

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.8
(124 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
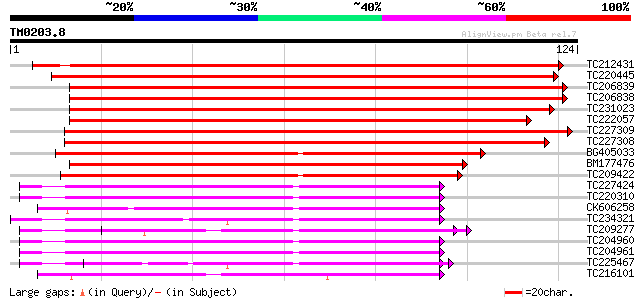
Score E
Sequences producing significant alignments: (bits) Value
TC212431 weakly similar to UP|Q9LLM1 (Q9LLM1) EF-hand Ca2+-bindi... 201 9e-53
TC220445 weakly similar to UP|Q9LLM1 (Q9LLM1) EF-hand Ca2+-bindi... 159 4e-40
TC206839 similar to UP|Q8LG35 (Q8LG35) EF-hand Ca2+-binding prot... 157 1e-39
TC206838 similar to GB|AAR16085.1|38304364|AY363867 KIC-related ... 155 6e-39
TC231023 weakly similar to UP|Q9LLM1 (Q9LLM1) EF-hand Ca2+-bindi... 149 4e-37
TC222057 weakly similar to GB|AAR16085.1|38304364|AY363867 KIC-r... 131 9e-32
TC227309 similar to GB|AAR17001.1|38325077|AY363866 KCBP interac... 130 2e-31
TC227308 similar to GB|AAR17001.1|38325077|AY363866 KCBP interac... 124 1e-29
BG405033 similar to GP|16648777|gb| AT4g27280/M4I22_90 {Arabidop... 119 4e-28
BM177476 weakly similar to GP|16648777|gb| AT4g27280/M4I22_90 {A... 119 4e-28
TC209422 weakly similar to UP|Q8LG35 (Q8LG35) EF-hand Ca2+-bindi... 115 7e-27
TC227424 UP|Q39890 (Q39890) Calmodulin, complete 48 1e-06
TC220310 homologue to UP|Q39890 (Q39890) Calmodulin, partial (61%) 47 2e-06
CK606258 47 3e-06
TC234321 similar to UP|Q40982 (Q40982) Calmodulin-like protein, ... 45 7e-06
TC209277 homologue to UP|Q43447 (Q43447) Calmodulin, complete 45 1e-05
TC204960 UP|Q6LEC4 (Q6LEC4) Calmodulin, complete 44 1e-05
TC204961 UP|Q6LEC4 (Q6LEC4) Calmodulin, complete 44 1e-05
TC225467 UP|O49184 (O49184) Calmodulin, complete 44 1e-05
TC216101 similar to UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, ... 44 1e-05
>TC212431 weakly similar to UP|Q9LLM1 (Q9LLM1) EF-hand Ca2+-binding protein
CCD1, partial (76%)
Length = 437
Score = 201 bits (510), Expect = 9e-53
Identities = 97/116 (83%), Positives = 110/116 (94%)
Frame = +1
Query: 6 TVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQD 65
T+ G G FEDLLPVMANKLGGEGL+KELCNGF++L+DK++GVITLESL+ NAA+LGLQD
Sbjct: 16 TMAGSG--FEDLLPVMANKLGGEGLMKELCNGFELLVDKEKGVITLESLRRNAALLGLQD 189
Query: 66 MKEDELVSMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHELDNN 121
+KEDELVSM+REGDLDGDGALTQMEFCVLMFRLSP+LMEESW WLEEALQHEL+NN
Sbjct: 190 LKEDELVSMMREGDLDGDGALTQMEFCVLMFRLSPDLMEESWFWLEEALQHELNNN 357
>TC220445 weakly similar to UP|Q9LLM1 (Q9LLM1) EF-hand Ca2+-binding protein
CCD1, partial (70%)
Length = 684
Score = 159 bits (401), Expect = 4e-40
Identities = 77/111 (69%), Positives = 92/111 (82%)
Frame = +3
Query: 10 KGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKED 69
K F D LPVMANKLGG+GLI ELCNGF++L D DRGVIT ESLK NAA+LGLQ + ++
Sbjct: 60 KPSHFHDYLPVMANKLGGDGLIDELCNGFNLLKDSDRGVITFESLKRNAALLGLQGLGDE 239
Query: 70 ELVSMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHELDN 120
+L SM+ +GD DGDGAL+Q+EFCVLMFRLSPELME S +WLEE LQ EL++
Sbjct: 240 DLRSMLGQGDFDGDGALSQLEFCVLMFRLSPELMEGSKLWLEEVLQQELEH 392
>TC206839 similar to UP|Q8LG35 (Q8LG35) EF-hand Ca2+-binding protein CCD1,
partial (74%)
Length = 552
Score = 157 bits (397), Expect = 1e-39
Identities = 75/109 (68%), Positives = 90/109 (81%)
Frame = +3
Query: 14 FEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVS 73
F+D LPVMANKLGG+GLI ELCNGFD+L D D+GVIT ESLK N+ +LGLQ + +++L S
Sbjct: 102 FQDYLPVMANKLGGDGLIDELCNGFDLLKDSDKGVITFESLKRNSYLLGLQGLSDEDLRS 281
Query: 74 MIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHELDNNY 122
M+ EGD DGDGAL QMEFCVLMFRLSPELME S +WLE+ LQ EL + +
Sbjct: 282 MVLEGDFDGDGALNQMEFCVLMFRLSPELMEGSKMWLEQVLQQELKDYF 428
>TC206838 similar to GB|AAR16085.1|38304364|AY363867 KIC-related protein
{Arabidopsis thaliana;} , partial (72%)
Length = 631
Score = 155 bits (391), Expect = 6e-39
Identities = 73/109 (66%), Positives = 90/109 (81%)
Frame = +1
Query: 14 FEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVS 73
F+D LPVMANKLGG+GLI ELCNGF++L D D+GVIT ESLK N+A+ GLQ + +++L S
Sbjct: 94 FQDYLPVMANKLGGDGLIDELCNGFNLLKDSDKGVITFESLKRNSALFGLQGLSDEDLRS 273
Query: 74 MIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHELDNNY 122
M+ EGD DGDGAL Q+EFCVLMFRLSPELME S +WLE+ LQ EL + +
Sbjct: 274 MVLEGDFDGDGALNQLEFCVLMFRLSPELMEGSKMWLEQVLQQELKDYF 420
>TC231023 weakly similar to UP|Q9LLM1 (Q9LLM1) EF-hand Ca2+-binding protein
CCD1, partial (74%)
Length = 747
Score = 149 bits (375), Expect = 4e-37
Identities = 71/106 (66%), Positives = 87/106 (81%)
Frame = +2
Query: 14 FEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVS 73
F D LP+MANKLGG+GLI ELCNGF++L D R VIT ESLK NAA+LGLQ + +++L
Sbjct: 206 FHDYLPMMANKLGGDGLIDELCNGFNLLKDSHREVITFESLKRNAALLGLQGLGDEDLRL 385
Query: 74 MIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHELD 119
M+R+GD DGDG L+Q+EFCVLMFRLSPELME S +WLEE LQ +L+
Sbjct: 386 MLRQGDFDGDGVLSQLEFCVLMFRLSPELMEGSKLWLEEVLQQDLE 523
>TC222057 weakly similar to GB|AAR16085.1|38304364|AY363867 KIC-related
protein {Arabidopsis thaliana;} , partial (78%)
Length = 902
Score = 131 bits (329), Expect = 9e-32
Identities = 65/101 (64%), Positives = 77/101 (75%)
Frame = +3
Query: 14 FEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVS 73
FED P M KLG EG +KEL NGF +LMD+++ VIT ESLK N+A+LGL+ M +DEL
Sbjct: 315 FEDYFPAMMEKLGTEGFMKELTNGFQLLMDREKKVITFESLKKNSALLGLEGMNDDELRC 494
Query: 74 MIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEAL 114
M+REGDLDGDGAL +MEFC LMFRLSP LM S LEEA+
Sbjct: 495 MLREGDLDGDGALDEMEFCTLMFRLSPALMNNSKELLEEAI 617
>TC227309 similar to GB|AAR17001.1|38325077|AY363866 KCBP interacting
Ca2+-binding protein {Arabidopsis thaliana;} , partial
(81%)
Length = 670
Score = 130 bits (327), Expect = 2e-31
Identities = 60/111 (54%), Positives = 83/111 (74%)
Frame = +1
Query: 13 EFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELV 72
EFEDLLPVMA KL E + ELC GF +L D + G+IT ESL+TN+A+LG++ M +++
Sbjct: 127 EFEDLLPVMAEKLDVESFVSELCGGFKLLSDPETGLITSESLRTNSALLGMEGMSKEDAE 306
Query: 73 SMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHELDNNYS 123
+M+R+GDLDGDG L + EFC+LM RLSP +ME++ WLE+A+ EL + S
Sbjct: 307 AMVRQGDLDGDGKLNETEFCILMVRLSPGMMEDAESWLEKAIDEELSKSSS 459
>TC227308 similar to GB|AAR17001.1|38325077|AY363866 KCBP interacting
Ca2+-binding protein {Arabidopsis thaliana;} , partial
(77%)
Length = 611
Score = 124 bits (310), Expect = 1e-29
Identities = 58/106 (54%), Positives = 78/106 (72%)
Frame = +2
Query: 13 EFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELV 72
EFEDLLPVMA KL E + ELC GF++L D + G+IT ESL N+ +LG+ M ++E
Sbjct: 101 EFEDLLPVMAKKLDVETFVSELCGGFNLLADPETGLITGESLMRNSTLLGMDGMSKEEAE 280
Query: 73 SMIREGDLDGDGALTQMEFCVLMFRLSPELMEESWIWLEEALQHEL 118
+M+R+GDLDGDG L + EFC+LM RLSP +ME++ WL +AL+ EL
Sbjct: 281 TMVRQGDLDGDGKLNETEFCILMVRLSPGIMEDAEAWLHKALEQEL 418
>BG405033 similar to GP|16648777|gb| AT4g27280/M4I22_90 {Arabidopsis
thaliana}, partial (55%)
Length = 415
Score = 119 bits (298), Expect = 4e-28
Identities = 58/94 (61%), Positives = 73/94 (76%)
Frame = +3
Query: 11 GLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDE 70
G +FED P M +LG EG I ELCNGF +LMD + G+IT ESLK + +LGL ++++DE
Sbjct: 69 GFDFEDCFPSMIARLGAEGFIGELCNGFQLLMDVNMGLITFESLKMSTNLLGL-EVRDDE 245
Query: 71 LVSMIREGDLDGDGALTQMEFCVLMFRLSPELME 104
L+ M+ EGDLDGDGAL QMEFC+LMFRLSP LM+
Sbjct: 246 LLCMLMEGDLDGDGALNQMEFCILMFRLSPCLMD 347
>BM177476 weakly similar to GP|16648777|gb| AT4g27280/M4I22_90 {Arabidopsis
thaliana}, partial (66%)
Length = 425
Score = 119 bits (298), Expect = 4e-28
Identities = 57/87 (65%), Positives = 68/87 (77%)
Frame = +2
Query: 14 FEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVS 73
FED P M KLG EG +KEL NGF +LMD+++ VIT ESLK N+A+LGL+ M +DEL
Sbjct: 161 FEDYFPAMMEKLGTEGFMKELTNGFQLLMDREKKVITFESLKKNSALLGLEGMSDDELRC 340
Query: 74 MIREGDLDGDGALTQMEFCVLMFRLSP 100
M+REGDLDGD AL +MEFC LMFRLSP
Sbjct: 341 MLREGDLDGDDALDEMEFCTLMFRLSP 421
>TC209422 weakly similar to UP|Q8LG35 (Q8LG35) EF-hand Ca2+-binding protein
CCD1, partial (32%)
Length = 627
Score = 115 bits (287), Expect = 7e-27
Identities = 56/88 (63%), Positives = 69/88 (77%)
Frame = +3
Query: 12 LEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDEL 71
L+FED P M ++G EG I ELCNGF +LMD ++G+IT ESLK N +LGL D+ +DEL
Sbjct: 129 LDFEDYFPSMIARMGAEGFIGELCNGFRLLMDVNKGLITFESLKLNCYLLGL-DVMDDEL 305
Query: 72 VSMIREGDLDGDGALTQMEFCVLMFRLS 99
M+ EGDLDGDGAL+QMEFC+LMFRLS
Sbjct: 306 ACMLMEGDLDGDGALSQMEFCILMFRLS 389
>TC227424 UP|Q39890 (Q39890) Calmodulin, complete
Length = 1424
Score = 47.8 bits (112), Expect = 1e-06
Identities = 30/93 (32%), Positives = 47/93 (50%)
Frame = +3
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ EF++ L +MA K+ +EL F + G I+ L+ LG
Sbjct: 834 GNGTI-----EFDEFLSLMAKKVKDTDAEEELKEAFKVFDKDQNGYISASELRHVMINLG 998
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + ++E+ MI+E DLDGDG + EF +M
Sbjct: 999 -EKLTDEEVEQMIKEADLDGDGQVNYEEFVKMM 1094
Score = 36.2 bits (82), Expect = 0.004
Identities = 26/56 (46%), Positives = 31/56 (54%), Gaps = 1/56 (1%)
Frame = +3
Query: 41 LMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
L DKD G IT+E L T L Q+ E+EL MI E D DG+G + EF LM
Sbjct: 711 LFDKDGDGCITVEELATVIRSLD-QNPTEEELQDMISEVDADGNGTIEFDEFLSLM 875
>TC220310 homologue to UP|Q39890 (Q39890) Calmodulin, partial (61%)
Length = 542
Score = 47.4 bits (111), Expect = 2e-06
Identities = 30/93 (32%), Positives = 47/93 (50%)
Frame = +1
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ EF++ L +MA K+ +EL F + G I+ L+ LG
Sbjct: 4 GNGTI-----EFDEFLSLMAKKVKDTDAEEELKEAFKVFDKDQNGYISASELRHVMINLG 168
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + ++E+ MI+E DLDGDG + EF +M
Sbjct: 169-EKLTDEEVEQMIKEADLDGDGQVNYDEFVKMM 264
>CK606258
Length = 627
Score = 46.6 bits (109), Expect = 3e-06
Identities = 31/90 (34%), Positives = 49/90 (54%), Gaps = 1/90 (1%)
Frame = +2
Query: 7 VNGKG-LEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQD 65
++G G ++FE+ L +M K+ E L +EL F + G I+ L+ LG +
Sbjct: 269 IDGNGSIDFEEFLNIMGRKMK-ETLAEELREAFKVFDRDQNGYISATELRHVMMNLG-ER 442
Query: 66 MKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ ++E MIRE DLDGDG ++ EF +M
Sbjct: 443 LTDEEAEQMIREADLDGDGQVSFEEFSRIM 532
Score = 36.2 bits (82), Expect = 0.004
Identities = 26/66 (39%), Positives = 36/66 (54%), Gaps = 1/66 (1%)
Frame = +2
Query: 41 LMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
L+DKD G IT++ L T L KE E+ MI E D+DG+G++ EF +M R
Sbjct: 152 LIDKDSDGFITVDELATIIRSLEGNPTKE-EIQDMISEVDIDGNGSIDFEEFLNIMGRKM 328
Query: 100 PELMEE 105
E + E
Sbjct: 329 KETLAE 346
>TC234321 similar to UP|Q40982 (Q40982) Calmodulin-like protein, partial
(96%)
Length = 626
Score = 45.4 bits (106), Expect = 7e-06
Identities = 32/96 (33%), Positives = 51/96 (52%), Gaps = 1/96 (1%)
Frame = +3
Query: 1 MAGTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDR-GVITLESLKTNAA 59
M G GT+ EF + L +MA K+ +EL F + DKD G I+ L++
Sbjct: 240 MDGNGTI-----EFGEFLNLMARKMKETEAEEELKEAF-RVFDKDHDGYISPSELRSVMR 401
Query: 60 VLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+G + + ++E+ M++E DLDGDG + EF +M
Sbjct: 402 TIG-EKVTDEEVEQMVKEADLDGDGLIDYEEFVRMM 506
Score = 34.7 bits (78), Expect = 0.012
Identities = 26/66 (39%), Positives = 34/66 (51%), Gaps = 1/66 (1%)
Frame = +3
Query: 41 LMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRLS 99
L DKD G IT+E L T L ++ +EL M+ E D+DG+G + EF LM R
Sbjct: 123 LFDKDGDGCITIEELSTAIRSLD-ENPTVEELQIMMNEVDMDGNGTIEFGEFLNLMARKM 299
Query: 100 PELMEE 105
E E
Sbjct: 300 KETEAE 317
>TC209277 homologue to UP|Q43447 (Q43447) Calmodulin, complete
Length = 873
Score = 44.7 bits (104), Expect = 1e-05
Identities = 29/96 (30%), Positives = 47/96 (48%)
Frame = +3
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ EF + L +MA K+ ++L F + G I+ L+ LG
Sbjct: 246 GNGTI-----EFVEFLNLMAKKMKETDAEEDLKEAFKVFDKDQNGYISASELRHVMINLG 410
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLMFRL 98
+ + ++E+ MI+E DLDGDG + EF +M +
Sbjct: 411 -EKLTDEEVEQMIKEADLDGDGQVNYDEFVKMMMTI 515
Score = 38.5 bits (88), Expect = 8e-04
Identities = 34/83 (40%), Positives = 42/83 (49%), Gaps = 2/83 (2%)
Frame = +3
Query: 21 MANKLGGE--GLIKELCNGFDMLMDKDRGVITLESLKTNAAVLGLQDMKEDELVSMIREG 78
MA+ L E G IKE FD D G IT+E L T L Q+ E+EL MI E
Sbjct: 69 MADVLSEEQIGEIKEAFGLFDKDGD---GCITVEELATVIRSLD-QNPTEEELQDMINEV 236
Query: 79 DLDGDGALTQMEFCVLMFRLSPE 101
D DG+G + +EF LM + E
Sbjct: 237 DTDGNGTIEFVEFLNLMAKKMKE 305
>TC204960 UP|Q6LEC4 (Q6LEC4) Calmodulin, complete
Length = 1012
Score = 44.3 bits (103), Expect = 1e-05
Identities = 29/93 (31%), Positives = 46/93 (49%)
Frame = +2
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ +F + L +MA K+ +EL F + G I+ L+ LG
Sbjct: 266 GNGTI-----DFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLG 430
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + ++E+ MIRE D+DGDG + EF +M
Sbjct: 431 -EKLTDEEVDEMIREADVDGDGQINYEEFVKVM 526
Score = 37.4 bits (85), Expect = 0.002
Identities = 34/78 (43%), Positives = 40/78 (50%), Gaps = 1/78 (1%)
Frame = +2
Query: 21 MANKLGGEGLIKELCNGFDMLMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGD 79
MA++L E I E F L DKD G IT + L T LG Q+ E EL MI E D
Sbjct: 89 MADQLTDEQ-ISEFKEAFS-LFDKDGDGCITTKELGTVMRSLG-QNPTEAELQDMINEVD 259
Query: 80 LDGDGALTQMEFCVLMFR 97
DG+G + EF LM R
Sbjct: 260 ADGNGTIDFPEFLNLMAR 313
>TC204961 UP|Q6LEC4 (Q6LEC4) Calmodulin, complete
Length = 928
Score = 44.3 bits (103), Expect = 1e-05
Identities = 29/93 (31%), Positives = 46/93 (49%)
Frame = +2
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ +F + L +MA K+ +EL F + G I+ L+ LG
Sbjct: 272 GNGTI-----DFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLG 436
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + ++E+ MIRE D+DGDG + EF +M
Sbjct: 437 -EKLTDEEVDEMIREADVDGDGQINYEEFVKVM 532
Score = 37.4 bits (85), Expect = 0.002
Identities = 34/78 (43%), Positives = 40/78 (50%), Gaps = 1/78 (1%)
Frame = +2
Query: 21 MANKLGGEGLIKELCNGFDMLMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMIREGD 79
MA++L E I E F L DKD G IT + L T LG Q+ E EL MI E D
Sbjct: 95 MADQLTDEQ-ISEFKEAFS-LFDKDGDGCITTKELGTVMRSLG-QNPTEAELQDMINEVD 265
Query: 80 LDGDGALTQMEFCVLMFR 97
DG+G + EF LM R
Sbjct: 266 ADGNGTIDFPEFLNLMAR 319
>TC225467 UP|O49184 (O49184) Calmodulin, complete
Length = 1048
Score = 44.3 bits (103), Expect = 1e-05
Identities = 29/93 (31%), Positives = 46/93 (49%)
Frame = +3
Query: 3 GTGTVNGKGLEFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
G GT+ +F + L +MA K+ +EL F + G I+ L+ LG
Sbjct: 303 GNGTI-----DFPEFLNLMARKMKDTDSEEELKEAFRVFDKDQNGFISAAELRHVMTNLG 467
Query: 63 LQDMKEDELVSMIREGDLDGDGALTQMEFCVLM 95
+ + ++E+ MIRE D+DGDG + EF +M
Sbjct: 468 -EKLTDEEVDEMIREADVDGDGQINYEEFVKVM 563
Score = 38.5 bits (88), Expect = 8e-04
Identities = 35/82 (42%), Positives = 42/82 (50%), Gaps = 1/82 (1%)
Frame = +3
Query: 17 LLPVMANKLGGEGLIKELCNGFDMLMDKDR-GVITLESLKTNAAVLGLQDMKEDELVSMI 75
L P MA++L + I E F L DKD G IT + L T LG Q+ E EL MI
Sbjct: 114 LSPPMADQLTDDQ-IAEFKEAFS-LFDKDGDGCITTKELGTVMRSLG-QNPTEAELQDMI 284
Query: 76 REGDLDGDGALTQMEFCVLMFR 97
E D DG+G + EF LM R
Sbjct: 285 NEVDADGNGTIDFPEFLNLMAR 350
>TC216101 similar to UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, partial
(65%)
Length = 796
Score = 44.3 bits (103), Expect = 1e-05
Identities = 35/94 (37%), Positives = 47/94 (49%), Gaps = 5/94 (5%)
Frame = +1
Query: 7 VNGKGL----EFEDLLPVMANKLGGEGLIKELCNGFDMLMDKDRGVITLESLKTNAAVLG 62
VNG G EF DL + + E ++E N FD D G IT+E L+ A LG
Sbjct: 370 VNGDGCVDMDEFGDLYESIMEERDEEEDMREAFNVFDQNRD---GFITVEELRRVLASLG 540
Query: 63 LQDMKE-DELVSMIREGDLDGDGALTQMEFCVLM 95
L+ + DE MI + D+DGDG + EF +M
Sbjct: 541 LKQGRTLDECKKMITKVDVDGDGMVNYKEFRQMM 642
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.138 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,567,138
Number of Sequences: 63676
Number of extensions: 44845
Number of successful extensions: 327
Number of sequences better than 10.0: 75
Number of HSP's better than 10.0 without gapping: 304
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 320
length of query: 124
length of database: 12,639,632
effective HSP length: 85
effective length of query: 39
effective length of database: 7,227,172
effective search space: 281859708
effective search space used: 281859708
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 53 (25.0 bits)
Lotus: description of TM0203.8


