

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.9
(529 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
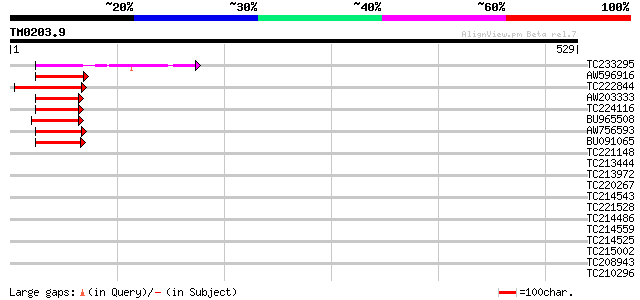
Sequences producing significant alignments: (bits) Value
TC233295 similar to UP|Q7KVQ0 (Q7KVQ0) CG4038-PA, partial (6%) 65 7e-11
AW596916 54 1e-07
TC222844 weakly similar to UP|Q9FFW4 (Q9FFW4) Similarity to heat... 52 5e-07
AW203333 50 2e-06
TC224116 49 7e-06
BU965508 49 7e-06
AW756593 48 9e-06
BU091065 45 1e-04
TC221148 similar to UP|Q8W104 (Q8W104) AT3g54650/T5N23_10, parti... 37 0.022
TC213444 37 0.028
TC213972 similar to UP|Q8W104 (Q8W104) AT3g54650/T5N23_10, parti... 36 0.037
TC220267 similar to UP|Q6K235 (Q6K235) F-box-like protein, parti... 34 0.18
TC214543 32 0.70
TC221528 weakly similar to UP|Q6YIA0 (Q6YIA0) Disease resistance... 31 1.6
TC214486 31 1.6
TC214559 31 1.6
TC214525 weakly similar to UP|Q9LU84 (Q9LU84) Arabidopsis thalia... 31 1.6
TC215002 similar to UP|O48601 (O48601) NADPH:isoflavone reductas... 30 2.7
TC208943 weakly similar to UP|DR30_ARATH (Q9LZ25) Probable disea... 30 2.7
TC210296 weakly similar to UP|DR29_ARATH (Q9SZA7) Probable disea... 29 5.9
>TC233295 similar to UP|Q7KVQ0 (Q7KVQ0) CG4038-PA, partial (6%)
Length = 557
Score = 65.1 bits (157), Expect = 7e-11
Identities = 50/156 (32%), Positives = 68/156 (43%), Gaps = 2/156 (1%)
Frame = +3
Query: 25 DRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTIDPHAFSSRNLQS 84
DRIS LPD +L ILSLLP K A T ILSKRW+ LW LDF + S
Sbjct: 66 DRISHLPDVLLLQILSLLPTKQAVITGILSKRWRPLWPAVSVLDF----------DDESS 215
Query: 85 SDFETSGHPLASTMDFITQVLSTRDKDS--DIRALCFRARLSFSRLNSLIRSAVRHNVKE 142
+F G L +F+ VL D + R C S + + + R +
Sbjct: 216 PEFHHPG-GLTGFAEFVYSVLLLHDAPAIERFRLRCANPNYSARDIATWLCHVARRRAER 392
Query: 143 LDIEVTPEVRTDEYFNFPRCLIASKTLRVLKLRSGF 178
+++ ++ Y PRCL T+ V+KL F
Sbjct: 393 VELSLS----LSRYVALPRCLFHCDTVSVMKLNGVF 488
>AW596916
Length = 455
Score = 54.3 bits (129), Expect = 1e-07
Identities = 25/49 (51%), Positives = 32/49 (65%)
Frame = +3
Query: 25 DRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRTID 73
DR+SDLPD VL HI+ + +K A QT +LSKRWK LW +L + D
Sbjct: 264 DRLSDLPDFVLLHIMKFMSMKHAVQTCVLSKRWKELWKRLTNLALHSSD 410
>TC222844 weakly similar to UP|Q9FFW4 (Q9FFW4) Similarity to heat shock
transcription factor HSF30, partial (6%)
Length = 746
Score = 52.4 bits (124), Expect = 5e-07
Identities = 25/67 (37%), Positives = 42/67 (62%)
Frame = +1
Query: 5 SAKRKKMAQIMENEAKAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTF 64
S ++K +A+ ++ E + DRIS LPD++L H+++ + K+A QT +LSKRW L
Sbjct: 436 SERKKNIAREVDGENEDHRPDRISALPDSLLFHMINFMDTKSAVQTCVLSKRWNDLSKCL 615
Query: 65 PDLDFRT 71
+L F +
Sbjct: 616 TNLTFNS 636
>AW203333
Length = 418
Score = 50.1 bits (118), Expect = 2e-06
Identities = 23/45 (51%), Positives = 30/45 (66%)
Frame = +1
Query: 25 DRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDF 69
D IS LP +LH ILS +P + A +TS+LSK W W+T+P L F
Sbjct: 85 DMISTLPKTILHDILSRMPEEDAVRTSVLSKSWAEAWSTYPILYF 219
>TC224116
Length = 520
Score = 48.5 bits (114), Expect = 7e-06
Identities = 21/45 (46%), Positives = 30/45 (66%)
Frame = +2
Query: 25 DRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDF 69
D ISDLP +++ IL LPI+ A +TSILS +W++ W + L F
Sbjct: 287 DLISDLPQSIIESILVQLPIRDAVRTSILSSKWRYKWASITQLVF 421
>BU965508
Length = 447
Score = 48.5 bits (114), Expect = 7e-06
Identities = 22/49 (44%), Positives = 31/49 (62%)
Frame = +2
Query: 21 AAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDF 69
A D ISDLP +++ IL LPI+ A +TSILS +W++ W + L F
Sbjct: 284 AMGPDLISDLPQSIIESILVQLPIRDAVRTSILSSKWRYKWASITRLVF 430
>AW756593
Length = 370
Score = 48.1 bits (113), Expect = 9e-06
Identities = 21/47 (44%), Positives = 29/47 (61%)
Frame = +3
Query: 25 DRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFRT 71
D+IS+LPD +L H++ + + A QT +LSKRW LW L F T
Sbjct: 63 DKISELPDNILLHMMDFMDTREAVQTCVLSKRWNNLWKRLSTLLFNT 203
>BU091065
Length = 460
Score = 44.7 bits (104), Expect = 1e-04
Identities = 18/46 (39%), Positives = 31/46 (67%)
Frame = +3
Query: 25 DRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFLWTTFPDLDFR 70
+RI DLP V+ IL LP + +TS+L++ W+++W++ P +FR
Sbjct: 90 NRIGDLPMNVIVSILQRLPFQDLVKTSVLARAWRYMWSSTPRREFR 227
>TC221148 similar to UP|Q8W104 (Q8W104) AT3g54650/T5N23_10, partial (26%)
Length = 1001
Score = 37.0 bits (84), Expect = 0.022
Identities = 22/53 (41%), Positives = 27/53 (50%), Gaps = 2/53 (3%)
Frame = +3
Query: 221 LRNLHLQLCFGLRVLRVGCLALRDLSLEKCINLH--GLDVVTPKLERLRVSEC 271
L+ L L C GL L + C L DL+L C NLH L + P LE + S C
Sbjct: 324 LKKLSLWGCSGLDALYLNCPQLNDLNLNSCRNLHPERLLLQCPSLENVHASGC 482
>TC213444
Length = 530
Score = 36.6 bits (83), Expect = 0.028
Identities = 15/36 (41%), Positives = 24/36 (66%)
Frame = +2
Query: 12 AQIMENEAKAAATDRISDLPDAVLHHILSLLPIKTA 47
+Q ++N + DR+SD+PD ++HHILS + K A
Sbjct: 419 SQNVDNCEMEESEDRLSDMPDCIIHHILSFMETKDA 526
>TC213972 similar to UP|Q8W104 (Q8W104) AT3g54650/T5N23_10, partial (24%)
Length = 858
Score = 36.2 bits (82), Expect = 0.037
Identities = 22/53 (41%), Positives = 27/53 (50%), Gaps = 2/53 (3%)
Frame = +1
Query: 221 LRNLHLQLCFGLRVLRVGCLALRDLSLEKCINLH--GLDVVTPKLERLRVSEC 271
L+ L L C GL L + C L DL+L C NLH L + P LE + S C
Sbjct: 283 LKKLSLWGCTGLDALYLNCPRLIDLNLNSCSNLHPGRLLLQCPNLENVHASGC 441
>TC220267 similar to UP|Q6K235 (Q6K235) F-box-like protein, partial (7%)
Length = 586
Score = 33.9 bits (76), Expect = 0.18
Identities = 18/56 (32%), Positives = 28/56 (49%), Gaps = 7/56 (12%)
Frame = +2
Query: 13 QIMENEAKAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRW-------KFLW 61
++ E K A + LPD +L IL+ LPI + + +SKRW +F+W
Sbjct: 287 ELSEEGEKEATAVSLDVLPDDLLERILAYLPIASIFRAGCVSKRWHEIVNSERFVW 454
>TC214543
Length = 687
Score = 32.0 bits (71), Expect = 0.70
Identities = 18/55 (32%), Positives = 28/55 (50%)
Frame = +3
Query: 219 PFLRNLHLQLCFGLRVLRVGCLALRDLSLEKCINLHGLDVVTPKLERLRVSECFH 273
P L++L L C L +R +L LSL C + L++ P LE++ + C H
Sbjct: 24 PMLKSLVLDNCESLTSVRFISTSLVSLSLGGCRAITSLELTCPNLEKVILDGCDH 188
>TC221528 weakly similar to UP|Q6YIA0 (Q6YIA0) Disease resistance
protein-like protein MsR1, partial (32%)
Length = 720
Score = 30.8 bits (68), Expect = 1.6
Identities = 24/89 (26%), Positives = 37/89 (40%), Gaps = 6/89 (6%)
Frame = +1
Query: 189 TFQSLHTLSLTLGPVDHQAPLSDLFTESAFPFLRNLHLQLCFGLRVLRVGCLALRDLSLE 248
T ++L LSL L ++ AFP L +L++ C L L G + L +
Sbjct: 400 TLKNLKKLSLFLCNTRQAFENGNMLISDAFPNLEDLNIDYCKDLIELPKGVCDITSLKML 579
Query: 249 KCINLHGLDVVTPK------LERLRVSEC 271
N H L + + L+ LR+S C
Sbjct: 580 SITNCHKLSALPQQFGNLENLKLLRLSSC 666
>TC214486
Length = 1070
Score = 30.8 bits (68), Expect = 1.6
Identities = 17/48 (35%), Positives = 25/48 (51%)
Frame = +1
Query: 224 LHLQLCFGLRVLRVGCLALRDLSLEKCINLHGLDVVTPKLERLRVSEC 271
L+L L L+ + V CL L L+L C +L L + P+L L + C
Sbjct: 556 LNLSLSANLKEVDVACLNLSWLNLSNCSSLEVLKLECPRLTSLFLQSC 699
>TC214559
Length = 762
Score = 30.8 bits (68), Expect = 1.6
Identities = 17/48 (35%), Positives = 25/48 (51%)
Frame = +1
Query: 224 LHLQLCFGLRVLRVGCLALRDLSLEKCINLHGLDVVTPKLERLRVSEC 271
L+L L L+ + V CL L L+L C +L L + P+L L + C
Sbjct: 190 LNLSLSANLKEVDVACLNLSWLNLSNCSSLEVLKLECPRLTSLFLQSC 333
>TC214525 weakly similar to UP|Q9LU84 (Q9LU84) Arabidopsis thaliana genomic
DNA, chromosome 3, P1 clone: MPE11, partial (20%)
Length = 458
Score = 30.8 bits (68), Expect = 1.6
Identities = 14/63 (22%), Positives = 32/63 (50%)
Frame = +1
Query: 1 METRSAKRKKMAQIMENEAKAAATDRISDLPDAVLHHILSLLPIKTAAQTSILSKRWKFL 60
++ + K + + I + A AT + D + + ++ PIK +T++L +W+ +
Sbjct: 211 IDAKIIKEEMLEAIAPLDRGADATPQDQQTIDQIARELEAVTPIKEPLKTNLLDGKWELI 390
Query: 61 WTT 63
+TT
Sbjct: 391 YTT 399
>TC215002 similar to UP|O48601 (O48601) NADPH:isoflavone reductase , complete
Length = 1359
Score = 30.0 bits (66), Expect = 2.7
Identities = 12/35 (34%), Positives = 19/35 (54%)
Frame = -3
Query: 357 LQKRITSMCSSPWLQMLSNSNFYIEPLYNLKSLEL 391
L + +C+ WLQ + S FY+ +N K L+L
Sbjct: 322 LPSHLPRLCAFQWLQWVPKSGFYLSQPWNWKKLKL 218
>TC208943 weakly similar to UP|DR30_ARATH (Q9LZ25) Probable disease
resistance protein At5g04720, partial (38%)
Length = 1320
Score = 30.0 bits (66), Expect = 2.7
Identities = 33/129 (25%), Positives = 48/129 (36%), Gaps = 9/129 (6%)
Frame = +3
Query: 149 PEVRTDEYFNFPRCLIASKTLRVLKLRSGFRLP------PSSIMKETFQSLHTLSLTLGP 202
P +R N+ + V K S R P + ++L L + L
Sbjct: 798 PNLRALIIINYSATYACLLNVSVFKNLSNLRSPWLEKVSTPELSSIVLENLGKLFIVLCK 977
Query: 203 VDHQAPLSDLFTESAFPFLRNLHLQLCFGLRVLR---VGCLALRDLSLEKCINLHGLDVV 259
V++ ++ FP L L L C L L G +L++LSL C NL L V
Sbjct: 978 VNNSLVEKEVDLAQVFPNLLELTLDHCDDLTQLPSSICGMKSLQNLSLTNCHNLTELPVE 1157
Query: 260 TPKLERLRV 268
KL L +
Sbjct: 1158LGKLRSLEI 1184
>TC210296 weakly similar to UP|DR29_ARATH (Q9SZA7) Probable disease
resistance protein At4g33300, partial (13%)
Length = 794
Score = 28.9 bits (63), Expect = 5.9
Identities = 26/92 (28%), Positives = 38/92 (41%), Gaps = 3/92 (3%)
Frame = +2
Query: 180 LPPSSIMKETFQSLHTLSLTLGPVDHQAPLSDLFTESAFPFLRNLHLQLCFGLRVLR--- 236
+P + ++L L + L V+ ++ FP L L L C L L
Sbjct: 62 VPXPELSSIVLENLGKLFIVLCKVNXXXVEKEVDLAQVFPNLLELTLDHCDDLIQLPSSI 241
Query: 237 VGCLALRDLSLEKCINLHGLDVVTPKLERLRV 268
G +L++LSL C NL L V KL L +
Sbjct: 242 CGMKSLQNLSLTNCHNLTQLPVELGKLRSLEI 337
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.134 0.397
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,069,300
Number of Sequences: 63676
Number of extensions: 365734
Number of successful extensions: 1982
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 1973
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1981
length of query: 529
length of database: 12,639,632
effective HSP length: 102
effective length of query: 427
effective length of database: 6,144,680
effective search space: 2623778360
effective search space used: 2623778360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0203.9


