEPQI+HLLQ+AEAIRKDYPNEDWLHLT LIHDLGK+L+LP
Sbjct: 368 WECCELLNEVTDDSDPDLDEPQIQHLLQSAEAIRKDYPNEDWLHLTALIHDLGKILMLPS 547
Query: 152 FGGLPQW 158
FGGLPQW
Sbjct: 548 FGGLPQW 568
>TC207600 similar to UP|O82200 (O82200) Expressed protein, partial (35%)
Length = 468
Score = 160 bits (404), Expect = 9e-40
Identities = 77/147 (52%), Positives = 101/147 (68%), Gaps = 11/147 (7%)
Frame = +2
Query: 1 MTIIIDQPDHFGSDVEGKTVPIDEKELVLDGGF-----------VMPHTNSFGHTFRDYN 49
M ++I+Q S++ + +LVLDGGF +P NS+GH+FR+Y
Sbjct: 41 MVVLIEQ-----SELTRPQAEENNDDLVLDGGFSLLKSPSHVGFTVPDINSYGHSFRNYY 205
Query: 50 AESERQEGVENFYRKNHIYQSVDFVKKMREEYGKLNRVEMSMWECCELLNEVVDESDPDL 109
ESER + VE YR+ H Q+ +F KKMREEYGKLN+VE+ +WECCE+L+++VD SDPDL
Sbjct: 206 DESERNKSVEELYRQQHTNQTYEFAKKMREEYGKLNKVEVGIWECCEMLDKIVDASDPDL 385
Query: 110 DEPQIEHLLQTAEAIRKDYPNEDWLHL 136
+E QI+H LQTAEA R DYPNEDWLHL
Sbjct: 386 EESQIQHALQTAEAARNDYPNEDWLHL 466
>BG363587
Length = 270
Score = 91.7 bits (226), Expect(2) = 4e-29
Identities = 41/58 (70%), Positives = 46/58 (78%)
Frame = +1
Query: 243 YALHREGAYKHLMNEEDFENLEWLHIFNKYDLYSKSKVRIDVEKVKPYYLSLIEKYFP 300
YALH+EGAY H MNEE ENL+WL IFNKY LY+K K + VEKVKPYY+SLI YFP
Sbjct: 97 YALHKEGAYTHFMNEEXVENLKWLKIFNKYHLYTKXKPLVXVEKVKPYYVSLIXNYFP 270
Score = 54.3 bits (129), Expect(2) = 4e-29
Identities = 20/31 (64%), Positives = 26/31 (83%)
Frame = +2
Query: 190 PAYNTRYGMYSEKCGLNNVLMSWGHDDYMYL 220
PAY+T+ G+Y+ CGL N+ MSWGHDDYMY+
Sbjct: 8 PAYSTKNGIYTXGCGLXNIXMSWGHDDYMYM 100
>TC226141 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, partial
(57%)
Length = 1062
Score = 102 bits (255), Expect = 2e-22
Identities = 54/101 (53%), Positives = 61/101 (59%), Gaps = 39/101 (38%)
Frame = +3
Query: 244 ALHREGAYKHLMNEEDFENLEWLHIF---------------------------------- 269
ALHREGAYKHL+N+ED ENL+WLHIF
Sbjct: 9 ALHREGAYKHLLNDEDVENLKWLHIFK*HYILLIKYIGEW*LATKCVLCKDISIFSLTFF 188
Query: 270 -----NKYDLYSKSKVRIDVEKVKPYYLSLIEKYFPAKLKW 305
+KYDLYSKSKVRIDVE+VKPYYLSLI++YFPAKLKW
Sbjct: 189 GIYTCSKYDLYSKSKVRIDVERVKPYYLSLIDQYFPAKLKW 311
>BE474715
Length = 154
Score = 78.2 bits (191), Expect = 4e-15
Identities = 35/50 (70%), Positives = 42/50 (84%)
Frame = +2
Query: 68 YQSVDFVKKMREEYGKLNRVEMSMWECCELLNEVVDESDPDLDEPQIEHL 117
YQS+D KKM +EY LNRVEMS+WECC+LLN+VVD+ DPDLD+P IE L
Sbjct: 5 YQSIDLEKKMIDEYA*LNRVEMSIWECCDLLNDVVDDMDPDLDDP*IETL 154
>TC207601 similar to UP|Q9MA30 (Q9MA30) T5E21.2, partial (20%)
Length = 694
Score = 57.4 bits (137), Expect = 8e-09
Identities = 22/31 (70%), Positives = 28/31 (89%)
Frame = +3
Query: 192 YNTRYGMYSEKCGLNNVLMSWGHDDYMYLVA 222
YNT+ G+Y E CGL+NV+MSWGHD+YMYLV+
Sbjct: 3 YNTKKGIYDEGCGLDNVMMSWGHDEYMYLVS 95
Score = 54.7 bits (130), Expect(2) = 7e-14
Identities = 34/91 (37%), Positives = 42/91 (45%), Gaps = 41/91 (45%)
Frame = +1
Query: 240 HSFYALHREGAYKHLMNEEDFENLEWLHIF------------------------------ 269
++ AL++ GAYKHL +EED +NL+WL IF
Sbjct: 421 YNLAALYQTGAYKHLYSEEDVKNLKWLEIFQ*ANYFCFRF*HISLYLVFNLT**IHNGTD 600
Query: 270 -----------NKYDLYSKSKVRIDVEKVKP 289
+KYDLYSKS V IDVEKVKP
Sbjct: 601 YVSYKLCDTKHSKYDLYSKSNVLIDVEKVKP 693
Score = 39.7 bits (91), Expect(2) = 7e-14
Identities = 18/23 (78%), Positives = 19/23 (82%)
Frame = +3
Query: 221 VAKENKTTLPSAALFIIRYHSFY 243
V K N TTLPS ALFIIRYHSF+
Sbjct: 252 VTKGNNTTLPSEALFIIRYHSFH 320
>BM086270
Length = 413
Score = 40.8 bits (94), Expect = 8e-04
Identities = 17/23 (73%), Positives = 20/23 (86%)
Frame = +2
Query: 142 DLGKVLLLPVFGGLPQWAVVGDT 164
DLGK+L LP FG LPQWAVVG++
Sbjct: 290 DLGKILALPSFGELPQWAVVGES 358
>TC226809 similar to GB|AAP37699.1|30725354|BT008340 At5g20700 {Arabidopsis
thaliana;} , partial (27%)
Length = 1148
Score = 33.1 bits (74), Expect = 0.16
Identities = 14/39 (35%), Positives = 23/39 (58%)
Frame = +3
Query: 8 PDHFGSDVEGKTVPIDEKELVLDGGFVMPHTNSFGHTFR 46
P H GSD G+ P +++++V G PH +FGH ++
Sbjct: 111 PQHQGSDHVGEEAPANDRKVVGVVGLERPHGGTFGHHWK 227
>AW596821
Length = 430
Score = 28.5 bits (62), Expect = 4.0
Identities = 13/47 (27%), Positives = 23/47 (48%), Gaps = 2/47 (4%)
Frame = +3
Query: 150 PVFGGLPQWAVVGDTFPVGCRFDESIVHHKHFKENPD--YNNPAYNT 194
P G +P W + T+ CR+ + + H+ F +P N P Y++
Sbjct: 120 PT*G*VPWWTLSSPTWTKPCRYPHTTISHQTFPISPPPWKNTPGYDS 260
>BM308800
Length = 429
Score = 27.3 bits (59), Expect = 8.8
Identities = 9/14 (64%), Positives = 12/14 (85%)
Frame = -2
Query: 256 NEEDFENLEWLHIF 269
NEE F +++WLHIF
Sbjct: 392 NEETFSHMDWLHIF 351
>BE610370
Length = 303
Score = 27.3 bits (59), Expect = 8.8
Identities = 14/45 (31%), Positives = 25/45 (55%)
Frame = +2
Query: 132 DWLHLTGLIHDLGKVLLLPVFGGLPQWAVVGDTFPVGCRFDESIV 176
+W+++ G++ L K+ LL +FGG+ W + T P C +V
Sbjct: 17 EWVNIQGVVILLVKMNLLAIFGGVNIWVHI--TSPDICSMGSLLV 145
>BQ453609 homologue to GP|13605607|gb| T3B23.2/T3B23.2 {Arabidopsis
thaliana}, partial (11%)
Length = 405
Score = 27.3 bits (59), Expect = 8.8
Identities = 11/30 (36%), Positives = 20/30 (66%)
Frame = -2
Query: 270 NKYDLYSKSKVRIDVEKVKPYYLSLIEKYF 299
N++++ SK VRI++ +K YYL + + F
Sbjct: 140 NEFEV*SKGYVRINICFIKTYYLQFLHQLF 51
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.139 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,258,898
Number of Sequences: 63676
Number of extensions: 235124
Number of successful extensions: 1046
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 1037
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1044
length of query: 305
length of database: 12,639,632
effective HSP length: 97
effective length of query: 208
effective length of database: 6,463,060
effective search space: 1344316480
effective search space used: 1344316480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0213.1


