

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0209.3
(1547 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
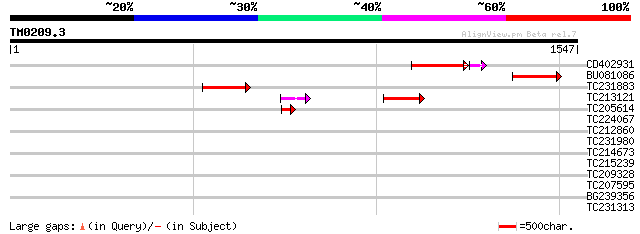
Score E
Sequences producing significant alignments: (bits) Value
CD402931 similar to GP|9081793|dbj P0489A01.23 {Oryza sativa (ja... 166 4e-46
BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza s... 139 1e-32
TC231883 weakly similar to UP|Q7QBQ3 (Q7QBQ3) AgCP4056 (Fragment... 128 2e-29
TC213121 103 6e-22
TC205614 54 5e-07
TC224067 38 0.030
TC212860 weakly similar to UP|PIF1_YEAST (P07271) DNA repair and... 37 0.051
TC231980 similar to UP|Q8S562 (Q8S562) KAP-2, partial (19%) 33 0.74
TC214673 homologue to UP|Q9LNS1 (Q9LNS1) F1L3.2, partial (5%) 32 1.7
TC215239 UP|H3_PEA (P02300) Histone H3, partial (39%) 32 2.2
TC209328 similar to UP|KAPS_CATRO (O49204) Adenylyl-sulfate kina... 31 3.7
TC207595 similar to GB|AAG45492.1|12061244|AY013245 36I5.4 {Oryz... 31 4.8
BG239356 weakly similar to GP|974782|emb| cobalamine-independent... 31 4.8
TC231313 UP|Q9FXZ2 (Q9FXZ2) Resistance protein LM12 (Fragment), ... 30 8.2
>CD402931 similar to GP|9081793|dbj P0489A01.23 {Oryza sativa (japonica
cultivar-group)}, partial (5%)
Length = 617
Score = 166 bits (419), Expect(2) = 4e-46
Identities = 91/156 (58%), Positives = 112/156 (71%)
Frame = +2
Query: 1096 GTGKTFVWNTLSAAVRSKGLIVLNVASSGIASLLLPGGRTAHSRFSILISINEVSTCNLR 1155
GT KTF+W TLSA +RSKGL+ + +ASLLLP G TAHS FSI + I + STCN+
Sbjct: 11 GTSKTFLWKTLSAGMRSKGLMSSSCLKR-LASLLLPRG*TAHSTFSIPLVIKDDSTCNIN 187
Query: 1156 QGSPKAELLQKASLIIWDETPMMNKHCFEALDRSLNDVMKTRANYGNDRPFRGKVVVLGG 1215
QGS +++LL LIIWDETP+MNK CFEALDR+L D+M ++ +PF GK VVLGG
Sbjct: 188 QGSARSKLLLHTKLIIWDETPVMNKFCFEALDRTLQDIMASQNKDNATKPFGGK-VVLGG 364
Query: 1216 DFRQILPVISKGSRADMVGSTVTSSYLWKYCKVMKL 1251
DFRQILPVI KGSR D+VGS + +S KV+KL
Sbjct: 365 DFRQILPVIRKGSRQDIVGSAINASK-----KVLKL 457
Score = 39.3 bits (90), Expect(2) = 4e-46
Identities = 21/48 (43%), Positives = 29/48 (59%)
Frame = +1
Query: 1254 NMRLQSASS*TSATEIREFAQWILKVGDETVDTIDEDETTIEIPSDLL 1301
NMRL + ++ +IREF WIL + D D +EDE I+IP +LL
Sbjct: 463 NMRLGTTNNNEDRDDIREFVDWILMIRDGNRD--EEDEGEIDIPKNLL 600
>BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza sativa
(japonica cultivar-group)}, partial (7%)
Length = 426
Score = 139 bits (349), Expect = 1e-32
Identities = 70/137 (51%), Positives = 93/137 (67%), Gaps = 3/137 (2%)
Frame = +2
Query: 1371 DEDSEVHAEWF---TSEFLNDVQCSGIPNHRLILKERVPIMLLRNIDQAGGLCNGTRLRV 1427
++ + +E F T+EF+N + S +PNH++ LK IMLLRN+DQ GLCN TRL +
Sbjct: 14 EKSDTIESEGFSTITTEFINSLSTSRLPNHKIKLKVDSRIMLLRNLDQNEGLCNSTRLII 193
Query: 1428 THLTQYIIVATVLSGIRLGKTEYIPRITLTPSDSGLPFKFSRRQFPVTLCFAMTINKSQG 1487
T +II A ++SG G T YIPR+ +PS S PFK RR+FP+ + +AMTINKSQG
Sbjct: 194 TMFADHIIEAKIMSGKGQGNTVYIPRLATSPSQSPWPFKLIRRKFPIIVSYAMTINKSQG 373
Query: 1488 QSLSHVGLYLPRPVFTH 1504
Q L+ VGLYLP PVF+H
Sbjct: 374 QLLASVGLYLPTPVFSH 424
>TC231883 weakly similar to UP|Q7QBQ3 (Q7QBQ3) AgCP4056 (Fragment), partial
(17%)
Length = 1263
Score = 128 bits (321), Expect = 2e-29
Identities = 62/132 (46%), Positives = 87/132 (64%)
Frame = +1
Query: 525 FPAGMPTIEFQKRGFPHAHILLWL*PQHKIKTGDDIDKHISAELPDPKLYPKLYKAVSSY 584
F M TIEFQKRG PH +LL+L P ++ + D+ID ISAE+P + P+LY V ++
Sbjct: 865 FKEVMHTIEFQKRGLPHIQLLLFLHPDNQYPSSDEIDHIISAEIPSHEDDPELYTLVQNH 1044
Query: 585 MIHGPCGPIDPKSVCMVDGKCSKHFPKKFQNCTTVDDDGFPIYKRRTTRITVTKKGVPLD 644
M+HGPCG + S CM +GKCS+ +PK F T +D +G+ +Y+RR T+ K V +D
Sbjct: 1045MVHGPCGILQSHSPCMKEGKCSRFYPKIFLPNTLLDLNGYLVYRRRNDGRTILKNSVIVD 1224
Query: 645 NGFVVPYNPKLL 656
N ++VPYN KLL
Sbjct: 1225NRYLVPYNAKLL 1260
>TC213121
Length = 713
Score = 103 bits (257), Expect = 6e-22
Identities = 50/114 (43%), Positives = 73/114 (63%)
Frame = +2
Query: 1019 ARSLKDFPSLSYPTFAEIMNFENKFVADELNYNKEEMSKIHEELVRSLTGEQKAAYKKVL 1078
A S ++FP +SYP + + NK + +EL Y+K+ ++ SLT E + + K++
Sbjct: 269 ANSHRNFPLMSYPIGYIVNPYGNKLIYNELAYDKDILAA*FSRCYHSLTDEHASIFNKIM 448
Query: 1079 DVVMSNKGGFLFLYGFGGTGKTFVWNTLSAAVRSKGLIVLNVASSGIASLLLPG 1132
++ + GG FLYG+GGTGKTFVW TLS+ + S IVL +ASSGI S+LLPG
Sbjct: 449 HMIATQSGGVYFLYGYGGTGKTFVWKTLSSTIHSNSGIVLTMASSGILSMLLPG 610
Score = 50.4 bits (119), Expect = 6e-06
Identities = 32/82 (39%), Positives = 43/82 (52%)
Frame = +3
Query: 738 IVGAKCRYLTPCEATWRTFKFDIHDRWPPVVRLGFHLPNQQKVLFKESDDFEEVVQKCSR 797
+V C Y++P E + F +H R P V L FHL NQ KES ++ K +
Sbjct: 15 LVNVICIYVSPPETC*KISAFPMHGRAPTVECLYFHLENQ*P--*KESQQIGSMLSKSTI 188
Query: 798 KETKFLAWMKANKRYPEGRNLT 819
KE+ F A M +NK Y GR+LT
Sbjct: 189 KESIFTACMDSNKMYHHGRDLT 254
>TC205614
Length = 742
Score = 53.9 bits (128), Expect = 5e-07
Identities = 22/36 (61%), Positives = 27/36 (74%)
Frame = +2
Query: 743 CRYLTPCEATWRTFKFDIHDRWPPVVRLGFHLPNQQ 778
C+Y++PCEA WR F F I+ R P V RL FHLPN+Q
Sbjct: 632 CQYISPCEACWRIFSFQIYGRIPIVERLYFHLPNEQ 739
>TC224067
Length = 415
Score = 38.1 bits (87), Expect = 0.030
Identities = 23/41 (56%), Positives = 30/41 (73%), Gaps = 2/41 (4%)
Frame = -2
Query: 1476 LCFAMTINKSQGQSLSHV-GLYLPRPV-FTHGQLYVALSRV 1514
+C+ M INKS GQ+LS V G+ LPRP +HGQ YV L++V
Sbjct: 360 VCYTMKINKSHGQTLSQVGGVLLPRPN**SHGQ-YVTLNQV 241
>TC212860 weakly similar to UP|PIF1_YEAST (P07271) DNA repair and recombination
protein PIF1, mitochondrial precursor, partial (8%)
Length = 596
Score = 37.4 bits (85), Expect = 0.051
Identities = 22/43 (51%), Positives = 29/43 (67%)
Frame = +2
Query: 1478 FAMTINKSQGQSLSHVGLYLPRPVFTHGQLYVALSRVKSRKGL 1520
+AM+I+K QG +L V L R F G +YVALSRV+S +GL
Sbjct: 5 WAMSIHKCQGMTLERVHTDLSR-AFGCGMVYVALSRVRSLEGL 130
>TC231980 similar to UP|Q8S562 (Q8S562) KAP-2, partial (19%)
Length = 590
Score = 33.5 bits (75), Expect = 0.74
Identities = 12/36 (33%), Positives = 23/36 (63%)
Frame = -3
Query: 927 ILTTNAMSRPKEVWEKTWRILSDGILYERRKALNIP 962
+L T S+ ++VW++TW ++D I+Y +K+ P
Sbjct: 582 LLLTATXSKSEQVWDQTWHWMADDIVYNYKKSSTSP 475
>TC214673 homologue to UP|Q9LNS1 (Q9LNS1) F1L3.2, partial (5%)
Length = 445
Score = 32.3 bits (72), Expect = 1.7
Identities = 14/31 (45%), Positives = 21/31 (67%)
Frame = -2
Query: 280 KHMLLSFLFSTRFFSRMVKTDFEKISPSVSC 310
KH L+ FLFST ++ KTD++ +S +V C
Sbjct: 444 KHHLVFFLFSTILLTKNTKTDYQGLSINVHC 352
>TC215239 UP|H3_PEA (P02300) Histone H3, partial (39%)
Length = 340
Score = 32.0 bits (71), Expect = 2.2
Identities = 12/18 (66%), Positives = 16/18 (88%)
Frame = -3
Query: 1472 FPVTLCFAMTINKSQGQS 1489
FP+ +CFAMT NKS+GQ+
Sbjct: 326 FPLIVCFAMTTNKSEGQT 273
>TC209328 similar to UP|KAPS_CATRO (O49204) Adenylyl-sulfate kinase,
chloroplast precursor (APS kinase)
(Adenosine-5'phosphosulfate kinase) (ATP
adenosine-5'-phosphosulfate 3'-phosphotransferase) ,
partial (69%)
Length = 1056
Score = 31.2 bits (69), Expect = 3.7
Identities = 25/90 (27%), Positives = 40/90 (43%)
Frame = +1
Query: 1081 VMSNKGGFLFLYGFGGTGKTFVWNTLSAAVRSKGLIVLNVASSGIASLLLPGGRTAHSRF 1140
++ KG ++L G G+GKT + LS ++ SKG + S +L G H
Sbjct: 235 LLQQKGCVIWLTGLSGSGKTTIACALSRSLHSKGKL----------SYILDGDNIRHGLN 384
Query: 1141 SILISINEVSTCNLRQGSPKAELLQKASLI 1170
L E + N+R+ A+L A +I
Sbjct: 385 QDLSFRAEDRSENIRRIGEVAKLFADAGVI 474
>TC207595 similar to GB|AAG45492.1|12061244|AY013245 36I5.4 {Oryza sativa
(japonica cultivar-group);} , partial (12%)
Length = 607
Score = 30.8 bits (68), Expect = 4.8
Identities = 15/29 (51%), Positives = 19/29 (64%)
Frame = +3
Query: 152 FFFLPVSIYPSFLNYISSNICNYMFLCCV 180
FFF+ +SI+ SFLN I SN+ FL V
Sbjct: 498 FFFI*MSIFLSFLNQIKSNVSYLYFLTLV 584
>BG239356 weakly similar to GP|974782|emb| cobalamine-independent methionine
synthase {Solenostemon scutellarioides}, partial (15%)
Length = 382
Score = 30.8 bits (68), Expect = 4.8
Identities = 18/59 (30%), Positives = 30/59 (50%)
Frame = +2
Query: 402 FCHRLLPGDIAICLTIARMQWPYARNLATRTSS*HSLAIQLGVKYKGFYNLGITSRRTD 460
+ HRLL G + + +A + W Y RN R + + +A + + + F GIT +TD
Sbjct: 47 YTHRLLNGMLTV--PVATLNWNYVRNDQPRFETTYHIASCIKDEVEDFEKAGITVIQTD 217
>TC231313 UP|Q9FXZ2 (Q9FXZ2) Resistance protein LM12 (Fragment), complete
Length = 1314
Score = 30.0 bits (66), Expect = 8.2
Identities = 38/149 (25%), Positives = 65/149 (43%), Gaps = 13/149 (8%)
Frame = +1
Query: 1039 FENKFVADELNYNKEEMSKIHEELVRSLTGEQKAAY--KKVLDVVMSNKGGFLFLYGFGG 1096
+E KF+ + + + + H + L G + K +LDV + + ++G G
Sbjct: 481 YEYKFIKEIVESVSNKFNHDHLHVSDVLVGLESPVLEVKSLLDVGRDDVVHMVGIHGLAG 660
Query: 1097 TGKTFVWNTLSAAVRSKGLIVLNVASSGIASLLLPG-GRTAHS--------RFSILISIN 1147
GKT TL+ AV + ++A AS L RT+++ F + +
Sbjct: 661 VGKT----TLAIAVYN------SIAGHFEASCFLENVKRTSNTINGLEKLQSFLLSKTAG 810
Query: 1148 EVSTCNLRQGSP--KAELLQKASLIIWDE 1174
E+ N R+G P K +L QK L+I D+
Sbjct: 811 EIKLTNWREGIPIIKRKLKQKKILLILDD 897
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.331 0.145 0.455
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 71,987,638
Number of Sequences: 63676
Number of extensions: 1066122
Number of successful extensions: 5751
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 5651
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5747
length of query: 1547
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1437
effective length of database: 5,635,272
effective search space: 8097885864
effective search space used: 8097885864
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0209.3


