

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0203.7
(824 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
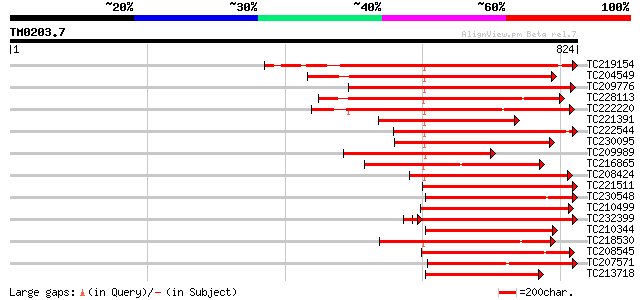
Score E
Sequences producing significant alignments: (bits) Value
TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.1... 401 e-112
TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-l... 392 e-109
TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-l... 383 e-106
TC228113 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kin... 350 1e-96
TC222220 weakly similar to UP|Q9ZP16 (Q9ZP16) Receptor-like prot... 339 3e-93
TC221391 similar to UP|O81906 (O81906) Serine/threonine kinase-l... 299 3e-81
TC222544 weakly similar to UP|O23743 (O23743) SFR1 protein precu... 299 3e-81
TC230095 weakly similar to UP|Q9XED4 (Q9XED4) Receptor-like prot... 272 5e-73
TC209989 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kin... 270 2e-72
TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial... 269 4e-72
TC208424 weakly similar to UP|O65466 (O65466) Serine/threonine k... 266 2e-71
TC221511 similar to UP|Q40096 (Q40096) Receptor protein kinase, ... 265 5e-71
TC230548 weakly similar to UP|Q8LLI4 (Q8LLI4) S-locus receptor-l... 255 5e-68
TC210499 weakly similar to UP|Q9LDS6 (Q9LDS6) Serine/threonine k... 248 8e-66
TC232399 similar to UP|Q39086 (Q39086) Receptor kinase , partial... 246 2e-65
TC210344 similar to UP|Q9AVE0 (Q9AVE0) SRKb, partial (21%) 241 7e-64
TC218530 similar to UP|Q9SGU0 (Q9SGU0) T6H22.8.1 protein, partia... 238 8e-63
TC208545 weakly similar to UP|Q9S972 (Q9S972) ARK2 product/recep... 235 7e-62
TC207571 weakly similar to UP|O65470 (O65470) Serine/threonine k... 228 1e-59
TC213718 similar to UP|Q70I29 (Q70I29) S-receptor kinase-like pr... 226 2e-59
>TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.110 precursor
- Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (31%)
Length = 1363
Score = 401 bits (1030), Expect = e-112
Identities = 222/464 (47%), Positives = 297/464 (63%), Gaps = 10/464 (2%)
Frame = +1
Query: 371 CYAYSYEEYEKARQGDSVDPNAICWIWSQDLNNLE--EEYEGGCDLHVRVAFSDIEGDSY 428
C AY+ A G C +W DL +++ E G LH+R+ S++E
Sbjct: 4 CMAYTNYNISGAGSG--------CVMWFGDLLDIKLYSVAESGRRLHIRLPPSELE---- 147
Query: 429 QTERNLSSPTIILVTFTAVVFLIILSSTCIYLRRRRQAKIRGIKLYGSERYVRDLIESGR 488
+ ++ S II+ T A ++L+ IY R + D ++ +
Sbjct: 148 -SIKSKKSSKIIIGTSVAAPLGVVLAICFIYRRN-----------------IADKSKTKK 273
Query: 489 LQEDDAKAIDIPHFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCS 548
+ + +D+P F + +I AT+NF + NK+G+GGFGPVYKGK GGQEIAVKRLSS S
Sbjct: 274 SIDRQLQDVDVPLFDMLTITAATDNFLLNNKIGEGGFGPVYKGKLVGGQEIAVKRLSSLS 453
Query: 549 GQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD------- 601
GQG+ EF EV LIA+LQHRNLV+LLG C++G EK+LVYEY+ N SL++FIFD
Sbjct: 454 GQGITEFITEVKLIAKLQHRNLVKLLGCCIKGQEKLLVYEYVVNGSLNSFIFDQIKSKLL 633
Query: 602 -WDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKD 660
W RF IILGIARGLLYLH+DSRLRIIHRDLKASN+LLDE+ NPKISDFG+AR FGG
Sbjct: 634 DWPRRFNIILGIARGLLYLHQDSRLRIIHRDLKASNVLLDEKLNPKISDFGMARAFGGDQ 813
Query: 661 TVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLL 720
T GNT+RVVGTYGYM+PEYA DG+FS+KSDVFSFG+++LEI+ G +N L+L+
Sbjct: 814 TEGNTNRVVGTYGYMAPEYAFDGNFSIKSDVFSFGILLLEIVCGIKNKSLCHENQTLNLV 993
Query: 721 GYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSE 780
GYAW LWK L ID + +C E ++C++V LLC+Q+ P +RPTM++V+ MLGSE
Sbjct: 994 GYAWALWKEQNALQLIDSGIKDSCVIPEVLRCIHVSLLCVQQYPEDRPTMTSVIQMLGSE 1173
Query: 781 SNTLPSPREPAFVIRRCPSSRASTSSKMETFSRNDLTVTLENGR 824
+ + P+EP F RR + E S ++LT++L +GR
Sbjct: 1174MD-MVEPKEPGFFPRRI----LKEGNLKEMTSNDELTISLFSGR 1290
>TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-like protein,
partial (54%)
Length = 2032
Score = 392 bits (1008), Expect = e-109
Identities = 198/369 (53%), Positives = 256/369 (68%), Gaps = 8/369 (2%)
Frame = +3
Query: 434 LSSPTIILVTFTAVVFLIILSSTCIYLRRRRQAKIRGIKLYGSERYVRDLIESGRLQEDD 493
+S+ TI+ + V ++I +L RR + K +G G Y D
Sbjct: 627 ISAGTIVAIVVPITVAVLIFIVGICFLSRRARKKQQGSVKEGKTAY-------------D 767
Query: 494 AKAIDIPHFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLE 553
+D F +I ATN F+ NKLG+GGFG VYKG GQ +AVKRLS SGQG E
Sbjct: 768 IPTVDSLQFDFSTIEAATNKFSADNKLGEGGFGEVYKGTLSSGQVVAVKRLSKSSGQGGE 947
Query: 554 EFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMR 605
EFKNEVV++A+LQHRNLVRLLG+C++G+EK+LVYEY+PN+SLD +FD W R
Sbjct: 948 EFKNEVVVVAKLQHRNLVRLLGFCLQGEEKILVYEYVPNKSLDYILFDPEKQRELDWGRR 1127
Query: 606 FKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNT 665
+KII GIARG+ YLHEDSRLRIIHRDLKASNILLD + NPKISDFG+ARIFG T GNT
Sbjct: 1128YKIIGGIARGIQYLHEDSRLRIIHRDLKASNILLDGDMNPKISDFGMARIFGVDQTQGNT 1307
Query: 666 DRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWH 725
R+VGTYGYM+PEYA+ G FSVKSDV+SFGV+++EI+SGK+N+ FYQ + LL YAW
Sbjct: 1308SRIVGTYGYMAPEYAMHGEFSVKSDVYSFGVLLMEILSGKKNSSFYQTDGAEDLLSYAWQ 1487
Query: 726 LWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLP 785
LWK G L+ +D L ++ E ++ +++GLLC+QEDP +RPTM+ ++ ML S + TLP
Sbjct: 1488LWKDGTPLELMDPILRESYNQNEVIRSIHIGLLCVQEDPADRPTMATIVLMLDSNTVTLP 1667
Query: 786 SPREPAFVI 794
+P +PAF +
Sbjct: 1668TPTQPAFFV 1694
>TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-like protein
, partial (47%)
Length = 1305
Score = 383 bits (983), Expect = e-106
Identities = 188/338 (55%), Positives = 246/338 (72%), Gaps = 8/338 (2%)
Frame = +2
Query: 493 DAKAIDIPHFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGL 552
+ A++ F +I AT F+ ANKLG+GGFG VYKG P GQE+AVKRLS SGQG
Sbjct: 17 EISAVESLRFDFSTIEAATQKFSEANKLGEGGFGEVYKGLLPSGQEVAVKRLSKISGQGG 196
Query: 553 EEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDM 604
EEFKNEV ++A+LQHRNLVRLLG+C+EG+EK+LVYE++ N+SLD +FD W
Sbjct: 197 EEFKNEVEIVAKLQHRNLVRLLGFCLEGEEKILVYEFVANKSLDYILFDPEKQKSLDWTR 376
Query: 605 RFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGN 664
R+KI+ GIARG+ YLHEDSRL+IIHRDLKASN+LLD + NPKISDFG+ARIFG T N
Sbjct: 377 RYKIVEGIARGIQYLHEDSRLKIIHRDLKASNVLLDGDMNPKISDFGMARIFGVDQTQAN 556
Query: 665 TDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAW 724
T+R+VGTYGYMSPEYA+ G +S KSDV+SFGV++LEIISGKRN+ FY+ + LL YAW
Sbjct: 557 TNRIVGTYGYMSPEYAMHGEYSAKSDVYSFGVLILEIISGKRNSSFYETDVAEDLLSYAW 736
Query: 725 HLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTL 784
LWK L+ +DQ+L ++ E ++C+++GLLC+QEDPI+RPTM++V+ ML S S TL
Sbjct: 737 KLWKDEAPLELMDQSLRESYTRNEVIRCIHIGLLCVQEDPIDRPTMASVVLMLDSYSVTL 916
Query: 785 PSPREPAFVIRRCPSSRASTSSKMETFSRNDLTVTLEN 822
P +PAF I K++ + N + ++ +
Sbjct: 917 QVPNQPAFYINSRTEPNMPKGLKIDQSTTNSTSKSVND 1030
>TC228113 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kinase homolog
RK20-1, partial (53%)
Length = 1492
Score = 350 bits (898), Expect = 1e-96
Identities = 187/401 (46%), Positives = 253/401 (62%), Gaps = 44/401 (10%)
Frame = +1
Query: 450 LIILSSTCIYLRRRRQAKIRGIKLYGSERYVRDLIESGRLQEDDAKAIDIPHFHLESILD 509
++ L CIY RR + K +K + + D+ + F+ ++I
Sbjct: 4 VVPLICLCIYSRRSKARKSSLVKQHEDD--------------DEIEIAQSLQFNFDTIRV 141
Query: 510 ATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRN 569
AT +F+ +NKLGQGGFG VY+G+ GQ IAVKRLS S QG EFKNEV+L+A+LQHRN
Sbjct: 142 ATEDFSDSNKLGQGGFGAVYRGRLSDGQMIAVKRLSRESSQGDTEFKNEVLLVAKLQHRN 321
Query: 570 LVRLLGYCVEGDEKMLVYEYMPNRSLDAFIF----------------------------- 600
LVRLLG+C+EG E++L+YEY+PN+SLD FIF
Sbjct: 322 LVRLLGFCLEGKERLLIYEYVPNKSLDYFIFGRS**VHN*TVALYFTSGSGQRLNIHIPL 501
Query: 601 ---------------DWDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNP 645
+W+MR+KII G+ARGLLYLHEDS LRIIHRDLKASNILL+EE NP
Sbjct: 502 KLMAVYADPTKKAQLNWEMRYKIITGVARGLLYLHEDSHLRIIHRDLKASNILLNEEMNP 681
Query: 646 KISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGK 705
KI+DFG+AR+ T NT+R+VGTYGYM+PEYA+ G FS+KSDVFSFGV+VLEIISG
Sbjct: 682 KIADFGMARLVLMDQTQANTNRIVGTYGYMAPEYAMHGQFSMKSDVFSFGVLVLEIISGH 861
Query: 706 RNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPI 765
+N+G E+ LL +AW W+ G + +D +L+ + E ++C+++GLLC+QE+
Sbjct: 862 KNSGIRHGENVEDLLSFAWRNWREGTAVKIVDPSLNNNSR-NEMLRCIHIGLLCVQENLA 1038
Query: 766 ERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSS 806
+RPTM+ ++ ML S S +LP P +PAF + S ++T S
Sbjct: 1039DRPTMTTIMLMLNSYSLSLPIPSKPAFYVSSRTGSISATQS 1161
>TC222220 weakly similar to UP|Q9ZP16 (Q9ZP16) Receptor-like protein kinase,
RLK3 precursor, partial (35%)
Length = 1304
Score = 339 bits (870), Expect = 3e-93
Identities = 182/394 (46%), Positives = 248/394 (62%), Gaps = 12/394 (3%)
Frame = +3
Query: 439 IILVTFTAVVFLIILSSTCIYLRRRRQAKIRGIKLYGSERYVRDLIESGRLQE---DDAK 495
++L+ VV +I+L C ++ +R + K + L+E +++
Sbjct: 42 VVLIVVPVVVSIILLLCVCYFILKRSRKKYNTL-----------------LRENFGEESA 170
Query: 496 AIDIPHFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEF 555
++ F L +I ATN F+ ++G+GGFG VYKG P G+EIAVK+LS SGQG EF
Sbjct: 171 TLESLQFGLVTIEAATNKFSYEKRIGEGGFGVVYKGVLPDGREIAVKKLSKSSGQGANEF 350
Query: 556 KNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFK 607
KNE++LIA+LQHRNLV LLG+C+E EKML+YE++ N+SLD F+FD W R+K
Sbjct: 351 KNEILLIAKLQHRNLVTLLGFCLEEHEKMLIYEFVSNKSLDYFLFDSHRSKQLNWSERYK 530
Query: 608 IILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDR 667
II GIA+G+ YLHE SRL++IHRDLK SN+LLD NPKISDFG+ARI G T+R
Sbjct: 531 IIEGIAQGISYLHEHSRLKVIHRDLKPSNVLLDSNMNPKISDFGMARIVAIDQLQGKTNR 710
Query: 668 VVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLW 727
+VGTYGYMSPEYA+ G FS KSDVFSFGV+VLEIIS KRNT +H+ LL Y W W
Sbjct: 711 IVGTYGYMSPEYAMHGQFSEKSDVFSFGVIVLEIISAKRNTRSVFSDHD-DLLSYTWEQW 887
Query: 728 KVGRTLDFIDQTL-SQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPS 786
L+ DQ++ ++ C E +KC+ +GLLC+QE P +RP ++ V+ L S LP
Sbjct: 888 MDEAPLNIFDQSIEAEFCDHSEVVKCIQIGLLCVQEKPDDRPKITQVISYLNSSITELPL 1067
Query: 787 PREPAFVIRRCPSSRASTSSKMETFSRNDLTVTL 820
P++P SS T S N+++V++
Sbjct: 1068PKKPIRQSGIVQKIAVGESSSGSTPSINEMSVSI 1169
>TC221391 similar to UP|O81906 (O81906) Serine/threonine kinase-like protein
, partial (25%)
Length = 642
Score = 299 bits (766), Expect = 3e-81
Identities = 147/213 (69%), Positives = 176/213 (82%), Gaps = 8/213 (3%)
Frame = +2
Query: 536 GQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSL 595
G+E+AVKRLS S QGLEEFKNE+VLIA+LQHRNLVRLLG C++G+EK+LVYEY+PN+SL
Sbjct: 2 GEEVAVKRLSRKSSQGLEEFKNEMVLIAKLQHRNLVRLLGCCIQGEEKILVYEYLPNKSL 181
Query: 596 DAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKI 647
D F+FD W RF+II GIARGLLYLH DSRLRIIHRDLKASNILLDE NPKI
Sbjct: 182 DCFLFDPVKQTQLDWAKRFEIIEGIARGLLYLHRDSRLRIIHRDLKASNILLDESMNPKI 361
Query: 648 SDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRN 707
SDFGLARIFGG NT+RVVGTYGYMSPEYA++G FS+KSDV+SFGV++LEI+SG++N
Sbjct: 362 SDFGLARIFGGNQNEANTNRVVGTYGYMSPEYAMEGLFSIKSDVYSFGVLLLEIMSGRKN 541
Query: 708 TGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTL 740
T F + + SL+GYAWHLW R ++ +D +L
Sbjct: 542 TSFRDTD-DSSLIGYAWHLWSEQRVMELVDPSL 637
>TC222544 weakly similar to UP|O23743 (O23743) SFR1 protein precursor ,
partial (27%)
Length = 861
Score = 299 bits (766), Expect = 3e-81
Identities = 158/277 (57%), Positives = 196/277 (70%), Gaps = 10/277 (3%)
Frame = +3
Query: 558 EVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKII 609
EVV+I++LQHRNLVRLLG C+E DE+MLVYE+MPN+SLD+F+FD W RF II
Sbjct: 3 EVVVISKLQHRNLVRLLGCCIERDEQMLVYEFMPNKSLDSFLFDPLQRKILDWKKRFNII 182
Query: 610 LGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIF-GGKDTVGNTDRV 668
GIARG+LYLH DSRLRIIHRDLKASNILLD+E +PKISDFGLARI G D NT RV
Sbjct: 183 EGIARGILYLHRDSRLRIIHRDLKASNILLDDEMHPKISDFGLARIVRSGDDDEANTKRV 362
Query: 669 VGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWK 728
VGTYGYM PEYA++G FS KSDV+ V++LEI+SG+RNT FY E LSL YAW LW
Sbjct: 363 VGTYGYMPPEYAMEGIFSEKSDVYXXXVLLLEIVSGRRNTSFYNNEQSLSLXXYAWKLWN 542
Query: 729 VGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPR 788
G ID + + ++C+++GLLC+QE ERPT+S V+ ML SE LP PR
Sbjct: 543 EGNIKSIIDLEIQDPMFEKSILRCIHIGLLCVQELTKERPTISTVVXMLISEITHLPPPR 722
Query: 789 EPAFVIRR-CPSSRASTSSKMETFSRNDLTVTLENGR 824
+ FV ++ C SS +S S+ S N++T++ GR
Sbjct: 723 QXXFVQKQNCQSSESSQKSQFN--SNNNVTISEIQGR 827
>TC230095 weakly similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kinase
homolog RK20-1, partial (30%)
Length = 819
Score = 272 bits (695), Expect = 5e-73
Identities = 135/241 (56%), Positives = 174/241 (72%), Gaps = 8/241 (3%)
Frame = +3
Query: 560 VLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFD--------WDMRFKIILG 611
+LIA+LQH+NLVRL+G+C E EK+L+YEY+PN+SLD F+FD W RFKI+ G
Sbjct: 6 LLIAKLQHKNLVRLVGFCQEDREKILIYEYVPNKSLDHFLFDSQKHRQLTWSERFKIVKG 185
Query: 612 IARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGT 671
IARG+LYLHEDSRL+IIHRD+K SN+LLD NPKISDFG+AR+ G T+RVVGT
Sbjct: 186 IARGILYLHEDSRLKIIHRDIKPSNVLLDYGINPKISDFGMARMVATDQIQGCTNRVVGT 365
Query: 672 YGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGR 731
YGYMSPEYA+ G FS KSDVFSFGV+VL+IISGK+N+ ++ LL YAW+ W+
Sbjct: 366 YGYMSPEYAMHGQFSEKSDVFSFGVMVLDIISGKKNSCSFESCRVDDLLSYAWNNWRDES 545
Query: 732 TLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPA 791
+D TL ++ E KC+ +GLLC+QE+P +RPTM ++ L + S +P P EPA
Sbjct: 546 PYQLLDSTLLESYVPNEVEKCMQIGLLCVQENPDDRPTMGTIVSYLSNPSFEMPFPLEPA 725
Query: 792 F 792
F
Sbjct: 726 F 728
>TC209989 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kinase homolog
RK20-1, partial (32%)
Length = 738
Score = 270 bits (690), Expect = 2e-72
Identities = 138/230 (60%), Positives = 168/230 (73%), Gaps = 8/230 (3%)
Frame = +3
Query: 485 ESGRLQEDDAKAIDIPHFHLESILDATNNFAIANKLGQGGFGPVYKGKFPGGQEIAVKRL 544
+ G L D + F +I ATNNF+ ANKLGQGGFG VYKG GQEIA+KRL
Sbjct: 48 DEGELANDIKTDDQLLQFEFATIKFATNNFSDANKLGQGGFGIVYKGTLSDGQEIAIKRL 227
Query: 545 SSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDAFIFDW-- 602
S S QG EFKNE++L RLQHRNLVRLLG+C E++L+YE++PN+SLD FIFD
Sbjct: 228 SINSNQGETEFKNEILLTGRLQHRNLVRLLGFCFARRERLLIYEFVPNKSLDYFIFDPNK 407
Query: 603 ------DMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIF 656
++R+KII GIARGLLYLHEDSRL ++HRDLK SNILLD E NPKISDFG+AR+F
Sbjct: 408 RVNLN*EIRYKIIRGIARGLLYLHEDSRLNVVHRDLKTSNILLDGELNPKISDFGMARLF 587
Query: 657 GGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKR 706
T +T +VGT+GYM+PEY G FS+KSDVFSFGV++LEI+ G+R
Sbjct: 588 EINQTEASTTTIVGTFGYMAPEYIKHGQFSIKSDVFSFGVMILEIVCGQR 737
>TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial (28%)
Length = 1358
Score = 269 bits (687), Expect = 4e-72
Identities = 140/271 (51%), Positives = 187/271 (68%), Gaps = 9/271 (3%)
Frame = +1
Query: 516 IANKLGQGGFGPVYKGKFPGGQEIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLG 575
+ANK+G+GGFGPVYKG F G IAVK+LSS S QG EF NE+ +I+ LQH +LV+L G
Sbjct: 1 VANKIGEGGFGPVYKGCFSDGTLIAVKQLSSKSRQGNREFLNEIGMISALQHPHLVKLYG 180
Query: 576 YCVEGDEKMLVYEYMPNRSLDAFIF---------DWDMRFKIILGIARGLLYLHEDSRLR 626
CVEGD+ +LVYEYM N SL +F DW R+KI +GIARGL YLHE+SRL+
Sbjct: 181 CCVEGDQLLLVYEYMENNSLARALFGAEEHQIKLDWTTRYKICVGIARGLAYLHEESRLK 360
Query: 627 IIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFS 686
I+HRD+KA+N+LLD++ NPKISDFGLA++ +D + R+ GT+GYM+PEYA+ G+ +
Sbjct: 361 IVHRDIKATNVLLDQDLNPKISDFGLAKL-DEEDNTHISTRIAGTFGYMAPEYAMHGYLT 537
Query: 687 VKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKA 746
K+DV+SFG+V LEII+G+ NT Q E S+L +A L + G +D +D+ L
Sbjct: 538 DKADVYSFGIVALEIINGRSNTIHRQKEESFSVLEWAHLLREKGDIMDLVDRRLGLEFNK 717
Query: 747 EECMKCVNVGLLCLQEDPIERPTMSNVLFML 777
EE + + V LLC RPTMS+V+ ML
Sbjct: 718 EEALVMIKVALLCTNVTAALRPTMSSVVSML 810
>TC208424 weakly similar to UP|O65466 (O65466) Serine/threonine kinase-like
protein , partial (34%)
Length = 913
Score = 266 bits (681), Expect = 2e-71
Identities = 131/244 (53%), Positives = 172/244 (69%), Gaps = 8/244 (3%)
Frame = +2
Query: 582 EKMLVYEYMPNRSLDAFIFD--------WDMRFKIILGIARGLLYLHEDSRLRIIHRDLK 633
EK+L+YEY+ N+SLD F+FD W R+KII+GIARG+ YLHEDS+LRIIHRDLK
Sbjct: 2 EKILIYEYITNKSLDHFLFDPVKQKELDWSRRYKIIVGIARGIQYLHEDSQLRIIHRDLK 181
Query: 634 ASNILLDEEKNPKISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFS 693
ASN+LLDE NPKISDFG+A+IF T NT R+VGTYGYMSPEYA+ G FSVKSDVFS
Sbjct: 182 ASNVLLDENMNPKISDFGMAKIFQADQTQVNTGRIVGTYGYMSPEYAMRGQFSVKSDVFS 361
Query: 694 FGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCV 753
FGV+VLEI+SGK+NT FYQ H LL +AW W + L+ +D TL + E +C+
Sbjct: 362 FGVLVLEIVSGKKNTDFYQSNHADDLLSHAWKNWTLQTPLELLDPTLRGSYSRNEVNRCI 541
Query: 754 NVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSSKMETFSR 813
++GLLC+QE+P +RP+M+ + ML S S T+ P++PA ++R +R + + +
Sbjct: 542 HIGLLCVQENPSDRPSMATIALMLNSYSVTMSMPQQPASLLRGRGPNRLNQGMDSDQSTT 721
Query: 814 NDLT 817
N T
Sbjct: 722 NQST 733
>TC221511 similar to UP|Q40096 (Q40096) Receptor protein kinase, partial
(21%)
Length = 765
Score = 265 bits (678), Expect = 5e-71
Identities = 131/224 (58%), Positives = 167/224 (74%)
Frame = +3
Query: 601 DWDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKD 660
DW RF II GIA+GLLYLH+DSRLRIIHRDLKASN+LLD E NPKISDFG+ARIFG
Sbjct: 6 DWSKRFNIICGIAKGLLYLHQDSRLRIIHRDLKASNVLLDSELNPKISDFGMARIFGVDQ 185
Query: 661 TVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLL 720
GNT R+VGTYGYM+PEYA DG FSVKSDVFSFGV++LEIISGKR+ G+Y H +L+
Sbjct: 186 QEGNTKRIVGTYGYMAPEYATDGLFSVKSDVFSFGVLLLEIISGKRSRGYYNQNHSQNLI 365
Query: 721 GYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSE 780
G+AW LWK GR L+ ID+++ + + + C++V LLC+Q++P +RP MS+VL ML SE
Sbjct: 366 GHAWKLWKEGRPLELIDKSIEDSSSLSQMLHCIHVSLLCVQQNPEDRPGMSSVLLMLVSE 545
Query: 781 SNTLPSPREPAFVIRRCPSSRASTSSKMETFSRNDLTVTLENGR 824
LP P++P F + + +ST SK + S N++T+TL R
Sbjct: 546 LE-LPEPKQPGFFGKYSGEADSST-SKQQLSSTNEITITLLEAR 671
>TC230548 weakly similar to UP|Q8LLI4 (Q8LLI4) S-locus receptor-like kinase
RLK14, partial (20%)
Length = 692
Score = 255 bits (652), Expect = 5e-68
Identities = 121/220 (55%), Positives = 165/220 (75%)
Frame = +3
Query: 605 RFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGN 664
RF II GIARGLLYLH+DSRLRIIHRDLKASN+LLD E NPKISDFGLAR+ GG G
Sbjct: 3 RFGIINGIARGLLYLHQDSRLRIIHRDLKASNVLLDNEMNPKISDFGLARMCGGDQIEGE 182
Query: 665 TDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAW 724
T RVVGTYGYM+PEYA DG FS+KSDVFSFGV++LEI+SGK+N+ + P +L+G+AW
Sbjct: 183 TSRVVGTYGYMAPEYAFDGIFSIKSDVFSFGVLLLEIVSGKKNSRLFYPNDYNNLIGHAW 362
Query: 725 HLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTL 784
LWK G + FID +L +C E ++C+++GLLC+Q P +RP M++V+ +L +E N L
Sbjct: 363 MLWKEGNPMQFIDSSLEDSCILYEALRCIHIGLLCVQHHPNDRPNMASVVVLLSNE-NAL 539
Query: 785 PSPREPAFVIRRCPSSRASTSSKMETFSRNDLTVTLENGR 824
P P++P+++ + + R S+S + S ND+T+++ + +
Sbjct: 540 PLPKDPSYLSKDISTERESSSKNFTSVSINDVTMSMMSAK 659
>TC210499 weakly similar to UP|Q9LDS6 (Q9LDS6) Serine/threonine kinase-like
protein , partial (27%)
Length = 717
Score = 248 bits (633), Expect = 8e-66
Identities = 117/223 (52%), Positives = 164/223 (73%)
Frame = +3
Query: 597 AFIFDWDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIF 656
+ + DW RF II GI++G+LYLH+ SRL+IIHRDLKASNILLDE NPKISDFGLAR+F
Sbjct: 6 SMLLDWKKRFNIIEGISQGILYLHKYSRLKIIHRDLKASNILLDENMNPKISDFGLARMF 185
Query: 657 GGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHE 716
+++ G T R+VGTYGYMSPEYA++G FS KSDV+SFGV++LEI+SG++NT FY +H
Sbjct: 186 MQQESTGTTSRIVGTYGYMSPEYAMEGTFSTKSDVYSFGVLLLEIVSGRKNTSFYDVDHL 365
Query: 717 LSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFM 776
L+L+G+AW LW G +L +D +L+ + +E +C++VGLLC++ +RPTMSNV+ M
Sbjct: 366 LNLIGHAWELWNQGESLQLLDPSLNDSFDPDEVKRCIHVGLLCVEHYANDRPTMSNVISM 545
Query: 777 LGSESNTLPSPREPAFVIRRCPSSRASTSSKMETFSRNDLTVT 819
L +ES + PR PAF + R ++S ++ S ++ T +
Sbjct: 546 LTNESAPVTLPRRPAFYVERKNFDGKTSSKELCVDSTDEFTAS 674
>TC232399 similar to UP|Q39086 (Q39086) Receptor kinase , partial (23%)
Length = 1167
Score = 246 bits (629), Expect = 2e-65
Identities = 129/241 (53%), Positives = 168/241 (69%), Gaps = 2/241 (0%)
Frame = +2
Query: 586 VYEYMPNRSLDAFIFDWDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNP 645
VY P++S + DW R II GI RGLLYLH DSRL+IIHRDLKASN+LL E NP
Sbjct: 314 VYFLDPSKSK---LLDWRKRCGIIEGIGRGLLYLHRDSRLKIIHRDLKASNVLLYEALNP 484
Query: 646 KISDFGLARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGK 705
KISDFG+ARIFGG + NT+RVVGTYGYMSPEYA+ G FS KSDVFSFGV+V+EI+SG+
Sbjct: 485 KISDFGMARIFGGTEDQANTNRVVGTYGYMSPEYAMQGLFSEKSDVFSFGVLVIEIVSGR 664
Query: 706 RNTGFYQPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPI 765
RN+ FY ++ LSLLG+AW W+ G L ID + ++ ++C+++GLLC+QE +
Sbjct: 665 RNSRFYDDDNALSLLGFAWIQWREGNILSVIDPEIYDVTHHKDILRCIHIGLLCVQERAV 844
Query: 766 ERPTMSNVLFMLGSESNTLPSPREPAFVIRRCPSSRASTSS--KMETFSRNDLTVTLENG 823
+RPTM+ V+ ML SE LP P +PAFV + + S SS + + S N +++T G
Sbjct: 845 DRPTMAAVISMLNSEVAFLPPPDQPAFVQSQNMLNLVSVSSEERQKLCSINGISITDIRG 1024
Query: 824 R 824
R
Sbjct: 1025R 1027
Score = 48.9 bits (115), Expect = 9e-06
Identities = 21/28 (75%), Positives = 23/28 (82%)
Frame = +3
Query: 573 LLGYCVEGDEKMLVYEYMPNRSLDAFIF 600
L G C EGDEKML+YEYM N+SLD FIF
Sbjct: 3 LFGCCAEGDEKMLIYEYMLNKSLDVFIF 86
>TC210344 similar to UP|Q9AVE0 (Q9AVE0) SRKb, partial (21%)
Length = 870
Score = 241 bits (616), Expect = 7e-64
Identities = 115/192 (59%), Positives = 145/192 (74%)
Frame = +2
Query: 605 RFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGN 664
R II GIARG+LYLHEDSRL+IIHRDLKASN+LLD + NPKISDFG+ARIF G + N
Sbjct: 8 RLDIINGIARGILYLHEDSRLKIIHRDLKASNVLLDYDMNPKISDFGMARIFAGSEGEAN 187
Query: 665 TDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAW 724
T +VGTYGYM+PEYA++G +S+KSDVF FGV++LEII+GKRN GFY SLL YAW
Sbjct: 188 TATIVGTYGYMAPEYAMEGLYSIKSDVFGFGVLLLEIITGKRNAGFYHSNKTPSLLSYAW 367
Query: 725 HLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTL 784
HLW G+ L+ ID +C +E ++ +++GLLC+QED +RPTMS+V+ ML +ES TL
Sbjct: 368 HLWNEGKGLELIDPMSVDSCPGDEFLRYMHIGLLCVQEDAYDRPTMSSVVLMLKNESATL 547
Query: 785 PSPREPAFVIRR 796
P P F + R
Sbjct: 548 GQPERPPFSLGR 583
>TC218530 similar to UP|Q9SGU0 (Q9SGU0) T6H22.8.1 protein, partial (25%)
Length = 1190
Score = 238 bits (607), Expect = 8e-63
Identities = 126/262 (48%), Positives = 173/262 (65%), Gaps = 6/262 (2%)
Frame = +3
Query: 538 EIAVKRLSSCSGQGLEEFKNEVVLIARLQHRNLVRLLGYCVEGDEKMLVYEYMPNRSLDA 597
EIAVK+LS S QG +F E+ I+ +QHRNLV+L G C+EG +++LVYEY+ N+SLD
Sbjct: 3 EIAVKQLSVGSHQGKSQFITEIATISAVQHRNLVKLYGCCIEGSKRLLVYEYLENKSLDQ 182
Query: 598 FIF------DWDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFG 651
+F +W R+ I LG+ARGL YLHE+SRLRI+HRD+KASNILLD E PKISDFG
Sbjct: 183 ALFGKCLTLNWSTRYDICLGVARGLTYLHEESRLRIVHRDVKASNILLDYELIPKISDFG 362
Query: 652 LARIFGGKDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFY 711
LA+++ K T +T V GT GY++PEYA+ GH + K+DVFSFGVV LE++SG+ N+
Sbjct: 363 LAKLYDDKKTHISTG-VAGTIGYLAPEYAMRGHLTEKADVFSFGVVALELVSGRPNSDSS 539
Query: 712 QPEHELSLLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMS 771
++ LL +AW L + +D +D LS+ EE + V + LLC Q P RP+MS
Sbjct: 540 LEGEKVYLLEWAWQLHEKNCIIDLVDDRLSE-FNEEEVKRVVGIALLCTQTSPTLRPSMS 716
Query: 772 NVLFMLGSESNTLPSPREPAFV 793
V+ ML + +P ++
Sbjct: 717 RVVAMLSGDIEVSTVTSKPGYL 782
>TC208545 weakly similar to UP|Q9S972 (Q9S972) ARK2 product/receptor-like
serine/threonine protein kinase ARK2, partial (20%)
Length = 1253
Score = 235 bits (599), Expect = 7e-62
Identities = 115/223 (51%), Positives = 159/223 (70%)
Frame = +2
Query: 599 IFDWDMRFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGG 658
+ DW RF II GI++GLLYLH+ SRL++IHRDLKASNILLDE NPKISDFGLAR+F
Sbjct: 344 LLDWKKRFNIIEGISQGLLYLHKYSRLKVIHRDLKASNILLDENMNPKISDFGLARMFTR 523
Query: 659 KDTVGNTDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELS 718
+++ NT R+VGTYGYMSPEYA++G FSVKSDV+SFGV++LEI+SG+RNT FY + L+
Sbjct: 524 QESTTNTSRIVGTYGYMSPEYAMEGVFSVKSDVYSFGVLLLEIVSGRRNTSFYDGDRFLN 703
Query: 719 LLGYAWHLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLG 778
L+G+AW LW G L ID +L+++ +E +C+++GLLC++++ RP MS ++ ML
Sbjct: 704 LIGHAWELWNEGACLKLIDPSLTESPDLDEVQRCIHIGLLCVEQNANNRPLMSQIISML- 880
Query: 779 SESNTLPSPREPAFVIRRCPSSRASTSSKMETFSRNDLTVTLE 821
S N + P+ PAF +S++ T S +T + E
Sbjct: 881 SNKNPITLPQRPAFYFGSETFDGIISSTEFCTDSTKAITTSRE 1009
>TC207571 weakly similar to UP|O65470 (O65470) Serine/threonine kinase-like
protein , partial (23%)
Length = 1117
Score = 228 bits (580), Expect = 1e-59
Identities = 111/217 (51%), Positives = 152/217 (69%)
Frame = +2
Query: 608 IILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGNTDR 667
II GIA+GLLYLH+ SRL++IHRDLKASNILLD E N KISDFG+ARIFG + + NT+R
Sbjct: 2 IIGGIAQGLLYLHKYSRLKVIHRDLKASNILLDHEMNAKISDFGMARIFGVRVSEENTNR 181
Query: 668 VVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAWHLW 727
VVGTYGYM+PEYA+ G S+K+DVFSFGV++LEI+S K+N Y +H L+L+GY LW
Sbjct: 182 VVGTYGYMAPEYAMKGVVSIKTDVFSFGVLLLEILSSKKNNSRYHSDHPLNLIGY---LW 352
Query: 728 KVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFMLGSESNTLPSP 787
GR L+ ID TL+ C E +C+++GLLC+Q+ +RPTM +++ L +++ LP P
Sbjct: 353 NAGRALELIDSTLNGLCSQNEVFRCIHIGLLCVQDQATDRPTMVDIVSFLSNDTIQLPQP 532
Query: 788 REPAFVIRRCPSSRASTSSKMETFSRNDLTVTLENGR 824
+PA+ I ++ E S ND+T++ R
Sbjct: 533 MQPAYFINEVVEESELPYNQQEFHSENDVTISSTRAR 643
>TC213718 similar to UP|Q70I29 (Q70I29) S-receptor kinase-like protein 2,
partial (20%)
Length = 525
Score = 226 bits (577), Expect = 2e-59
Identities = 109/172 (63%), Positives = 130/172 (75%)
Frame = +1
Query: 605 RFKIILGIARGLLYLHEDSRLRIIHRDLKASNILLDEEKNPKISDFGLARIFGGKDTVGN 664
R +II GIARGLLYLH+DSRLRIIHRDLK SNILLD + NPKISDFGLAR FGG N
Sbjct: 10 RLQIIDGIARGLLYLHQDSRLRIIHRDLKVSNILLDNDMNPKISDFGLARTFGGDQAEAN 189
Query: 665 TDRVVGTYGYMSPEYALDGHFSVKSDVFSFGVVVLEIISGKRNTGFYQPEHELSLLGYAW 724
T+RV+GTYGYM PEYAL G FS+KSDVFSFGV+VLEIISG++N F EH L+LL +AW
Sbjct: 190 TNRVMGTYGYMPPEYALHGRFSIKSDVFSFGVIVLEIISGRKNRNFQDSEHHLNLLSHAW 369
Query: 725 HLWKVGRTLDFIDQTLSQTCKAEECMKCVNVGLLCLQEDPIERPTMSNVLFM 776
LW + L+ ID L E ++C++VGLLC+Q+ P RP MS+V+ M
Sbjct: 370 RLWIEEKPLELIDDLLDDPVSPHEILRCIHVGLLCVQQTPENRPNMSSVVLM 525
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.137 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 40,376,921
Number of Sequences: 63676
Number of extensions: 637672
Number of successful extensions: 5007
Number of sequences better than 10.0: 1005
Number of HSP's better than 10.0 without gapping: 4163
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4232
length of query: 824
length of database: 12,639,632
effective HSP length: 105
effective length of query: 719
effective length of database: 5,953,652
effective search space: 4280675788
effective search space used: 4280675788
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 63 (28.9 bits)
Lotus: description of TM0203.7


